Abstract
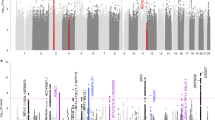
Very few genetic variants have been associated with depression and neuroticism, likely because of limitations on sample size in previous studies. Subjective well-being, a phenotype that is genetically correlated with both of these traits, has not yet been studied with genome-wide data. We conducted genome-wide association studies of three phenotypes: subjective well-being (n = 298,420), depressive symptoms (n = 161,460), and neuroticism (n = 170,911). We identify 3 variants associated with subjective well-being, 2 variants associated with depressive symptoms, and 11 variants associated with neuroticism, including 2 inversion polymorphisms. The two loci associated with depressive symptoms replicate in an independent depression sample. Joint analyses that exploit the high genetic correlations between the phenotypes (|ρ^| ≈ 0.8) strengthen the overall credibility of the findings and allow us to identify additional variants. Across our phenotypes, loci regulating expression in central nervous system and adrenal or pancreas tissues are strongly enriched for association.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
27 June 2016
In the version of this article initially published, the following co-authors and their affiliations were incorrectly omitted from the author list: Gudmar Thorleifsson, Sven Bergmann, Gyda Bjornsdottir, David C. Liewald, John M. Starr, Kari Stefansson and Unnur Thorsteinsdottir. In addition, the middle initial for co-author Andreas J. Forstner was also omitted. The errors have been corrected in the HTML and PDF versions of the article.
29 August 2016
In the version of this article initially published, one of the affiliations listed for author Maciej Trzaskowski, to the Department of Public Health, Faculty of Medicine, University of Split, Split, Croatia, was included in error. The correct affiliation for this author is the Queensland Brain Institute, University of Queensland, Brisbane, Queensland, Australia. The error has been corrected in the HTML and PDF versions of the article.
References
Kendler, K.S. & Myers, J. The genetic and environmental relationship between major depression and the five-factor model of personality. Psychol. Med. 40, 801–806 (2010).
Weiss, A., Bates, T.C. & Luciano, M. Happiness is a personal(ity) thing: the genetics of personality and well-being in a representative sample. Psychol. Sci. 19, 205–210 (2008).
Bartels, M., Cacioppo, J.T. & van Beijsterveldt, T.C.E.M. & Boomsma, D.I. Exploring the association between well-being and psychopathology in adolescents. Behav. Genet. 43, 177–190 (2013).
Genetics of Personality Consortium. Meta-analysis of genome-wide association studies for neuroticism, and the polygenic association with major depressive disorder. JAMA Psychiatry 72, 642–650 (2015).
Hyman, S. Mental health: depression needs large human-genetics studies. Nature 515, 189–191 (2014).
Rietveld, C.A. et al. Common genetic variants associated with cognitive performance identified using the proxy-phenotype method. Proc. Natl. Acad. Sci. USA 111, 13790–13794 (2014).
Kahneman, D. & Deaton, A. High income improves evaluation of life but not emotional well-being. Proc. Natl. Acad. Sci. USA 107, 16489–16493 (2010).
Kahneman, D. & Riis, J. in The Science of Well-Being (eds. Uppter, F., Baylis, N. & Keverne, B.) 285–301 (Oxford University Press, 2005).
Bartels, M. & Boomsma, D.I. Born to be happy? The etiology of subjective well-being. Behav. Genet. 39, 605–615 (2009).
Bulik-Sullivan, B.K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
Major Depressive Disorder Working Group of the Psychiatric GWAS Consortium. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Eysenck, H.J. & Eysenck, S.B.G. Manual of the Eysenck Personality Questionnaire (Hodder and Stroughton, 1975).
Tian, C. et al. Analysis and application of European genetic substructure using 300 K SNP information. PLoS Genet. 4, e4 (2008).
Steinberg, K.M. et al. Structural diversity and African origin of the 17q21.31 inversion polymorphism. Nat. Genet. 44, 872–880 (2012).
Smith, D.J. et al. Genome-wide analysis of over 106,000 individuals identifies 9 neuroticism-associated loci. bioRxiv doi:10.1101/032417 (2016).
Meuwissen, T.H., Hayes, B.J. & Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 157, 1819–1829 (2001).
Vilhjálmsson, B.J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Roberts, R.E., Kaplan, G.A., Shema, S.J. & Strawbridge, W.J. Are the obese at greater risk for depression? Am. J. Epidemiol. 152, 163–170 (2000).
Glassman, A.H. et al. Smoking, smoking cessation, and major depression. J. Am. Med. Assoc. 264, 1546–1549 (1990).
