Abstract
Bats are the only mammals capable of powered flight, but little is known about the genetic determinants that shape their wings. Here we generated a genome for Miniopterus natalensis and performed RNA-seq and ChIP-seq (H3K27ac and H3K27me3) analyses on its developing forelimb and hindlimb autopods at sequential embryonic stages to decipher the molecular events that underlie bat wing development. Over 7,000 genes and several long noncoding RNAs, including Tbx5-as1 and Hottip, were differentially expressed between forelimb and hindlimb, and across different stages. ChIP-seq analysis identified thousands of regions that are differentially modified in forelimb and hindlimb. Comparative genomics found 2,796 bat-accelerated regions within H3K27ac peaks, several of which cluster near limb-associated genes. Pathway analyses highlighted multiple ribosomal proteins and known limb patterning signaling pathways as differentially regulated and implicated increased forelimb mesenchymal condensation in differential growth. In combination, our work outlines multiple genetic components that likely contribute to bat wing formation, providing insights into this morphological innovation.
Similar content being viewed by others
Main
The order Chiroptera, commonly known as bats, is the only group of mammals to have evolved the capability of flight. They are estimated to have diverged from their arboreal ancestors ∼51 million years ago1. Their adaptions for flight include substantial specialization of the forelimb, characterized by the notable extension of digits II–V, a decrease in wing bone mineralization along the proximal-distal axis, and the retention and expansion of interdigit webbing, which is controlled by a novel complex of muscles2,3. Bat hindlimbs are comparatively short, with free, symmetrical digits, providing an informative contrast that can be used to highlight the genetic processes involved in bat wing formation. Previous studies that examined gene expression in developing bat forelimbs and hindlimbs reported differential expression of several genes, including Tbx3, Brinp3, Meis2, the 5′ HoxD genes and components of the Shh-Fgf signaling loop, suggesting that multiple genes and processes are involved in generating these morphological innovations4,5,6,7,8. Gene regulatory elements are thought to be important drivers of these changes: for example, replacement of the mouse Prx1 limb enhancer with the equivalent bat sequence resulted in elongated forelimbs9. However, an integrated understanding of how changes in regulatory elements, various genes and signaling pathways combine to collectively shape the bat wing remains largely elusive.
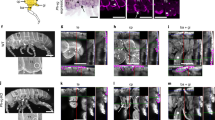
To characterize the genetic differences that underlie divergence in bat forelimb and hindlimb development, we used a comprehensive, genome-wide strategy. We generated a de novo whole-genome assembly for the vesper bat, M. natalensis, for which a well-characterized stage-by-stage morphological comparison between developing bat and mouse limbs is available10. In this species, the developing forelimb noticeably diverges from the hindlimb from developmental stages CS15 and CS16, with clear morphological differences seen at a subsequent stage, CS17 (ref. 10). This developmental window is equivalent to embryonic day (E) 12.0 to E13.5 in mouse4,10. M. natalensis embryos were obtained and transcriptomic (RNA-seq) data and ChIP-seq data for both an active (acetylation of histone H3 at lysine 27, H3K27ac; refs. 11,12) and a repressive (trimethylation of histone H3 at lysine 27, H3K27me3; ref. 13) mark were generated for these three developmental stages (Fig. 1).
Results
The M. natalensis genome
High-coverage genomes for three bat species (Pteropus alecto14, Myotis davidii14 and Myotis brandtii15) and low-coverage genomes for two bat species (Myotis lucifugus and Pteropus vampyrus16) have been published. However, the evolutionary distance of these species from M. natalensis (43 million years since the last common ancestor) precludes the use of their genomes in RNA-seq and ChIP-seq data analyses. We thus generated a draft genome from an adult M. natalensis male at 77× coverage, named Mnat.v1. The quality of the Mnat.v1 genome is comparable to that of the high-coverage bat genomes (Supplementary Table 1). It has an estimated heterozygosity level of 0.13%, with repetitive regions making up 33% of the genome. We annotated 24,239 genes (including protein-coding genes and long noncoding RNAs (lncRNAs)) in Mnat.v1. Of the highly conserved genes used by the Core Eukaryotic Genes Mapping Approach (CEGMA)17, 92.7% were found in their entirety, with an additional 3.3% partially detected, further confirming Mnat.v1 to be a reliable substrate for subsequent genomic analyses.
Differentially expressed limb transcripts
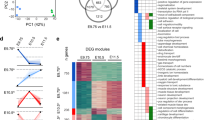
To identify gene expression differences that could be involved in the morphological divergence in bat limb development, we examined the transcriptomes of whole-autopod tissue from forelimbs and hindlimbs at three sequential developmental stages (CS15, CS16 and CS17). Principal-component analysis (PCA) showed an expected segregation pattern, with principal component 1 (PC1) reflecting the developmental stage and principal component 2 (PC2) reflecting the tissue type (forelimb or hindlimb; Fig. 2a). We found 2,952 genes differentially expressed in forelimb and hindlimb and 5,164 genes differentially expressed in comparisons of any two sequential stages (adjusted P value ≤ 0.01; Online Methods). Pairwise tests for differential expression directly comparing forelimb and hindlimb at each stage (for example, CS15 forelimb versus CS15 hindlimb) contributed an additional 1,596 genes. Combined, these analyses identified 7,172 differentially expressed genes (adjusted P value ≤ 0.01; Fig. 2b and Supplementary Table 2).
(a) PCA using the 3,000 genes with the highest variance in expression. PC1 is stage dependent, and PC2 is tissue dependent (forelimb or hindlimb); these components explain 57.1% and 13.3% of the variance, respectively. FL, forelimb; HL, hindlimb. (b) Gene-wise hierarchical clustering heat map of all 7,172 differentially expressed genes (adjusted P value ≤ 0.01) showing segregation into five groups. The z-score scale represents mean-subtracted regularized log-transformed read counts. Cluster 1 (n = 64) includes genes with increased expression throughout the stages. Cluster 11 (n = 465) includes genes upregulated in hindlimb. Cluster 30 (n = 718) includes genes upregulated in forelimb. Each box in the box plots is the interquartile range (IQR), the line is the median and the whiskers show the furthest data points from the median within 1.5 times the IQR. Enriched GO terms are shown to the right. (c) Heat map of the genes from the DNA binding (GO:0003677) and regulation of transcription, DNA dependent (GO:0006355) GO terms that displayed the most significant differences (adjusted P value ≤ 0.01) and greatest fold change (fold change ≥2) in expression between forelimb and hindlimb. The z-score scale represents a sample of the mean-subtracted average of the regularized log-transformed read counts in each sample. Mllt3 and Lhx8 are highlighted by red and purple asterisks, respectively. (d) In situ hybridization for Mllt3 and Lhx8 in stage-matched forelimbs and hindlimbs from bat and mouse. Bat Mllt3 expression shows a shift toward the distal autopod in the future location of digits III–V, which elongate in bats. Bat expression of Lhx8 is strongest in the most proximal region of the autopod, especially along the anterior and posterior edges of the limb. Scale bars, 0.5 mm.
Differentially expressed genes were grouped by their expression profiles across the samples into 38 manually defined clusters using hierarchical clustering (Supplementary Fig. 1). These clusters were functionally annotated, with several Gene Ontology (GO) terms correlated with differential expression (Fig. 2b and Supplementary Table 3). Grouping the genes displaying the most significant differential expression on the basis of biological functions of interest (for example, DNA binding and transcriptional regulation, limb morphogenesis, bone morphogenesis, apoptotic process and others) identified both genes with known roles in limb development and genes with potentially new functions in bat wing development (Supplementary Fig. 2). For example, the genes differentially expressed in forelimb and hindlimb involved in DNA binding and transcriptional regulation included Hoxd10, Hoxd11, Meis2, Pitx1, Tbx4 and Tbx5, comprising all genes that were previously shown to be differentially expressed in bats5,6,7 along with several genes showing higher expression in forelimb that have not yet been characterized (Fig. 2c). We also observed hindlimb-specific increased expression for several genes, notably Msx1 and Msx2, both of which are key genes involved in apoptotic activity during interdigit tissue regression18.
