Abstract
Translocation events are frequent in cancer and may create chimeric fusions or 'regulatory rearrangements' that drive oncogene overexpression. Here we identify super-enhancer translocations that drive overexpression of the oncogenic transcription factor MYB as a recurrent theme in adenoid cystic carcinoma (ACC). Whole-genome sequencing data and chromatin maps highlight distinct chromosomal rearrangements that juxtapose super-enhancers to the MYB locus. Chromosome conformation capture confirms that the translocated enhancers interact with the MYB promoter. Remarkably, MYB protein binds to the translocated enhancers, creating a positive feedback loop that sustains its expression. MYB also binds enhancers that drive different regulatory programs in alternate cell lineages in ACC, cooperating with TP63 in myoepithelial cells and a Notch program in luminal epithelial cells. Bromodomain inhibitors slow tumor growth in ACC primagraft models in vivo. Thus, our study identifies super-enhancer translocations that drive MYB expression and provides insight into downstream MYB functions in alternate ACC lineages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
Gene Expression Omnibus
References
Ohno, H. et al. Molecular analysis of a chromosomal translocation, t(9;14)(p13;q32), in a diffuse large-cell lymphoma cell line expressing the Ki-1 antigen. Proc. Natl. Acad. Sci. USA 87, 628–632 (1990).
Rabbitts, T.H. Chromosomal translocations in human cancer. Nature 372, 143–149 (1994).
Gröschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014).
Northcott, P.A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Tomlins, S.A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Adelstein, D.J., Koyfman, S.A., El-Naggar, A.K. & Hanna, E.Y. Biology and management of salivary gland cancers. Semin. Radiat. Oncol. 22, 245–253 (2012).
Ho, A.S. et al. The mutational landscape of adenoid cystic carcinoma. Nat. Genet. 45, 791–798 (2013).
Ramsay, R.G. & Gonda, T.J. MYB function in normal and cancer cells. Nat. Rev. Cancer 8, 523–534 (2008).
Persson, M. et al. Recurrent fusion of MYB and NFIB transcription factor genes in carcinomas of the breast and head and neck. Proc. Natl. Acad. Sci. USA 106, 18740–18744 (2009).
Mitani, Y. et al. Comprehensive analysis of the MYB-NFIB gene fusion in salivary adenoid cystic carcinoma: incidence, variability, and clinicopathologic significance. Clin. Cancer Res. 16, 4722–4731 (2010).
Stephens, P.J. et al. Whole exome sequencing of adenoid cystic carcinoma. J. Clin. Invest. 123, 2965–2968 (2013).
Moskaluk, C.A. et al. Development and characterization of xenograft model systems for adenoid cystic carcinoma. Lab. Invest. 91, 1480–1490 (2011).
Rivera, C.M. & Ren, B. Mapping human epigenomes. Cell 155, 39–55 (2013).
Queimado, L. et al. In vitro transformation of cell lines from human salivary gland tumors. Int. J. Cancer 81, 793–798 (1999).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Quintana, A.M., Liu, F., O'Rourke, J.P. & Ness, S.A. Identification and regulation of c-Myb target genes in MCF-7 cells. BMC Cancer 11, 30 (2011).
Zhao, L. et al. Integrated genome-wide chromatin occupancy and expression analyses identify key myeloid pro-differentiation transcription factors repressed by Myb. Nucleic Acids Res. 39, 4664–4679 (2011).
Mansour, M.R. et al. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Bell, D., Bell, A., Roberts, D., Weber, R.S. & El-Naggar, A.K. Developmental transcription factor EN1—a novel biomarker in human salivary gland adenoid cystic carcinoma. Cancer 118, 1288–1292 (2012).
Yamamoto, Y., Saka, T., Makimoto, K. & Takahashi, H. Histological changes during progression of adenoid cystic carcinoma. J. Laryngol. Otol. 106, 1016–1020 (1992).
Sato, K. et al. Adenoid cystic carcinoma of the maxillary sinus with gradual histologic transformation to high-grade adenocarcinoma: a comparative report with dedifferentiated carcinoma. Virchows Arch. 448, 204–208 (2006).
Barbareschi, M. et al. p63, a p53 homologue, is a selective nuclear marker of myoepithelial cells of the human breast. Am. J. Surg. Pathol. 25, 1054–1060 (2001).
Nguyen, B.C. et al. Cross-regulation between Notch and p63 in keratinocyte commitment to differentiation. Genes Dev. 20, 1028–1042 (2006).
Yalcin-Ozuysal, O. et al. Antagonistic roles of Notch and p63 in controlling mammary epithelial cell fates. Cell Death Differ. 17, 1600–1612 (2010).
Stoeck, A. et al. Discovery of biomarkers predictive of GSI response in triple-negative breast cancer and adenoid cystic carcinoma. Cancer Discov. 4, 1154–1167 (2014).
Robinson, D.R. et al. Functionally recurrent rearrangements of the MAST kinase and Notch gene families in breast cancer. Nat. Med. 17, 1646–1651 (2011).