Shahab, L. & West, R. Differences in happiness between smokers, ex-smokers and never smokers: cross-sectional findings from a national household survey. Drug Alcohol Depend. 121, 38–44 (2012).
Rugulies, R. Depression as a predictor for coronary heart disease. a review and meta-analysis. Am. J. Prev. Med. 23, 51–61 (2002).
Finucane, H.K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Wood, A.R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Stetler, C. & Miller, G.E. Depression and hypothalamic-pituitary-adrenal activation: a quantitative summary of four decades of research. Psychosom. Med. 73, 114–126 (2011).
Seeman, P. Dopamine D2 receptors as treatment targets in schizophrenia. Clin. Schizophr. Relat. Psychoses 4, 56–73 (2010).
Vallone, D., Picetti, R. & Borrelli, E. Structure and function of dopamine receptors. Neurosci. Biobehav. Rev. 24, 125–132 (2000).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Kathiresan, S. et al. Common variants at 30 loci contribute to polygenic dyslipidemia. Nat. Genet. 41, 56–65 (2009).
UK Parkinson's Disease Consortium & Wellcome Trust Case Control Consortium 2. Dissection of the genetics of Parkinson's disease identifies an additional association 5′ of SNCA and multiple associated haplotypes at 17q21. Hum. Mol. Genet. 20, 345–353 (2011).
Höglinger, G.U. et al. Identification of common variants influencing risk of the tauopathy progressive supranuclear palsy. Nat. Genet. 43, 699–705 (2011).
Sullivan, P. Don't give up on GWAS. Mol. Psychiatry 17, 2–3 (2012).
International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Winkler, T.W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Cáceres, A. & González, J.R. Following the footprints of polymorphic inversions on SNP data: from detection to association tests. Nucleic Acids Res. 43, e53 (2015).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Sonnega, A. et al. Cohort profile: the Health and Retirement Study (HRS). Int. J. Epidemiol. 43, 576–585 (2014).
Willemsen, G. et al. The Adult Netherlands Twin Register: twenty-five years of survey and biological data collection. Twin Res. Hum. Genet. 16, 271–281 (2013).
van Beijsterveldt, C.E.M. et al. The Young Netherlands Twin Register (YNTR): longitudinal twin and family studies in over 70,000 children. Twin Res. Hum. Genet. 16, 252–267 (2013).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP–trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Penninx, B.W.J.H. et al. The Netherlands Study of Depression and Anxiety (NESDA): rationale, objectives and methods. Int. J. Methods Psychiatr. Res. 17, 121–140 (2008).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Fehrmann, R.S.N. et al. Gene expression analysis identifies global gene dosage sensitivity in cancer. Nat. Genet. 47, 115–125 (2015).
Acknowledgements
We are grateful to P.M. Visscher for advice, support, and feedback. We thank S. Cunningham and N. Galla for research assistance. This research was carried out under the auspices of the Social Science Genetic Association Consortium (SSGAC). The SSGAC seeks to facilitate studies that investigate the influence of genes on human behavior, well-being, and social–scientific outcomes using large GWAS meta-analyses. The SSGAC also provides opportunities for replication and promotes the collection of accurately measured, harmonized phenotypes across cohorts. The SSGAC operates as a working group within the CHARGE Consortium. This research has also been conducted using the UK Biobank Resource. The study was supported by funding from the US National Science Foundation (EAGER: 'Workshop for the Formation of a Social Science Genetic Association Consortium'), a supplementary grant from the National Institute of Health Office of Behavioral and Social Science Research, the Ragnar Söderberg Foundation (E9/11), the Swedish Research Council (421-2013-1061), the Jan Wallander and Tom Hedelius Foundation, an ERC Consolidator Grant (647648 EdGe), the Pershing Square Fund of the Foundations of Human Behavior, and the NIA/NIH through grants P01-AG005842, P01-AG005842-20S2, P30-AG012810, and T32-AG000186-23 to NBER and R01-AG042568-02 to the University of Southern California. A full list of acknowledgments is provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
M.B., D.J.B., D.C., J.-E.D.N., P.D.K., and R.F.K. designed and oversaw the study. A.O. and B.M.L.B. were responsible for quality control and meta-analyses. Bioinformatics analyses were carried out by J.P.B., T.E., M.A.F., J.R.G., J.J.L. S.F.W.M., M.G.N., and H.-J.W. Other follow-up analyses were conducted by M.A.F., J.P.B., P.T., A.O., B.M.L.B., and R.K.L. Especially major contributions to writing and editing were made by M.B., D.J.B., J.P.B., D.C.C., J.-E.D.N., P.D.K., A.J.O., and P.T. All authors contributed to and critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
University of Groningen, University Medical Center Groningen, Groningen, the Netherlands
Integrated supplementary information
Supplementary Figure 1 Illustrating the tradeoff between maximizing sample size and having more precise and uniform phenotype measures across cohorts.