We characterized a limited number of differentially expressed genes of interest in both mouse and bat embryos using whole-mount in situ hybridization (Supplementary Fig. 3). Among these, Mllt3 was chosen for its strong forelimb expression at CS15 and CS16 (Fig. 2c) and was found to be uniquely expressed in bat forelimb in a region restricted to the distal edge where digits III–V are slated to develop (Fig. 2d). Mllt3 is thought to be a Hox gene regulator, with Mllt3-null mouse mutants exhibiting axial defects19; however, no gross skeletal limb abnormalities were observed in homozygous knockout mice (Online Methods and Supplementary Fig. 4). Lhx8, a known regulator of neuronal development20, had higher expression in CS16 and CS17 forelimbs (Fig. 2c). Whole-mount in situ hybridization analysis showed localized Lhx8 expression in the posterior portion of the wrist region, specifically in the junction between the base of digit V and the plagiopatagium, whereas no expression was detected in mouse limbs (Fig. 2d). Together, these experiments support our RNA-seq analyses and highlight genes that have previously uncharacterized roles in limb development.
Bat-specific lncRNAs
lncRNAs have been shown to be important developmental regulators in several tissues, including the limb21. To find potential lncRNAs associated with bat limb development, we annotated transcripts that did not show similarity to known protein-coding genes, identifying 227 potential lncRNAs (Supplementary Table 4). Among these, 188 exhibited some sequence conservation across mammals, 12 of which were similar to characterized lncRNAs in lncRNAdb v2.0 (ref. 22). Five putative lncRNAs were identified as being conserved only in bats, and 34 were only present in M. natalensis. Within this data set, eight known lncRNAs showed differential expression between forelimb and hindlimb, including Hottip and an uncharacterized lncRNA, Tbx5-as1 (Fig. 3). Hottip is thought to be required for activation of 5′ HoxA genes, which are important regulators of autopod patterning during limb development21. Both Hottip and Hoxa13 showed elevated hindlimb expression in all three stages examined (Fig. 3a). A comparison of their expression patterns showed both to be more strongly expressed in interdigit tissue of the bat hindlimb. Although Hottip expression was concentrated in distal interdigit tissue, Hoxa13 expression was more apparent in digit tips (Fig. 3b,c). The bat Tbx5-as1 transcript maps close to Tbx5 (Fig. 4a), in the antisense orientation and is similar to human Tbx5 antisense RNA1-as1 transcript (GenBank, NR_038440). Tbx5-as1 was the most differentially expressed lncRNA, with elevated expression in the forelimb relative to the hindlimb across all stages (Fig. 3a). Although its role is unknown, its associated gene, Tbx5, is required for forelimb bud initiation, with inactivation in mice abolishing forelimb skeletal formation23,24. In support of coupled activity for these transcripts, the expression patterns of Tbx5 and Tbx5-as1 were similar: both were restricted to the base of digits I–V during late CS16 (CS16L) and CS17, with clear expression in proximal interdigit tissue (Fig. 3d,e).
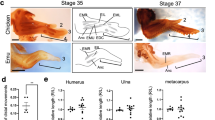
(a) RNA-seq fragments per kilobase of exon per million mapped reads (FPKM) results for Tbx5, Tbx5-as1, Hoxa13 and Hottip. Asterisks denote significant forelimb versus hindlimb expression differences by stage (adjusted P value ≤ 0.01). (b) In situ hybridization for Hoxa13 at CS16L and CS17, showing lower expression in the forelimb than in the hindlimb but retained expression in digit tips in forelimb. (c) In situ hybridization for Hottip at CS15 and CS16L, showing expression in the interdigit webbing in both forelimb and hindlimb but lower levels in forelimb. (d,e) In situ hybridization for Tbx5 at CS16L and CS17L (d) and Tbx5-as1 at CS16L and CS17 (e) showing both transcripts to be restricted to the base of digits I–V, with robust expression in proximal interdigit tissue at CS17. Scale bars, 0.5 mm.
(a,b) RNA-seq and ChIP-seq tracks for Tbx5 (a) and Pitx1 (b). RNA-seq and ChIP-seq tracks are represented as forelimb coverage minus hindlimb coverage in 100-bp intervals for each stage. For RNA-seq data, intervals with forelimb minus hindlimb expression >0 are shown in dark blue and those with values <0 are shown in light blue. Likewise, for ChIP-seq data, intervals with forelimb minus hindlimb enrichment values >0 are colored in dark green (H3K27ac) and dark red (H3K27me3) and those with values <0 are colored in light green (H3K27ac) and light red (H3K27me3). The scale has been normalized to range from 0 to 1, where 0 is essentially background noise and 1 corresponds to the top 0.1% of signal. (c) Heat map of H3K27ac (green) and H3K27me3 (red) enrichment scores, including a dendrogram of region-wise hierarchical clustering. The heat map shows all regions with differential enrichment between forelimb and hindlimb in both chromatin marks in at least two stages per mark (2,475 such regions at adjusted P value ≤ 0.05). The z score represents the mean-subtracted log2-transformed value of the signal to noise–normalized enrichment score + 1. The hierarchical clusters of the regions were segregated into 17 separate clusters; 2 such clusters are shown as examples, cluster 11 (n = 258 peaks) with higher hindlimb H3K27ac and forelimb H3K27me3 levels and cluster 9 (n = 108 peaks) with higher forelimb H3K27ac and hindlimb H3K27me3 levels. RNA-seq expression levels of the ChIP-seq peaks neighboring genes are plotted next to the histone mark data. Enrichment score distribution is shown as box plots for each cluster, and the enrichment for GO categories of the nearest gene for each region is displayed to the right.
ChIP-seq highlights forelimb regulatory regions
Changes in gene regulatory elements have been shown to be important drivers of morphological adaptations25, including the bat wing9. To identify regulatory elements that could be involved in controlling gene expression in developing bat limbs, we performed low-cell-number ChIP-seq using antibodies for both H3K27ac (active regions14,15) and H3K27me3 (repressed regions13) on autopods from CS15, CS16 and CS17 forelimbs and hindlimbs, identifying numerous putative regulatory regions (Supplementary Table 5). Using the Genomic Regions Enrichment of Annotations Tool (GREAT)26, after converting peaks to mouse genomic coordinates, we found significant enrichment for the H3K27ac peaks in several limb development–associated categories among GO morphological processes, mouse phenotypes and Mouse Genome Informatics (MGI) expression data (Supplementary Fig. 5). To further validate our ChIP-seq results, we also examined genes known to be specifically expressed in the forelimb (Tbx5) or hindlimb (Pitx1) and observed correspondence with H3K27ac and H3K27me3 peak presence and RNA expression (Fig. 4a,b).
