Abstract
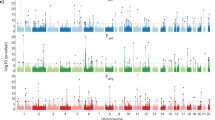
We simultaneously investigated the genetic landscape of ankylosing spondylitis, Crohn's disease, psoriasis, primary sclerosing cholangitis and ulcerative colitis to investigate pleiotropy and the relationship between these clinically related diseases. Using high-density genotype data from more than 86,000 individuals of European ancestry, we identified 244 independent multidisease signals, including 27 new genome-wide significant susceptibility loci and 3 unreported shared risk loci. Complex pleiotropy was supported when contrasting multidisease signals with expression data sets from human, rat and mouse together with epigenetic and expressed enhancer profiles. The comorbidities among the five immune diseases were best explained by biological pleiotropy rather than heterogeneity (a subgroup of cases genetically identical to those with another disease, possibly owing to diagnostic misclassification, molecular subtypes or excessive comorbidity). In particular, the strong comorbidity between primary sclerosing cholangitis and inflammatory bowel disease is likely the result of a unique disease, which is genetically distinct from classical inflammatory bowel disease phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Parkes, M., Cortes, A., van Heel, D.A. & Brown, M.A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 14, 661–673 (2013).
Najarian, D.J. & Gottlieb, A.B. Connections between psoriasis and Crohn's disease. J. Am. Acad. Dermatol. 48, 805–821, quiz 822–824 (2003).
Loftus, E.V. Jr. et al. PSC-IBD: a unique form of inflammatory bowel disease associated with primary sclerosing cholangitis. Gut 54, 91–96 (2005).
Jensen, A.B. et al. Temporal disease trajectories condensed from population-wide registry data covering 6.2 million patients. Nat. Commun. 5, 4022 (2014).
Bhattacharjee, S. et al. A subset-based approach improves power and interpretation for the combined analysis of genetic association studies of heterogeneous traits. Am. J. Hum. Genet. 90, 821–835 (2012).
Stahl, E.A. et al. Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat. Genet. 42, 508–514 (2010).
Grant, S.F. et al. Follow-up analysis of genome-wide association data identifies novel loci for type 1 diabetes. Diabetes 58, 290–295 (2009).
Wang, Z. et al. Imputation and subset-based association analysis across different cancer types identifies multiple independent risk loci in the TERT-CLPTM1L region on chromosome 5p15.33. Hum. Mol. Genet. 23, 6616–6633 (2014).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Fehrmann, R.S. et al. Trans-eQTLs reveal that independent genetic variants associated with a complex phenotype converge on intermediate genes, with a major role for the HLA. PLoS Genet. 7, e1002197 (2011).
Nelis, M. et al. Genetic structure of Europeans: a view from the North-East. PLoS One 4, e5472 (2009).
Trynka, G. et al. Disentangling the effects of colocalizing genomic annotations to functionally prioritize non-coding variants within complex-trait loci. Am. J. Hum. Genet. 97, 139–152 (2015).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
FANTOM Consortium and the RIKEN PMI and CLST (DGT). A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Pers, T.H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Fehrmann, R.S. et al. Gene expression analysis identifies global gene dosage sensitivity in cancer. Nat. Genet. 47, 115–125 (2015).
Kamburov, A. et al. ConsensusPathDB: toward a more complete picture of cell biology. Nucleic Acids Res. 39, D712–D717 (2011).
Han, B. et al. Using genotype data to distinguish pleiotropy from heterogeneity: deciphering coheritability in autoimmune and neuropsychiatric diseases. bioRxiv doi:10.1101/030783 (6 November 2015).
Lee, S.H., Wray, N.R., Goddard, M.E. & Visscher, P.M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Lee, S.H., Yang, J., Goddard, M.E., Visscher, P.M. & Wray, N.R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism–derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Chen, G.B. et al. Estimation and partitioning of (co)heritability of inflammatory bowel disease from GWAS and immunochip data. Hum. Mol. Genet. 23, 4710–4720 (2014).
Grant, A.J., Lalor, P.F., Salmi, M., Jalkanen, S. & Adams, D.H. Homing of mucosal lymphocytes to the liver in the pathogenesis of hepatic complications of inflammatory bowel disease. Lancet 359, 150–157 (2002).
Adams, D.H. & Eksteen, B. Aberrant homing of mucosal T cells and extra-intestinal manifestations of inflammatory bowel disease. Nat. Rev. Immunol. 6, 244–251 (2006).
Apetoh, L. et al. Toll-like receptor 4–dependent contribution of the immune system to anticancer chemotherapy and radiotherapy. Nat. Med. 13, 1050–1059 (2007).