This figure plots RMSE  versus n2 for various assumed values of R2 for β1 and of rg,LD, assuming that n1 = 200,000,
versus n2 for various assumed values of R2 for β1 and of rg,LD, assuming that n1 = 200,000,  and var(β2) = 7.24 × 10–7. A dotted line below the blue line indicates that including an additional independent cohort of size n2 with an imperfect phenotype measure with a cohort of size n1 tends to increase the accuracy and precision of the estimate of β1. See the Supplementary Note for additional details.
and var(β2) = 7.24 × 10–7. A dotted line below the blue line indicates that including an additional independent cohort of size n2 with an imperfect phenotype measure with a cohort of size n1 tends to increase the accuracy and precision of the estimate of β1. See the Supplementary Note for additional details.
Supplementary Figure 2 Local Manhattan plots of the association between SNPs on chromosome 8 and neuroticism in UKB.
Results are shown without controlling for the inversion-tagging principal component (PC) (a) and conditional on the PC (b). (c–e) Results without controls for the PC for individuals with inversion genotype 0 (c), inversion genotype 1 (d), and inversion genotype 2 (e). The gray background area indicates the inversion region. Note that the sample sizes differ in c–e.
Supplementary Figure 3 Proxy-phenotype analyses: test of SNPs associated with subjective well-being at P < 1 × 10–4 for association with depressive symptoms and neuroticism.
In both analyses, the second-stage sample is restricted to non-overlapping cohorts. For interpretational ease, for each SNP, we choose the reference allele for subjective well-being to be the effect-increasing allele. See Supplementary Tables 16 and 17 for detailed results and the Supplementary Note for additional details.
Supplementary Figure 4 Quantile–quantile plots for primary subjective well-being analysis, post hoc subjective well-being analysis using 1000 Genomes Project SNPs, life satisfaction analysis, and positive affect analysis.
(a) Primary subjective well-being analysis. (b) Post hoc subjective well-being analysis using 1000 Genomes Project SNPs. (c) Life satisfaction analysis. (d) Positive affect analysis. The estimated LD Score intercepts used to adjust the standard errors were 1.012, 1.008, 1.007, and 1.011, respectively. The gray-shaded areas in the quantile−quantile plots represent the 95% confidence intervals under the null hypothesis.
Supplementary Figure 5 Manhattan plots for life satisfaction analysis, positive affect analysis, and post hoc subjective well-being analysis using 1000 Genomes Project SNPs.
SNPs are plotted on the x axis according to their position on each chromosome against association with the phenotype on the y axis (shown as –log10 (P value)). The solid line indicates the threshold for genome-wide significance (P = 5 × 10–8), and the dashed line indicates the threshold for suggestive hits (P = 1 × 10–5). Each independent genome-wide significant association (lead SNP) is marked by ×.
Supplementary Figure 6 LocusZoom plots for reference SNPs rs13185787 and r6579956.
The plots are based on the summary statistics from our post hoc GWAS of subjective well-being using 1000 Genomes Project SNPs and on build hg19 (1000 Genomes Project November 2014 EUR). rs13185787 and r6579956 are the two genome-wide signals from our post hoc GWAS. rs13185787 and r6579956 are genome-wide signals from our baseline GWAS of subjective well-being using HapMap 2 SNPs that are in LD with rs13185787 (r2 = 0.95) and rs6579956 (r2 = 0.39), respectively.
Supplementary Figure 7 Quantile–quantile plots for depressive symptoms and neuroticism GWAS.
(a) Depressive symptoms. (b) Neuroticism. The estimated LD Score intercept used to adjust the standard errors was 1.008 for depressive symptoms. No adjustment was applied to the neuroticism results because the estimated intercept was below 1 (0.999). The gray-shaded areas in the quantile−quantile plots represent the 95% confidence intervals under the null hypothesis.
Supplementary Figure 8 LD Score regression plots based on the summary statistics from the GWAS of subjective well-being, depressive symptoms, and neuroticism.