We next set out to analyze differences between forelimb and hindlimb active and repressed ChIP-seq peaks. Differential enrichment analysis was carried out on H3K27ac and H3K27me3 peaks separately, identifying 14,553 and 19,352 differentially enriched regions, respectively (pairwise false discovery rate (FDR) ≤ 0.05; Online Methods and Supplementary Table 6). Of these regions, 2,475 were differentially enriched between forelimb and hindlimb for both H3K27ac and H3K27me3 signal. These regions were analyzed using hierarchical clustering based on H3K27ac and H3K27me3 enrichment (Fig. 4c), from which 17 manually defined clusters were identified with distinct H3K27ac and H3K27me3 enrichment patterns (Supplementary Fig. 6). GO term analysis of the nearest gene for each region showed enrichment for terms associated with limb development. For example, cluster 9 showed higher H3K27ac levels in forelimb and H3K27me3 levels in hindlimb, whereas cluster 11 showed the opposite pattern. The regulatory marks for both clusters had a general correspondence with RNA-seq expression levels for the neighboring genes and included fitting GO biological term enrichment for developmental processes (Fig. 4c and Supplementary Table 7).
Bat accelerated regions
To identify genomic changes that might be associated with the innovation of the bat wing, we used a comparative genomics approach27 that leveraged the growing number of bat genomes14,15,16. Whole-genome alignments were generated using the repeat-masked genomes of 18 other species, including 6 bats, 9 non-bat mammals and 3 non-mammal vertebrates. We next used phyloP28 to identify accelerated sequences, sequences that are conserved in vertebrates but changed significantly in the common ancestor of the bat lineage and were marked by H3K27ac in all ChIP-seq experiments. This analysis identified 2,796 bat accelerated regions (BARs; FDR ≤ 0.05) with an average size of 240 bp (Supplementary Table 8). Genomic regions over-represented for BARs were identified by comparison to vertebrate conserved regions overlapping H3K27ac peaks. Genes contained in these regions were subjected to functional annotation clustering, showing enrichment for categories relating to transcription factors, chromatin conformation and DNA binding (FDR ≤ 0.05; Supplementary Table 8). The region most highly enriched for BARs included the genes Lrrn1 (leucine-rich repeat neuronal 1) and Crbn (cereblon) (Supplementary Fig. 7a). Lrrn1 is expressed at significantly higher levels in bat hindlimb than forelimb (adjusted P value = 6.73 × 10−11; Supplementary Table 2) and was also shown to be expressed in developing mouse limb29. It was also shown to be important for midbrain-hindbrain boundary formation regulated by Fgf8 (ref. 30). Crbn is a known thalidomide target, thought to be important in limb outgrowth through its regulation of Fgfs31, but did not show significant expression differences between forelimb and hindlimb (Supplementary Table 2). Another BAR-dense region was around Fgf2 and Spry1 (Supplementary Fig. 7b). Fgf2 is known to have regenerative capabilities in the limb32 and was both the most highly expressed and had the most significant fold change in expression between forelimb and hindlimb across all stages among the Fgf genes in our study (Supplementary Fig. 7c). Spry1 was shown to be involved in limb muscle and tendon development33 but did not have significant expression differences between forelimb and hindlimb (Supplementary Fig. 7c). Combined, our ChIP-seq and BAR analyses highlight specific candidate sequences and genomic regions that might have had a role in development of the bat wing.
Wing developmental pathways
We next used Ingenuity Pathway Analysis (IPA) to identify signaling pathways that were differentially activated across the data set and could contribute to the differences in patterning between bat forelimb and hindlimb autopods. Interestingly, the top pathway in our analysis, showing strong hindlimb activation, was the elongation initiation factor 2 (EIF2) signaling pathway (Fig. 5a and Supplementary Table 9), which has an important role in regulating the initiation of protein synthesis. A closer inspection showed that 41 ribosomal protein genes, which are coordinately downregulated in bat forelimb at CS15 and CS16 (Fig. 5b), were largely responsible for this score. Ribosomal protein expression has been shown to be highly heterogeneous across tissues during embryonic development, including in the limb34. Mutations in the loci encoding the ribosomal proteins RPL11, RPL35A, RPS7, RPS10 and RPS19 are known to lead to limb malformations in individuals with Diamond-Blackfan anemia35. The Rpl38 gene, whose encoded protein facilitates the translation of several HoxA genes by an IRES-dependent mechanism36 and that is mutated in tail short mice, which have skeletal patterning defects34, was downregulated in CS15 and CS16 bat forelimbs (Fig. 5b). Rictor, a negative upstream regulator of these ribosomal proteins, showed higher expression in forelimb (Supplementary Table 2). Rictor is a subunit of the mTORC2 complex, having a role in actin cytoskeleton organization, with conditional deletions in mice resulting in narrower and shorter limb bones37. Combined, our pathway analyses suggest that ribosomal proteins and their regulators could have an important role in bat wing development through the translational control of specific subsets of mRNA transcripts.
(a) The top ten canonical IPA-annotated pathways ranked by absolute IPA activation z scores for at least one developmental stage (**P < 0.001). The IPA z-score scale predicts the activation (or inhibition) state of the respective signaling pathways in bat forelimbs relative to hindlimbs. (b) Heat map showing the RNA-seq expression patterns of gene members of the EIF2 signaling pathway. The z-score scale represents a sample of the mean-subtracted average of the regularized log-transformed read counts in each sample. Genes that, when mutated, cause Diamond-Blackfan anemia are highlighted in bold, and Rpl38, which facilitates the translation of several HoxA genes, is indicated by an asterisk. (c) Differentially expressed genes from the Fgf, Wnt/β-catenin, Wnt-PCP and Bmp signaling pathways. Genes expressed more highly in forelimbs are on a white background, whereas genes expressed more highly in hindlimbs are on a gray background; activators are highlighted in green, and repressors are underlined and highlighted in red. Asterisks indicate genes that were manually assigned to the IPA curated pathways. (d) Differentially expressed genes known to be important regulators or markers of different stages of bone development, including mesenchymal condensation, chondrocyte differentiation, proliferation, maturation and hypertrophy. Genes that are established markers of a particular cell type are indicated in dark blue (mesenchymal condensations), light blue (proliferating chondrocytes) and gray (terminal chondrocytes), whereas positive regulators are depicted in green and repressors are depicted in red. The stages of bone development are aligned to embryonic bat limb developmental stages10.
Several pathways known to have an important role in limb and bone development, including Fgf, Wnt and Bmp signaling, were among the top ten IPA canonical pathways coordinately activated or repressed in bat forelimb as compared to hindlimb (Fig. 5a). Fgf proteins are known to mediate limb patterning by signaling the initial outgrowth of the limb bud from the apical ectodermal ridge38. This pathway showed consistent activation in CS15–CS17 forelimbs (Fig. 5c), with expression of Fgf2, Fgf7, Fgf19 and Hgf in forelimbs at CS16 and CS17 (Fig. 5c). We also observed higher hindlimb expression for several Fgf antagonists, including Spry2, Spry4 and Fgfrl1. Wnt ligands are secreted from the limb bud ectoderm and block cartilage formation in the periphery of the limb bud via the β-catenin pathway39. We observed overall suppression of the canonical Wnt/β-catenin pathway in forelimb versus hindlimb, with higher levels of several canonical Wnt pathway antagonists in the forelimb and canonical Wnt receptors in the hindlimb (Fig. 5c), including Lef1, which showed strong CS15 hindlimb expression in whole-mount in situ hybridization (Supplementary Fig. 3). The Wnt–planar cell polarity (PCP) pathway has an important role in elongation of the limb along the proximal-distal axis40 and was activated in bat forelimbs at all stages (Fig. 5c). This upregulation included the PCP pathway ligand Wnt11, which has been shown to antagonize the Wnt/β-catenin pathway41.
