Abstract
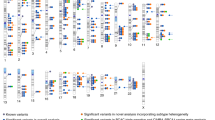
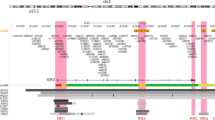
We conducted a three-stage genome-wide association study (GWAS) of breast cancer in 9,770 cases and 10,799 controls in the Cancer Genetic Markers of Susceptibility (CGEMS) initiative. In stage 1, we genotyped 528,173 SNPs in 1,145 cases of invasive breast cancer and 1,142 controls. In stage 2, we analyzed 24,909 top SNPs in 4,547 cases and 4,434 controls. In stage 3, we investigated 21 loci in 4,078 cases and 5,223 controls. Two new loci achieved genome-wide significance. A pericentromeric SNP on chromosome 1p11.2 (rs11249433; P = 6.74 × 10−10 adjusted genotype test, 2 degrees of freedom) resides in a large linkage disequilibrium block neighboring NOTCH2 and FCGR1B; this signal was stronger for estrogen-receptor–positive tumors. A second SNP on chromosome 14q24.1 (rs999737; P = 1.74 × 10−7) localizes to RAD51L1, a gene in the homologous recombination DNA repair pathway. We also confirmed associations with loci on chromosomes 2q35, 5p12, 5q11.2, 8q24, 10q26 and 16q12.1.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Colditz, G.A., Baer, H.J. & Tamimi, R.M. in Cancer Epidemiology and Prevention (eds. Schottenfeld, D. & Fraumeni, J.F.) Breast Cancer, 995–1012 (Oxford University Press, New York, 2006).
Miki, Y. et al. A strong candidate for the breast and ovarian-cancer susceptibility gene BRCA1. Science 266, 66–71 (1994).
Wooster, R. et al. Identification of the breast cancer susceptibility gene BRCA2. Nature 378, 789–792 (1995).
Rahman, N. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat. Genet. 39, 165–167 (2007).
Meijers-Heijboer, H. et al. Low-penetrance susceptibility to breast cancer due to CHEK2*1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 31, 55–59 (2002).
Erkko, H. et al. A recurrent mutation in PALB2 in Finnish cancer families. Nature 446, 316–319 (2007).
Renwick, A. et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat. Genet. 38, 873–875 (2006).
Seal, S. et al. Truncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility alleles. Nat. Genet. 38, 1239–1241 (2006).
Cox, A. et al. A common coding variant in CASP8 is associated with breast cancer risk. Nat. Genet. 39, 352–358 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Hunter, D.J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Stacey, S.N. et al. Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancer. Nat. Genet. 39, 865–869 (2007).
Stacey, S.N. et al. Common variants on chromosome 5p12 confer susceptibility to estrogen receptor-positive breast cancer. Nat. Genet. 40, 703–706 (2008).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Yu, K. et al. Population substructure and control selection in genome-wide association studies. PLoS ONE 3, e2551 (2008).
Falush, D., Stephens, M. & Pritchard, J.K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Skol, A.D., Scott, L.J., Abecasis, G.R. & Boehnke, M. Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat. Genet. 38, 209–213 (2006).
Gold, B. et al. Genome-wide association study provides evidence for a breast cancer risk locus at 6q22.33. Proc. Natl. Acad. Sci. USA 105, 4340–4345 (2008).
Staiger, H. et al. Novel meta-analysis-derived type 2 diabetes risk loci do not determine prediabetic phenotypes. PLoS ONE 3, e3019 (2008).
Xue, F. & Michels, K.B. Diabetes, metabolic syndrome, and breast cancer: a review of the current evidence. Am. J. Clin. Nutr. 86, s823–s835 (2007).
Li, X. & Heyer, W.D. Homologous recombination in DNA repair and DNA damage tolerance. Cell Res. 18, 99–113 (2008).
Antoniou, A.C. et al. RAD51 135G → C modifies breast cancer risk among BRCA2 mutation carriers: results from a combined analysis of 19 studies. Am. J. Hum. Genet. 81, 1186–1200 (2007).
Shlien, A. et al. Excessive genomic DNA copy number variation in the Li-Fraumeni cancer predisposition syndrome. Proc. Natl. Acad. Sci. USA 105, 11264–11269 (2008).
Garcia-Closas, M. et al. Heterogeneity of breast cancer associations with five susceptibility loci by clinical and pathological characteristics. PLoS Genet. 4, e1000054 (2008).
Pharoah, P.D., Antoniou, A.C., Easton, D.F. & Ponder, B.A. Polygenes, risk prediction, and targeted prevention of breast cancer. N. Engl. J. Med. 358, 2796–2803 (2008).
Pepe, M.S. & Janes, H.E. Gauging the performance of SNPs, biomarkers, and clinical factors for predicting risk of breast cancer. J. Natl. Cancer Inst. 100, 978–979 (2008).
Acknowledgements
The Nurses' Health Studies are supported by US National Institutes of Health grants CA65725, CA87969, CA49449, CA67262, CA50385 and 5UO1CA098233. We thank B. Egan, L. Egan, H. Judge Ellis, H. Ranu and P. Soule for assistance, and the participants in the Nurses' Health Studies. The WHI program is supported by contracts from the National Heart, Lung and Blood Institute, NIH. We thank the WHI investigators and staff for their dedication, and the study participants for making the program possible. A full listing of WHI investigators can be found at http://www.whi.org. The ACS study is supported by UO1 CA098710. We thank C. Lichtman for data management and the participants on the CPS-II. The US Radiologic Technologists Study (USRT) is supported by the Intramural Research Program of the Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, DHHS. The PLCO study is supported by the Intramural Research Program of the Division of Cancer Epidemiology and Genetics and contracts from the Division of Cancer Prevention, National Cancer Institute, NIH, DHHS. We thank P. Prorok, Division of Cancer Prevention, National Cancer Institute, the Screening Center investigators and staff of the Prostate, Lung, Colorectal, and Ovarian Cancer Screening Trial (PLCO), and T. Sheehy and staff at SAIC-Frederick. We acknowledge the study participants for their contributions to making this study possible. We thank the radiologic technologists who participated in the study; J. Reid of the American Registry of Radiologic Technologists for continued support of the study; D. Kampa and A. Iwan of the University of Minnesota for study coordination and data collection; B. Kopp and staff at SAIC-Frederick for biospecimen processing; and L. Bowen of Information Management Systems for data management.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Tables 1–4 and Supplementary Figures 1 and 2 (PDF 4197 kb)
Rights and permissions
About this article
Cite this article
Thomas, G., Jacobs, K., Kraft, P. et al. A multistage genome-wide association study in breast cancer identifies two new risk alleles at 1p11.2 and 14q24.1 (RAD51L1). Nat Genet 41, 579–584 (2009). https://doi.org/10.1038/ng.353
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.353
This article is cited by
-
Four differentially expressed genes can predict prognosis and microenvironment immune infiltration in lung cancer: a study based on data from the GEO
BMC Cancer (2022)
-
2q35-rs13387042 variant and the risk of breast cancer: a case–control study
Molecular Biology Reports (2022)
-
A truncating variant of RAD51B associated with primary ovarian insufficiency provides insights into its meiotic and somatic functions
Cell Death & Differentiation (2022)
-
Potential biomarkers and lncRNA-mRNA regulatory networks in invasive growth hormone-secreting pituitary adenomas
Journal of Endocrinological Investigation (2021)
-
Integrative genomic analysis implicates ERCC6 and its interaction with ERCC8 in susceptibility to breast cancer
Scientific Reports (2020)