Prohinar, P., Rallabhandi, P., Weiss, J.P. & Gioannini, T.L. Expression of functional D299G.T399I polymorphic variant of TLR4 depends more on coexpression of MD-2 than does wild-type TLR4. J. Immunol. 184, 4362–4367 (2010).
Isakov, N. & Altman, A. PKCθ-mediated signal delivery from the TCR/CD28 surface receptors. Front. Immunol. 3, 273 (2012).
Wachowicz, K. & Baier, G. Protein kinase Cθ: the pleiotropic T-cell signalling intermediate. Biochem. Soc. Trans. 42, 1512–1518 (2014).
Zanin-Zhorov, A. et al. Protein kinase Cθ mediates negative feedback on regulatory T cell function. Science 328, 372–376 (2010).
Anderson, C.A. et al. Meta-analysis identifies 29 additional ulcerative colitis risk loci, increasing the number of confirmed associations to 47. Nat. Genet. 43, 246–252 (2011).
Solovieff, N., Cotsapas, C., Lee, P.H., Purcell, S.M. & Smoller, J.W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Karlsen, T.H. & Boberg, K.M. Update on primary sclerosing cholangitis. J. Hepatol. 59, 571–582 (2013).
de Vries, A.B., Janse, M., Blokzijl, H. & Weersma, R.K. Distinctive inflammatory bowel disease phenotype in primary sclerosing cholangitis. World J. Gastroenterol. 21, 1956–1971 (2015).
Fairfax, B.P. et al. Genetics of gene expression in primary immune cells identifies cell type–specific master regulators and roles of HLA alleles. Nat. Genet. 44, 502–510 (2012).
International Genetics of Ankylosing Spondylitis Consortium (IGAS). Identification of multiple risk variants for ankylosing spondylitis through high-density genotyping of immune-related loci. Nat. Genet. 45, 730–738 (2013).
Tsoi, L.C. et al. Identification of 15 new psoriasis susceptibility loci highlights the role of innate immunity. Nat. Genet. 44, 1341–1348 (2012).
Liu, J.Z. et al. Dense genotyping of immune-related disease regions identifies nine new risk loci for primary sclerosing cholangitis. Nat. Genet. 45, 670–675 (2013).
Goyette, P. et al. High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat. Genet. 47, 172–179 (2015).
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat. Genet. 43, 1193–1201 (2011).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Price, A.L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–135, author reply 135–139 (2008).
Abraham, G. & Inouye, M. Fast principal component analysis of large-scale genome-wide data. PLoS One 9, e93766 (2014).
Taylor, J.E., Worsley, K.J. & Gosselin, F. Maxima of discretely sampled random fields, with an application to 'bubbles'. Biometrika 94, 1–18 (2007).
Tsoi, L.C. et al. Enhanced meta-analysis and replication studies identify five new psoriasis susceptibility loci. Nat. Commun. 6, 7001 (2015).
Liu, J.Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Morris, J.A., Randall, J.C., Maller, J.B. & Barrett, J.C. Evoker: a visualization tool for genotype intensity data. Bioinformatics 26, 1786–1787 (2010).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010).
Ng, P.C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Chang, C.C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Acknowledgements
We thank all patients with ankylosing spondylitis, Crohn's disease, PSC, psoriasis and ulcerative colitis, their families, healthy control individuals and clinicians for their participation in this study. We thank T. Wesse, T. Henke, S. Sedghpour Sabet, R. Vogler, G. Jacobs, I. Urbach, W. Albrecht, V. Pelkonen, K. Holm, H. Dahlen Sollid, B. Woldseth, J.A. Anmarkrud and L. Wenche Torbjørnsen for expert help. F. Braun, W. Kreisel, T. Berg and R. Günther are acknowledged for contributing German patients with PSC. B.A. Lie and the Norwegian Bone Marrow Donor Registry at Oslo University Hospital, Rikshospitalet, in Oslo and the Nord-Trøndelag Health Study (HUNT) are acknowledged for sharing the healthy Norwegian controls. This work was supported by the German Federal Ministry of Education and Research (BMBF) within the framework of the e:Med research and funding concept (SysInflame grant 01ZX1306A). This project received infrastructure support from Deutsche Forschungsgemeinschaft (DFG) Excellence Cluster 306 'Inflammation at Interfaces' and the PopGen Biobank. A.F. receives an endowment professorship by the Foundation for Experimental Medicine (Zurich, Switzerland). The Estonian Genome Center at the University of Tartu (EGCUT) received targeted financing from Estonian Research Council grant IUT20-60, the Center of Excellence in Genomics (EXCEGEN) and the University of Tartu (SP1GVARENG). We acknowledge EGCUT technical personnel, especially V. Soo and S. Smit. Data analyses were carried out in part at the High-Performance Computing Center of the University of Tartu. We acknowledge support from the UK Department of Health via National Institute for Health Research (NIHR) comprehensive Biomedical Research Centre awards to Guy's and St Thomas' National Health Service (NHS) Foundation Trust in partnership with King's College London and to Addenbrooke's Hospital in partnership with the University of Cambridge. The study was supported by the German Federal Ministry of Education and Research (BMBF), within the context of National Genome Research Network 2 (NGFN-2), National Genome Research Network plus (NGFNplus) and the Integrated Genome Research Network (IG) MooDS (grants 01GS08144 and 01GS08147). R.K.W. is supported by a VIDI grant (016.136.308) from the Netherlands Organization for Scientific Research (NWO). J.H. was supported by the Swedish Research Council (521-2011-2764). This work is supported in part by funding from the US NIH (1R01AR063759 (S.R.), 5U01GM092691-05 (S.R.), 1UH2AR067677-01 (S.R.), U19AI111224-01 (S.R.) and 1R01DK084960-05 (K.N.L.)) and Doris Duke Charitable Foundation grant 2013097. A.B.J. and S.B. acknowledge funding from the Novo Nordisk Foundation (grant NNF14CC0001) and the H2020 project MedBioinformatics (grant 634143). The study was supported by the Norwegian PSC Research Center. We thank G. Trynka for assistance in setting up GoShifter.