β-catenin signaling is known to suppress condensation of mesenchymal cells in endochondral bone development39. To test our prediction that β-catenin signaling is diminished and leads to larger fields of condensing mesenchymal cells (Fig. 5d), we stained sagittal sections of bat forelimb and hindlimb autopods using peanut agglutinin (PNA), a galactose-specific lectin that binds to cell surface markers on condensing precartilage mesenchymal cells42. Hematoxylin and eosin staining of matched sections demarcated the progression from condensation of mesenchymal cells (CS15) to differentiation of chondrocytes (CS16) and progression to mature chondrocytes (CS17) in both forelimb and hindlimb autopods (Supplementary Fig. 8). At CS15, PNA staining was more intense, centered on the emerging digit IV, in sections of forelimb as compared to hindlimb autopods (Fig. 6). By CS16, all five digits were clearly visible in both bat forelimb and hindlimb sections (Fig. 6 and Supplementary Fig. 8). Whereas PNA staining was diminished as chondrocytes differentiated and matured, forelimb digits showed more intense, continued recruitment of condensing mesenchymal cells in the distal domain of digits II–V at both CS16 and CS17 (Fig. 6 and Supplementary Fig. 8). These data suggest that the timing and size of the initial digit condensations and subsequent recruitment of mesenchymal condensations are different in bat forelimb and hindlimb autopods from CS15 onward and that the foundation for the rapid elongation of forelimb digits could be established earlier than CS20, as previously proposed43.
(a–l) PNA (green) and Hoechst (blue) staining of sagittal sections of bat CS15, CS16 and CS17 forelimb (a–c) and hindlimb (g–i) autopods. Higher magnification views are shown of the boxed regions, which correspond to the area with the strongest PNA staining at CS15 (d,j), digits III and IV at CS16 (e,k) and digit IV at CS17 (f,l). Ellipses indicate regions of maturing chondrocytes, determined by comparison to hematoxylin and eosin staining of adjacent sections (Supplementary Fig. 8). Scale bars: 500 μm in a–c and g–i, 100 μm in d–f and j–l.
In the limb, Bmp signaling regulates both bone formation and interdigit tissue regression44. We observed two distinct phases of Bmp signaling in our data sets (Fig. 5c). During digit initiation and specification (CS15), we observed high levels of the Bmp inhibitors Gremlin and Bmp3 in the hindlimb (which indicates a slight developmental lag at this stage), whereas the transcripts for Bmp receptors (Bmpr1a, Bmpr1b, Bmpr2 and Acvr1) and ligands (Bmp5 and Gdf5) were more abundant in the forelimb. Mutations in Bmp5 were shown to decrease mouse limb width45, and overexpression of GDF5 in chicken increases skeletal length46. The pattern of Bmp signaling started to switch at CS16, with CS17 forelimb showing higher levels of Bmp3 and Gremlin. Expression of these Bmp antagonists in the forelimb is consistent with the observed decrease in Msx1 and Msx2 expression. A similar suppression of the Bmp signaling pathway has been shown to have an important role in retention of interdigit webbing in duck47. Ranking genes from our differentially expressed signaling pathways for consistency across the RNA-seq and ChIP-seq data sets (Supplementary Table 9) found Msx2 (Bmp signaling) and Fzd10 (Wnt/β-catenin pathway) to be positively correlated for RNA-seq, H3K27ac and H3K27me3 signal. These genomic regions contain 8 and 12 BARs, respectively (within 500 kb of the transcription start site), suggesting that they could be important determinants of bat wing development.
Discussion
To identify genetic components that might contribute to bat wing development, we carried out whole-genome sequencing combined with RNA-seq and ChIP-seq for H3K27ac and H3K27me3 on developing bat forelimbs and hindlimbs at three key developmental time points. Overall, we found that multiple genetic components are likely to contribute to development of the bat wing. These include numerous gene expression changes, both in known limb developmental regulators and newly characterized ones, such as Mllt3 and Lhx8. lncRNAs could also have a strong influence on wing development, with observed forelimb-hindlimb expression differences for Hottip and Tbx5-as1, an uncharacterized lncRNA. Combined pathway analysis highlighted numerous signaling pathways that seem to be differentially activated. These include ribosomal proteins, whose alteration has been shown to result in limb malformations35. Suppression of the Wnt/β-catenin pathway in the forelimb is consistent with condensation of larger fields of digit mesenchymal cells in the developing bat wing. In contrast, Wnt-PCP signaling, which maintains the polarity of proliferating chondrocytes in the growth plate, was more active in forelimb (Fig. 6d) and could potentially set the foundation for extended digit growth. Interestingly, the Bmp signaling pathway showed two distinct phases, with the inhibitors Gremlin and Bmp3 expressed at high levels early in the hindlimb and at later stages in the forelimb, with different tissue and temporal identity for Bmp activators, fitting with the diverse roles of this pathway in chondrogenesis, osteogenesis and apoptosis. Combined, the differential activation of these pathways is consistent with changes in expression of key genes in long bone development, including enhanced expression of chondrogenic markers (for example, Sox6, Aggrecan and Mmp9) across CS15–CS17 (Fig. 6d). These expression changes could be driven by gene regulatory elements, with potential candidate sequences residing in our ChIP-seq data sets.
Our study obtained unique genomic data from wild-caught non-model organisms. Although restricted sample sizes, biological variation and gross tissue sampling may have reduced the scope of the experiments and the power of some of the analyses, we were able to generate robust genomic, transcriptomic and epigenomic data sets, which identified potential regulators of the processes involved in bat limb development. As bats are not currently amenable to transgenic experimentation, future functional characterization of the genes, lncRNAs and regulatory elements identified here could be performed in the mouse, with the potential to further understanding of their functional importance in the limb. Combined, our results uncover, on a genomic level, the molecular components and pathways that may have a key role in formation of the bat wing and provide a foundation for future studies examining such unique morphological innovations.
Methods
Genome assembly.
DNA was extracted from the leg muscle tissue of a single male M. natalensis using phenol-chloroform. The 4-μg protocol of the Nextera Mate-Pair Sample Preparation kit (Illumina) was used to generate libraries with insert sizes of 2 kb, 5–6 kb and 8–10-kb. For the libraries with insert sizes of 5–6 kb and 8–10 kb, multiple reactions were pooled (four and seven, respectively) before size selection. The smaller-insert libraries were generated with the TruSeq DNA LT Sample Preparation kit (Illumina), following the manufacturer's instructions. All libraries were sequenced on the Illumina HiSeq 2500 platform. The 175-bp and 300-bp paired reads were trimmed on either side to a minimum quality of 17 using Trimmomatic48. Trimmed reads were then used to calculate the 27-bp k-mer frequency using KmerFreq_HA in SOAPdenovo49. The 175-bp paired reads were then merged using their theoretical 25-bp overlaps and FLASH50. The remaining read pairs were trimmed to a minimum quality of 17 on the 3′ end before all reads were error corrected using Corrector_HA in SOAPdenovo. k-mers in a read with a frequency of 3 or lower were corrected to a more common k-mer. These changes were limited to two instances for the non-overlapping paired-end reads and four instances for the 175-bp reads. After these corrections, further erroneous k-mers were removed, to obtain a minimum read length of 60 bp. Duplicate reads were removed using FastUniq51. In combination, the reads totaled over 77× coverage, of which 17.5× was composed of long-insert mate pairs. Processed reads were assembled using SOAPdenovo49 and a k-mer size of 49. Pairs with one read mapping to a contig and one read mapping to a gap in a scaffold were used to fill in gaps with GapCloser (SOAPdenovo; submitted as whole-genome sequencing data under BioProject PRJNA283550). Heterozygosity was estimated using the Burrows-Wheeler Aligner (BWA)52 and SAMtools53. The coherency of the genomic sequence was tested with CEGMA17, using mammalian optimization.
Genome annotation.