Each point represents an LD Score quantile. The x and y coordinates of the point are the mean LD Score and the mean χ2 statistic of SNPs in that quantile, respectively. The fact that the intercepts are close to 1 and that the χ2 statistics increase linearly with the LD Scores for all three phenotypes suggests that the bulk of the inflation in the χ2 statistics for the three phenotypes is due to true polygenic signal and not to population stratification.
Supplementary Figure 9 Power of the sign test.
The estimated power of passing the GWAS/within-family sign test for subsets of approximately independent SNPs meeting various P-value thresholds. The x axis corresponds to the P-value threshold that defines the subset of SNPs used in the sign test. The y axis reports the expected power of the sign test given the posterior distribution of effect sizes corrected for winner’s curse. The non-monotonicity of this function is due to a tradeoff: as the P-value threshold increases, admitting a greater number of SNPs increases power, while admitting less significant SNPs decreases power.
Supplementary Figure 10 Histograms of the principal components tagging the inversions on chromosomes 8 and 17.
Superimposed over the histograms are the three-class normal mixtures fitted to the data with the purpose of assigning individuals to one of the three inversion genotypes for each inversion.
Supplementary Figure 11 Squared correlations between each SNP on chromosome 8 and the principal component tagging the inversion on chromosome 8.
(a) Squared correlations for all SNPs on chromosome 8. (b) Squared correlations for the SNPs from 6–14 Mb on chromosome 8. The red vertical lines are the borders chosen for the inversion in the UKB data. Note that SNPs around the borders of the inversion are missing because they do not pass quality control, likely as a result of the presence of the inversion.
Supplementary Figure 12 Local Manhattan plot for chromosome 17 (above horizontal line) and squared correlations between each SNP on chromosome 17 and the principal component tagging the inversion on chromosome 17 (below horizontal line).
Note the strong correspondence between the location of the genome-wide signals and the location of the inversion.
Supplementary Figure 13 Associations between inversion-tagging SNPs on chromosome 8 and neuroticism and subjective well-being.
(a) Associations with neuroticism are from the GPC (n = 63,661). (b) Associations with subjective well-being are from a meta-analysis of the cohorts that did not contribute to the neuroticism GWAS (n = 199,153). Using UKB data, we identified 126 SNPs whose pairwise R2 with the principal component tagging the inversion was above 0.75. Plotted are association statistics for the subsets of tagging SNPs that were available in the relevant lookup samples (111 SNPs for neuroticism and 53 SNPs for subjective well-being).
Supplementary Figure 14 Posterior credibility of GWAS results.
(a,c,e) The posterior probabilities that SNPs rs3756290 (a), rs2075677 (c), and rs4958581 (e) are non-null given the estimated effect of each SNP on subjective well-being. We use a mixture–Gaussian prior with point mass at 0, where the variance of the non-null SNPs is estimated by maximum likelihood using the GWAS data. The x axis corresponds to the fraction of SNPs that are assumed to be non-null by the prior, and the y axis corresponds to the probability that the corresponding SNP is non-null, given the prior and the estimated effect size. (b,d,f) The posterior mean effect sizes and 95% confidence intervals for SNPs rs3756290 (b), rs2075677 (d), and rs4958581 (f). The x axis corresponds to the fraction of SNPs that are assumed to be non-null by the prior, and the y axis corresponds to the posterior mean effect size (blue) and 95% confidence interval (red).
Supplementary Figure 15 Polygenic score prediction in HRS and NTR.
Predictive power of the polygenic score constructed from the subjective well-being GWAS results in two independent holdout cohorts (HRS and NTR). Predictive power is tested for subjective well-being, positive affect, life satisfaction, depressive symptoms, the Big Five personality traits (which include neuroticism), and height.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Note. (PDF 6039 kb)
Supplementary Tables 1–35
Supplementary Tables 1–35. (XLSX 532 kb)
Rights and permissions
About this article
Cite this article
Okbay, A., Baselmans, B., De Neve, JE. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat Genet 48, 624–633 (2016). https://doi.org/10.1038/ng.3552
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3552
This article is cited by
-
Genetic architecture distinguishes tinnitus from hearing loss
Nature Communications (2024)
-
Mendelian randomization studies of depression: evidence, opportunities, and challenges
Annals of General Psychiatry (2023)
-
Neuroticism mediates the association between childhood abuse and the well-being of community dwelling adult volunteers
BioPsychoSocial Medicine (2023)
-
A multivariate twin study of the genetic association between present moment attention and subjective wellbeing
Scientific Reports (2023)
-
A sex-specific genome-wide association study of depression phenotypes in UK Biobank
Molecular Psychiatry (2023)