Author information
Authors and Affiliations
Consortia
Contributions
D.E., L.J., S.L.S., A.C., J.B., B.H., Y.R.P., J.G.P., S.R., Y.W., T.E., H.-J.W., L.F., T.H.P., R.K.W., V.C., O.A.A., A.B.J., S.B. and A.M.D. performed statistical and computational analyses. M.H. performed computational analyses. T.F., A.M., M.D'A., J.H., W.L., F.D., A.J.F., A.H., S.S., U.M., B.D.J., K.N.L., R.C.T., S.W., M.W., E.E., J.T.E., J.N.W.N.B. and M.A.B. were involved in study subject recruitment and assembling phenotypic data. D.E. wrote the draft of the manuscript. D.E., D.P.M., T.H.K., J.C.B., M.P., M.A.B. and A.F. conceived, designed and managed the study. All authors reviewed, edited and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1, 4 and 7–17, Supplementary Figures 1–3 and 7–17, and Supplementary Note. (PDF 4199 kb)
Supplementary Figure 4
Regional association plots for 244 independent association signals within 169 genome-wide significant non-MHC risk loci. (PDF 33785 kb)
Supplementary Figure 5
Synthesis-View plots showing the multidisease association signals for 244 independent association signals within 169 genome-wide significant non-MHC risk loci. (PDF 14842 kb)
Supplementary Figure 6
Pairwise comparisons of variance explained per risk variant between ankylosing spondylitis (AS), Crohn's disease (CD), psoriasis (PS), primary sclerosing cholangitis (PSC) and ulcerative colitis (UC) for a maximum of 244 independent signals from 169 risk loci. (PDF 382 kb)
Supplementary Table 2
Twenty-seven newly identified single-disease associations with genome-wide significance. (XLSX 25 kb)
Supplementary Table 3
Summary of 169 non-MHC genome-wide significant susceptibility loci. (XLSX 512 kb)
Supplementary Table 5
Functional in silico annotations of risk SNPs. (XLSX 1217 kb)
Supplementary Table 6
Analysis of cis-eQTL data from whole peripheral samples of 2,360 unrelated individuals. (XLSX 156 kb)
Rights and permissions
About this article
Cite this article
Ellinghaus, D., Jostins, L., Spain, S. et al. Analysis of five chronic inflammatory diseases identifies 27 new associations and highlights disease-specific patterns at shared loci. Nat Genet 48, 510–518 (2016). https://doi.org/10.1038/ng.3528
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3528
This article is cited by
-
Sequence of Events in the Pathogenesis of Axial Spondyloarthritis: A Current Review—2023 SPARTAN Meeting Proceedings
Current Rheumatology Reports (2024)
-
Perturbational phenotyping of human blood cells reveals genetically determined latent traits associated with subsets of common diseases
Nature Genetics (2024)
-
Quantitative proteomic screening uncovers candidate diagnostic and monitoring serum biomarkers of ankylosing spondylitis
Arthritis Research & Therapy (2023)
-
A genome-wide cross-trait analysis identifies genomic correlation, pleiotropic loci, and causal relationship between sex hormone-binding globulin and rheumatoid arthritis
Human Genomics (2023)
-
A review article of inflammatory bowel disease treatment and pharmacogenomics
Beni-Suef University Journal of Basic and Applied Sciences (2023)