The M. natalensis genome was annotated using the Maker2 pipeline54. Repetitive regions comprised 33% of the genome and were soft-masked using RepeatMasker. Several transcriptome assemblies were used to annotate genes. These included draft assemblies of the M. natalensis forelimb and hindlimb RNA-seq data for each of the three time points (six assemblies) and a pooled assembly of all the RNA-seq data. Combined, these yielded 6.1 million transcripts that were aligned to the genome using BLAST55. In addition, 960,000 M. brandtii RNA-seq transcripts, from the liver, kidney and brain15, were aligned using relaxed BLASTN settings (75% coverage, 80% identity and an e-value cutoff of 5 × 10−9), and 51,778 mouse proteins from the RefSeq protein database were aligned using BLASTP. After alignment, Exonerate56 was used to clear up intron-exon boundaries. Ab initio gene prediction was performed by SNAP57, which was trained on the earlier annotation, and AUGUSTUS58, which was run using human optimization. Once annotation was complete, gene predictions with poor evidence (annotation edit distance (AED) >0.75) were ignored. Finally, PASA59 was used to identify and confirm alternatively spliced transcripts.
RNA extraction, sequencing and analysis.
RNA was extracted from paired forelimbs and hindlimbs from three individuals (biological replicates) at three developmental stages (CS15, CS16 and CS17) using the RNeasy Midi kit (Qiagen). All bat embryos were staged according to Hockman et al.10. Total RNA samples were enriched for polyadenylated transcripts using the Oligotex mRNA Mini kit (Qiagen), and strand-specific RNA-seq libraries60 were generated using PrepX RNA library preparation kits (IntegenX), following the manufacturer's protocol. After cleanup with AMPure XP beads (Beckman Coulter) and amplification with Phusion High-Fidelity polymerase (New England BioLabs), RNA libraries were sequenced on a HiSeq 2500 instrument to a depth of at least 30 million reads (submitted to the SRA under accession SRP051253). For de novo transcriptome analysis, raw reads were quality trimmed and adaptor sequences were removed using Trimmomatic48. Two de novo assembly strategies were employed (Supplementary Fig. 9). First, all three replicates for each tissue-stage combination were pooled and assembled separately using Trinity61. Second, reads from all stages and tissues were pooled and underwent digital downsampling and assembly using the Trinity pipeline61. All de novo assemblies were then used in the Maker2 pipeline54 to improve gene annotations. The sequences of 436 transcripts from 227 genes that did not have a match in the mammalian UniProt database were compared to sequences in lncRNAdb v2.0 (ref. 22) and the GENCODE v7 lncRNA gene annotation database62. These noncoding transcripts were also compared using BLAST55 to mouse, human, dog, horse, cat and other bat genomes to identify new lncRNA transcripts that were conserved either in bats or in a subset of mammals. The Coding Potential Calculator was used to score whether a transcript was likely to be coding or noncoding63. For differential expression analysis, raw sequencing reads were mapped to the M. natalensis draft genome using TopHat64. Read counts for each gene were calculated for each replicate using HTSeq65, and tests of differential expression were carried out using DESeq2 (ref. 66). After differential expression testing, genes with P values adjusted for multiple testing (FDR) <0.01 in any of the five differential expression tests were clustered for similar expression using the R package hclust and displayed in a heat map. Additionally, genes found to be differentially expressed in forelimb and hindlimb across all stages were grouped on the basis of specific GO categories and subjected to analysis by clustered heat maps.
In situ hybridization.
Mouse embryos (C57BL/6 strain UCT3) were supplied by the Animal Research Facility, University of Cape Town. M. natalensis embryos were collected from a maternity roost at the De Hoop Nature Reserve, South Africa (Cape Nature Conservation permit number AAA007-00133-0056) as previously described67. Ethical approval by the University of Cape Town for sampling of bats was granted by the Science Faculty Animal Ethics committee (2012/V41/NI and 2012/V39/NI) and for the use of mouse embryos was granted by the Health Science Animal Ethics Committee (FHS AEC 014/07). Bat and mouse embryos from equivalent stages were matched as described by Hockman et al.10. Fixation and storage of embryos and whole-mount in situ hybridization probe synthesis and conditions were as previously described6. The primers used to generate the probes for whole-mount in situ hybridization are summarized in Supplementary Table 10.
Mllt3 mouse skeletal preparations.
Skeletons of newborn Mllt3 homozygous knockouts19 and wild-type littermates were stained for cartilage with Alcian blue and for bone with Alizarin red as previously described68. Briefly, newborn mice were euthanized, skinned, eviscerated, fixed in 95% ethanol for several days and then incubated at 37 °C for 2 d in Alcian blue stain (15 mg of Alcian blue in 80 ml of 95% ethanol and 20 ml of glacial acetic acid). Samples were rinsed twice in 95% ethanol for 2 h each. Specimens were cleared in 1% potassium hydroxide for 4–5 h and counterstained overnight with Alizarin red stain (50 mg of Alizarin red in 1 l 2% potassium hydroxide). Finally, samples were cleared in 20% glycerol and 1% potassium hydroxide followed by 50% glycerol and 1% potassium hydroxide for several days for each step and then stored in 80% glycerol.
ChIP-seq analysis.
Developing bat forelimbs and hindlimbs (dissected from CS15, CS16 and CS17 embryos) were cross-linked with 1% formaldehyde for 10 min. Reactions were quenched with glycine, and samples were flash-frozen in the field. Cross-linked limbs were then combined in the laboratory into pools of 4–7 pairs per stage for chromatin sheering using a Covaris S2 sonicator. Sheared chromatin was used for immunoprecipitation with antibodies against active (H3K27ac; Abcam, ab4729) and repressive (H3K27me3; Millipore, 07-449) chromatin marks using the Diagenode LowCell# ChIP kit, following the manufacture's protocol. Libraries were prepared using the Rubicon ThruPLEX-FD Prep kit following the manufacturer's protocol and sequenced on an Illumina HiSeq 2500 instrument using single-end 50-bp reads to a sequencing depth of at least 25 million reads (submitted to the SRA under accession SRP051267). Uniquely mapping raw reads were aligned using Bowtie69 with default settings. Peak regions for each histone mark were called using SICER70, informed by the estimated average fragment size of the chromatin after shearing, as measured by Agilent 2100 Bioanalyzer. Peaks from all samples were merged using BEDTools71 and partitioned using BEDOPS72 (Supplementary Fig. 10). Differentially enriched regions in forelimb and hindlimb for each stage and histone mark were obtained following a methodology similar to MAnorm, which uses a linear model that assumes that peaks shared by samples can serve to normalize ChIP-seq data sets for differing signal-to-noise ratios. In this methodology, genomic regions not appearing as a peak in any sample were obtained from the partitioned regions using BEDTools and were used to normalize for the background noise present in each sample. Furthermore, a set of common regions for each histone mark was obtained, and these regions were used to normalize the ChIP-seq signal between all samples by creating a scaling factor based on the average signal in the shared peaks minus the average noise in non-peak regions. After removing duplicate reads with Picard MarkDuplicates, read counts were obtained with the BEDTools coverage command. The average noise from each region based on its genomic size was subtracted from each region's read counts. Noise-subtracted read counts were then normalized by multiplying each by the signal scaling factor and a read depth scaling factor, to create an enrichment score for each portioned region. Pairwise differential enrichment tests were carried out using a Bayesian model73, which is also used by MAnorm, followed by adjustment for multiple testing using the R package p.adjust.
Comparative genomics.
Whole-genome alignments were carried out using LASTZ74 with soft-masked genome assemblies from 18 species (E. fuscus, M. brandti, M. davidii, M. lucifugus, P. alecto, P. vampyrus, Bos taurus, Canis familiaris, Equus caballus, Felis catus, Homo sapiens, Loxodonta africana, Monodelphis domestica, Mus musculus, Sus scrofa, Danio rerio, Anolis carolinensis and Gallus gallus) using the repeat-masked M. natalensis genome as a reference. If no publically available repeat-masked genome was available, RepeatMasker was run using the mammal repeat database and default conditions. Alignment files were then chained, netted and converted to MAF files using UCSC Genome Browser utilities. Individual MAF files from each pairwise species alignment were combined into a multiple MAF file using the roast command, which is part of the Multiz-TBA package75. Tree models for both conserved and non-conserved sequences were created for the species used in the multiple MAF file by phyloFit76. These tree models were used in phastCons76 to identify vertebrate conserved sequences in the M. natalensis genome and to generate base-by-base conservation scores to be displayed in genome browsers. BARs were identified by using phyloP28,76 to test for acceleration in the common ancestor of the bat lineage over regions identified as vertebrate conserved sequences after filtering for quality alignments. Genomic regions enriched for BARs were identified by scanning the genome using a 100-kb sliding window with a step size of 50 kb while counting BARs and phyloP-tested regions within them. On average, phyloP found acceleration in 0.812% of the sequences tested. The expected number of BARs in each region was then set to be the number of sequences tested by phyloP in that region multiplied by 0.00812. Regions enriched for BARs were identified by comparing the average expected number of BARs to the observed number of BARs using a Poisson test. After correction for multiple testing, genes contained in or overlapping the genomic regions with significant over-representation of BARs were analyzed for functional annotation clustering using DAVID77,78 with the background set to the genes contained in regions with valid multiple-sequence alignments and H3K27ac peaks.
Ingenuity Pathway Analysis.
Pairwise differential expression testing for forelimb and hindlimb at each stage identified a total of 3,140 bat genes (FDR < 0.05). This list was filtered for genes that had an average FPKM value >2 and that had been mapped to a human Entrez Gene ID, resulting in 2,751 genes. IPA (Qiagen) was used to analyze this set of 2,751 genes to determine whether specific canonical signaling pathways and their upstream regulators were coordinately regulated across three developmental stages (CS15, CS16 and CS17) using fold change values. A Fisher's exact right-tailed test identified significantly enriched pathways, and a z score was computed to determine whether the pathway was activated or inhibited at each stage. IPA was also used to predict upstream regulators that would explain the patterns of differential gene expression observed across the data set.
Coherency (marked by 1) was tested by comparing significant differences between forelimb and hindlimb ChIP-seq and RNA-seq signals for genes that were differentially expressed in the top ten canonical IPA pathways (Fig. 5a). Significantly different acetylation marks were required to be antagonistic to their equivalent methylation marks, with at least a single mark being significantly different in the forelimb and hindlimb. RNA-seq levels in the forelimb and hindlimb were also required to be positively correlated with any significantly different acetylation marks.
Histochemistry.
Bat embryos were fixed in 4% paraformaldehyde for 3 h at room temperature, washed in PBS and stored in 30% sucrose in PBS at 4 °C for 5–6 d. Whole limbs were dissected from these embryos and embedded in tissue freezing medium (Leica Biosystems). These were sectioned to be 8 μm thick, using a Leica CM1850 cryotome at −17 °C, collected on Superfrost Plus (Thermo Scientific) slides and stored at −70 °C. Serial sections were stained with either hematoxylin and eosin or PNA. Slides containing sections were fixed in phosphate-buffered formalin for 5 min, washed in distilled water for 1 min and then stained with hematoxylin for 30 s. Slides were rinsed in 0.3% acid ethanol (70% ethanol) and running water and incubated in Scott's water for 1 min. After rinses in running water and 80% alcohol, slides were stained with acid-based eosin for 2.5 min. Slides were dehydrated through an ethanol series and dipped in xylene, and coverslips were secured with DPX mountant (Sigma).
For PNA staining, bat autopod sections were fixed for 10 min in acetone. Slides were washed three times in PBS and blocked in 3% BSA in PBS for 1 h at room temperature. Sections were incubated with 100 μg/ml FITC-conjugated PNA (Sigma, L7381) in 3% BSA in PBS at 4 °C overnight. Control slides were incubated in 3% BSA in PBS only. All slides were washed in PBS and stained for 10 min in 1 μg/ml Hoechst nuclear stain, before another three PBS washes. ProLong Gold antifade reagent (Life Technologies) was used to mount coverslips. Sections were photographed on a Nikon Ti-E inverted fluorescent microscope using the same standardized camera setting for all sections.
Accession codes.
The whole-genome shotgun assembly is available under BioProject PRJNA283550. RNA-seq data are available from the Sequence Read Archive (SRA) under accession PRJNA270639 (SRP051253). ChIP-seq data are available from the SRA under accession PRJNA270665 (SRP051267).
URLs.
Ingenuity Pathway Analysis (IPA), http://www.qiagen.com/ingenuity/; Picard MarkDuplicates, http://broadinstitute.github.io/picard; RepeatMasker, http://www.repeatmasker.org/.
Accession codes
Primary accessions
BioProject
Sequence Read Archive
Referenced accessions
NCBI Reference Sequence
References
Gunnell, G.F. & Simmons, N.B. Evolutionary History of Bats: Fossils, Molecules and Morphology (Cambridge Univ. Press, 2012).
Cooper, L.N., Cretekos, C.J. & Sears, K.E. The evolution and development of mammalian flight. Wiley Interdiscip. Rev. Dev. Biol. 1, 773–779 (2012).
Swartz, S.M. & Middleton, K.M. Biomechanics of the bat limb skeleton: scaling, material properties and mechanics. Cells Tissues Organs 187, 59–84 (2008).
Hockman, D. et al. A second wave of Sonic hedgehog expression during the development of the bat limb. Proc. Natl. Acad. Sci. USA 105, 16982–16987 (2008).
Wang, Z. et al. Unique expression patterns of multiple key genes associated with the evolution of mammalian flight. Proc. Biol. Soc. 281, 20133133 (2014).
Mason, M.K. et al. Retinoic acid–independent expression of Meis2 during autopod patterning in the developing bat and mouse limb. Evodevo 6, 6 (2015).
Wang, Z. et al. Digital gene expression tag profiling of bat digits provides robust candidates contributing to wing formation. BMC Genomics 11, 619 (2010).
Weatherbee, S.D., Behringer, R.R., Rasweiler, J.J. & IV & Niswander, L.A. Interdigital webbing retention in bat wings illustrates genetic changes underlying amniote limb diversification. Proc. Natl. Acad. Sci. USA 103, 15103–15107 (2006).
Cretekos, C.J. et al. Regulatory divergence modifies limb length between mammals. Genes Dev. 22, 141–151 (2008).
Hockman, D., Mason, M.K., Jacobs, D.S. & Illing, N. The role of early development in mammalian limb diversification: a descriptive comparison of early limb development between the Natal long-fingered bat (Miniopterus natalensis) and the mouse (Mus musculus). Dev. Dyn. 238, 965–979 (2009).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Creyghton, M.P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 107, 21931–21936 (2010).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Zhang, G. et al. Comparative analysis of bat genomes provides insight into the evolution of flight and immunity. Science 339, 456–460 (2013).
Seim, I. et al. Genome analysis reveals insights into physiology and longevity of the Brandt's bat Myotis brandtii. Nat. Commun. 4, 2212 (2013).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Lallemand, Y., Bensoussan, V., Cloment, C.S. & Robert, B. Msx genes are important apoptosis effectors downstream of the Shh/Gli3 pathway in the limb. Dev. Biol. 331, 189–198 (2009).
Collins, E.C. et al. Mouse Af9 is a controller of embryo patterning, like Mll, whose human homologue fuses with Af9 after chromosomal translocation in leukemia. Mol. Cell. Biol. 22, 7313–7324 (2002).
Zhao, Y. et al. The LIM-homeobox gene Lhx8 is required for the development of many cholinergic neurons in the mouse forebrain. Proc. Natl. Acad. Sci. USA 100, 9005–9010 (2003).
Wang, K.C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Quek, X.C. et al. lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 43, D168–D173 (2015).
Agarwal, P. et al. Tbx5 is essential for forelimb bud initiation following patterning of the limb field in the mouse embryo. Development 130, 623–633 (2003).
Rallis, C. et al. Tbx5 is required for forelimb bud formation and continued outgrowth. Development 130, 2741–2751 (2003).
Carroll, S.B. Evolution at two levels: on genes and form. PLoS Biol. 3, e245 (2005).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Booker, B. et al. Bat accelerated regions identify a bat forelimb specific enhancer in the HoxD locus. PLoS Genet. 12, e1005738 (2016).
Pollard, K.S., Hubisz, M.J., Rosenbloom, K.R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Homma, S., Shimada, T., Hikake, T. & Yaginuma, H. Expression pattern of LRR and Ig domain–containing protein (LRRIG protein) in the early mouse embryo. Gene Expr. Patterns 9, 1–26 (2009).
Tossell, K. et al. Lrrn1 is required for formation of the midbrain-hindbrain boundary and organiser through regulation of affinity differences between midbrain and hindbrain cells in chick. Dev. Biol. 352, 341–352 (2011).
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010).
Taylor, G.P., Anderson, R., Reginelli, A.D. & Muneoka, K. FGF-2 induces regeneration of the chick limb bud. Dev. Biol. 163, 282–284 (1994).
Eloy-Trinquet, S., Wang, H., Edom-Vovard, F. & Duprez, D. Fgf signaling components are associated with muscles and tendons during limb development. Dev. Dyn. 238, 1195–1206 (2009).
Kondrashov, N. et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 145, 383–397 (2011).
Ball, S. Diamond Blackfan anemia. Hematology (Am. Soc. Hematol. Educ. Program) 2011, 487–491 (2011).
Xue, S. et al. RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 517, 33–38 (2015).
Chen, J., Holguin, N., Shi, Y., Silva, M.J. & Long, F. mTORC2 signaling promotes skeletal growth and bone formation in mice. J. Bone Miner. Res. 30, 369–378 (2015).
Martin, G.R. The roles of FGFs in the early development of vertebrate limbs. Genes Dev. 12, 1571–1586 (1998).
Kozhemyakina, E., Lassar, A.B. & Zelzer, E. A pathway to bone: signaling molecules and transcription factors involved in chondrocyte development and maturation. Development 142, 817–831 (2015).
Gao, B. & Yang, Y. Planar cell polarity in vertebrate limb morphogenesis. Curr. Opin. Genet. Dev. 23, 438–444 (2013).
Bisson, J.A., Mills, B., Paul Helt, J.C., Zwaka, T.P. & Cohen, E.D. Wnt5a and Wnt11 inhibit the canonical Wnt pathway and promote cardiac progenitor development via the Caspase-dependent degradation of AKT. Dev. Biol. 398, 80–96 (2015).
Hall, B.K. & Miyake, T. Divide, accumulate, differentiate: cell condensation in skeletal development revisited. Int. J. Dev. Biol. 39, 881–893 (1995).
Sears, K.E., Behringer, R.R., Rasweiler, J.J. IV & Niswander, L.A. Development of bat flight: morphologic and molecular evolution of bat wing digits. Proc. Natl. Acad. Sci. USA 103, 6581–6586 (2006).
Pignatti, E., Zeller, R. & Zuniga, A. To BMP or not to BMP during vertebrate limb bud development. Semin. Cell Dev. Biol. 32, 119–127 (2014).
Kingsley, D.M. et al. The mouse short ear skeletal morphogenesis locus is associated with defects in a bone morphogenetic member of the TGF β superfamily. Cell 71, 399–410 (1992).
Francis-West, P.H. et al. Mechanisms of GDF-5 action during skeletal development. Development 126, 1305–1315 (1999).
Merino, R. et al. The BMP antagonist Gremlin regulates outgrowth, chondrogenesis and programmed cell death in the developing limb. Development 126, 5515–5522 (1999).
Bolger, A.M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Magoč, T. & Salzberg, S.L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Xu, H. et al. FastUniq: a fast de novo duplicates removal tool for paired short reads. PLoS One 7, e52249 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Slater, G.S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Johnson, A.D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24, 2938–2939 (2008).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Haas, B.J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Borodina, T., Adjaye, J. & Sultan, M. A strand-specific library preparation protocol for RNA sequencing. Methods Enzymol. 500, 79–98 (2011).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Harrow, J. et al. GENCODE: the reference human genome annotation for the ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P.T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Mason, M.K., Hockman, D., Jacobs, D.S. & Illing, N. Evaluation of maternal features as indicators of asynchronous embryonic development in Miniopterus natalensis. Acta Chiropt. 12, 161–171 (2010).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Manipulating the Mouse Embryo: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2002).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25, 1952–1958 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Audic, S. & Claverie, J.M. The significance of digital gene expression profiles. Genome Res. 7, 986–995 (1997).
Harris, R.S. Improved Pairwise Alignment of Genomic DNA. PhD thesis, Penn. State Univ. (2007).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Hubisz, M.J., Pollard, K.S. & Siepel, A. PHAST and RPHAST: phylogenetic analysis with space/time models. Brief. Bioinform. 12, 41–51 (2011).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang, W., Sherman, B.T. & Lempicki, R.A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Acknowledgements
We thank the Broad Institute Genomics Platform, Vertebrate Genome Biology group and K. Lindblad-Toh for making the data for Eptesicus fuscus available. We would also like to thank M. Kelley and E. Driver (National Institute on Deafness and Other Communication Disorders) for providing us with newborn Mllt3 mice and M. Petersen for assistance with sectioning bat autopods. This work was supported in part by National Institute of Child Health and Human Development grant 1R01HD059862. N.A. is also supported in part by National Institute of Diabetes and Digestive and Kidney Diseases award 1R01DK090382, National Institute of Neurological Disorders and Stroke award 1R01NS079231 and National Cancer Institute award 1R01CA197139. N.I. is supported by National Research Foundation (NRF; South Africa) grants (NBIG 86932), and C.M.-N. was supported by NRF grant 85207. L.C. and K.A.N. are supported by the Office of the Director, US National Institutes of Health, under award P51OD011092. T.F. and K.S.P. are supported by a gift from the San Simeon Fund and institutional funds from the J. David Gladstone Institutes.
Author information
Authors and Affiliations
Contributions
W.L.E., S.A.S., M.K.M., J.E.V., J.D.W., N.I. and N.A. conceived key aspects of the project and planned experiments. S.A.S., M.K.M., Z.G., A.V.P., C.M.-N. and N.I. collected bat embryos. S.A.S. and C.M.-N. extracted genomic DNA and RNA from bats. E.T., K.A.N. and L.C. generated genome sequencing libraries. W.L.E., S.A.S., M.K.M., Z.G., A.V.P., B.M.B., S.N., N.M., J.E.V. and N.I. performed experiments. T.F. and K.S.P. developed statistical methods. W.L.E., S.A.S., M.K.M., Z.G., T.F., L.C., K.S.P., J.D.W., N.I. and N.A. analyzed data. W.L.E., S.A.S., M.K.M., L.C., N.I. and N.A. wrote the manuscript. All authors commented on and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Gene clustering based on expression data from RNA-seq.
(a) Dendrogram of gene clustering denoting the 38 manually defined clusters. (b) Box plots showing expression changes (z scores) for each developmental stage (blue, CS15; green, CS16; red, CS17) and tissue (light, forelimbs (FL); dark, hindlimbs (HL)) in the 38 manually defined clusters.
Supplementary Figure 2 RNA-seq heat maps of genes differentially expressed in forelimbs and hindlimbs from selected GO categories.
The GO categories include limb morphogenesis (GO:0035108) (a), apoptotic process (GO:0006915) (b), cartilage development (GO:0051216) (c), bone morphogenesis (GO:0060349) (d), skeletal muscle tissue development (GO:0007519) (e), and stem cell proliferation and stem cell division (GO:0072089 and GO:0017145) (f).
Supplementary Figure 3 Whole-mount in situ hybridization of differentially expressed candidate transcription factors and lncRNAs in developing bat and mouse forelimbs and hindlimbs, at equivalent developmental stages.
(a) Hoxa13 has reduced expression in bat forelimbs as compared to hindlimbs at stages CS15–CS17. In mice, Hoxa13 expression is comparable in forelimbs and hindlimbs at all developmental stages. (b) Hottip, a lncRNA encoded near Hoxa13, displays a similar pattern of expression as Hoxa13, with reduced expression in bat forelimbs relative to hindlimbs. (c) Tbx5-as1, a lncRNA encoded near Tbx5, is expressed in a restricted region at the base of digits I–V during late CS16 (L) and CS17, with clear expression in proximal interdigit tissue. (d) At CS15, the transcription factor Mllt3 displays strong expression in the posterior-distal mesenchyme running from presumptive digits II–V in the forelimb, while expression in the hindlimb is restricted to the most proximal and posterior region of the autopod, extending to the zeugopod and presumptive calcar. At CS16, expression is faint between digits IV and V and is absent from the hindlimb. In mice at E11.5, there is similar expression for forelimb and hindlimb, with expression following the outer edge of the limb buds and concentrated in the posterior region. By E13.5, expression of Mllt3 is absent from the mouse limbs. (e) Expression of the transcription factor Lef1 is reduced in forelimb relative to hindlimb at CS15 and appears in a comet in the anterior region of the joint between the autopod and zeugopod. Expression is faint in both forelimb and hindlimb by CS16 and is completely absent from mouse limbs at both stages. (f) At CS15, there is weak expression of the transcription factor Lhx8 in the region of presumptive forelimb digits II–IV and in the posterior half of the hindlimb. At CS16L, Lhx8 expression is strong along the outer edges of the forelimb zeugopod, with weak expression in the interdigit tissue between digits III and IV, and is absent from hindlimbs. Lhx8 expression was not detectable in mouse limbs at equivalent stages. Scale bars, 0.5mm.
Supplementary Figure 4 Limb morphology of the Mllt3 knockout mouse.
(a–f) Newborn wild-type (+/+) (a–c) and Mllt3 homozygous knockout (Mllt3−/−; also known as Af9−/−) (d–f) littermates stained with Alizarin red and Alcian blue (b,e, forelimbs; c,f, hindlimbs). Scale bars, 1 mm.
Supplementary Figure 5 GREAT analysis of H3K27ac peak regions in bat forelimbs and hindlimbs after liftOver to mouse mm9 coordinates.
(a–f) Functional enrichment of bat H3K27ac peak regions for GO Biological Process categories (a, forelimb; d, hindlimb), MGI expression domains (b, forelimb; e, hindlimb) and mouse phenotypes (c, forelimb; f, hindlimb).
Supplementary Figure 6 Partitioned peak regions clustered on the basis of enrichment data from H3K27ac and H3K27me3 ChIP-seq analysis.
(a) Dendrogram of partitioned peak region clusters denoting the 17 manually defined clusters. (b) Box plots showing H3K27ac (green) and H3K27me3 (red) ChIP-seq–derived enrichment changes (z scores) and RNA-seq–derived expression changes of the nearest gene (blue) for each stage and tissue (dark, forelimb (F); light, hindlimb (H)) of the 17 manually defined clusters.
Supplementary Figure 7 Bat accelerated sequences.
(a) The top genomic locus enriched for BARs on M. natalensis scaffold 228, which contains the Crbn and Lrrn1 genes. RNA-seq and ChIP-seq tracks are represented as forelimb (FL) coverage minus hindlimb (HL) coverage in 100-bp intervals for each stage. For RNA-seq data, intervals with forelimb minus hindlimb expression values greater than zero are shown in dark blue and those with values less than zero are shown in light blue. Likewise, for ChIP-seq data, intervals with forelimb minus hindlimb enrichment values greater than zero are colored in dark green (H3K27ac) and dark red (H3K27me3) and those with values less than zero are colored in light green (H3K27ac) and light red (H3K27me3). A BAR track depicting BARs as blue lines and a vertebrate conservation track are also shown below. (b) The Fgf2-Spry1 genomic locus. (c) Fgf2 and Spry1 RNA-seq expression levels.
Supplementary Figure 8 Hematoxylin and eosin (H&E) staining of sagittal sections of bat CS15, CS16 and CS17 forelimbs and hindlimbs.
(a–l) H&E staining of sagittal sections of bat CS15, CS16 and CS17 forelimb (FL) (a–c) and hindlimb (HL) (g–i) autopods. These sections are adjacent to the sections shown in Figure 6. Higher magnification views are shown of the boxed regions, which correspond to the areas with the strongest PNA staining at CS15 (d and j), digits 3 and 4 at CS16 (e and k), and digit 4 at CS17 (f and l). Ellipses indicate regions of maturing chondrocytes. Scale bars: 500 μm in a–c and g–i, 100 μm in d–f and j–l.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10. (PDF 8744 kb)
Supplementary Table 1
M. natalensis genome assembly descriptive statistics. (XLSX 8 kb)
Supplementary Table 2
RNA-seq differential expression results, raw read counts and normalized read counts. (XLSX 13876 kb)
Supplementary Table 3
Functional annotation enrichment in differentially expressed gene clusters. (XLSX 1217 kb)
Supplementary Table 4
Predicted bat lncRNAs. (XLSX 32 kb)
Supplementary Table 5
ChIP-seq peak regions. (XLSX 16475 kb)
Supplementary Table 6
ChIP-seq differential enrichment analysis. (XLSX 53355 kb)
Supplementary Table 7
Functional annotation enrichment in differentially enriched ChIP-seq clusters. (XLSX 269 kb)
Supplementary Table 8
Analysis of bat accelerated regions (BARs). (XLSX 921 kb)
Supplementary Table 9
RNA-seq, ChIP-seq and BAR data for genes differentially expressed in canonical IPA and manually curated pathways. (XLSX 245 kb)
Supplementary Table 10
Primers used for mouse transgenic assays and WISH. (XLSX 9 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the articles Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Eckalbar, W., Schlebusch, S., Mason, M. et al. Transcriptomic and epigenomic characterization of the developing bat wing. Nat Genet 48, 528–536 (2016). https://doi.org/10.1038/ng.3537
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3537
This article is cited by
-
Boundary stacking interactions enable cross-TAD enhancer–promoter communication during limb development
Nature Genetics (2024)
-
Insights into the formation and diversification of a novel chiropteran wing membrane from embryonic development
BMC Biology (2023)
-
Gene expression changes during the evolution of the tetrapod limb
Biologia Futura (2022)
-
Evidence that DNA repair genes, a family of tumor suppressor genes, are associated with evolution rate and size of genomes
Human Genomics (2019)
-
Creating diversity in mammalian facial morphology: a review of potential developmental mechanisms
EvoDevo (2018)