Haydu, J.E. et al. An activating intragenic deletion in NOTCH1 in human T-ALL. Blood 119, 5211–5214 (2012).
Moll, U.M. & Slade, N. p63 and p73: roles in development and tumor formation. Mol. Cancer Res. 2, 371–386 (2004).
Mitani, Y. et al. Expression and regulation of the ΔN and TAp63 isoforms in salivary gland tumorigenesis: clinical and experimental findings. Am. J. Pathol. 179, 391–399 (2011).
Roe, J.S., Mercan, F., Rivera, K., Pappin, D.J. & Vakoc, C.R. BET bromodomain inhibition suppresses the function of hematopoietic transcription factors in acute myeloid leukemia. Mol. Cell 58, 1028–1039 (2015).
Knoechel, B. et al. An epigenetic mechanism of resistance to targeted therapy in T cell acute lymphoblastic leukemia. Nat. Genet. 46, 364–370 (2014).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Nicolaides, N.C., Gualdi, R., Casadevall, C., Manzella, L. & Calabretta, B. Positive autoregulation of c-myb expression via Myb binding sites in the 5′ flanking region of the human c-myb gene. Mol. Cell. Biol. 11, 6166–6176 (1991).
Nomura, T. et al. Negative autoregulation of c-Myb activity by homodimer formation through the leucine zipper. J. Biol. Chem. 268, 21914–21923 (1993).
Brill, L.B. II et al. Analysis of MYB expression and MYB-NFIB gene fusions in adenoid cystic carcinoma and other salivary neoplasms. Mod. Pathol. 24, 1169–1176 (2011).
Bell, D. et al. Clinical significance of Myb protein and downstream target genes in salivary adenoid cystic carcinoma. Cancer Biol. Ther. 12, 569–573 (2011).
Berger, M.F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Drier, Y. et al. Somatic rearrangements across cancer reveal classes of samples with distinct patterns of DNA breakage and rearrangement-induced hypermutability. Genome Res. 23, 228–235 (2013).
MacDonald, J.R., Ziman, R., Yuen, R.K., Feuk, L. & Scherer, S.W. The Database of Genomic Variants: a curated collection of structural variation in the human genome. Nucleic Acids Res. 42, D986–D992 (2014).
Ryan, R.J. et al. Detection of enhancer-associated rearrangements reveals mechanisms of oncogene dysregulation in B-cell lymphoma. Cancer Discov. 5, 1058–1071 (2015).
Thorvaldsdóttir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet. 4, e1000242 (2008).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Suvà, M.L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014).
Riggi, N. et al. EWS-FLI1 utilizes divergent chromatin remodeling mechanisms to directly activate or repress enhancer elements in Ewing sarcoma. Cancer Cell 26, 668–681 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Milacic, M. et al. Annotating cancer variants and anti-cancer therapeutics in Reactome. Cancers (Basel) 4, 1180–1211 (2012).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012).
Acknowledgements
We thank M. Rivera, N. Riggi, S. Puram, P. van Galen, J. Lohr and J. Kaufman for helpful discussions and critical comments on the manuscript; J. Voisine, R. Isenhart, R. Issner, H. Whitton, A. Spooner, M. Uziel, C. Epstein and N. Shoresh for technical assistance; T. Chan and V. Makarov for help with whole-genome sequencing data access; and the Salivary Gland Tumor Biorepository for providing the primary tumors (National Institute of Dental and Craniofacial Research (NIDCR) award reference HHSN268200900039C 04). This work was supported by the Adenoid Cystic Carcinoma Research Foundation (B.E.B. and B.K.), the Temares Family Foundation and the Howard Hughes Medical Institute. B.E.B. is an American Cancer Society Research Professor.
Author information
Authors and Affiliations
Contributions
B.K. and Y.D. designed and performed experiments and analyzed the data. B.K. and B.E.B. designed the experimental strategy and supervised the study and analysis. Y.D. carried out computational analyses. Y.D., B.K. and B.E.B. wrote the manuscript. J.C.A., M.J.C., K.E.W., S.M.G., C.D.C., S.J.R., L.M.S. and M.J.W. contributed to experiments and data analysis. A.H.A., R.J.H.R., M.J.K., W.C.F., L.Q., J.Q., J.E.B., C.A.M., A.K.E.-N. and J.E.B. provided reagents, contributed to analysis and gave conceptual advice. All authors discussed the results and implications and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.E.B. is a scientific founder of Tensha Therapeutics, which has licensed drug-like derivatives of the JQ1 bromodomain inhibitor from the Dana-Farber Cancer Institute. The remaining authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 PCR validation of rearrangements in four ACC primagrafts.
PCR results across genomic breakpoints (Table 1 and Supplementary Table 1) of high-confidence rearrangements (HC), MYB rearrangements and low-confidence rearrangements (LC) are shown for four ACC primagrafts. Control (C) refers to the same primer used in a different sample—ACC X2 for ACC X9 and ACC X9 for all others.
Supplementary Figure 2 ACC primagrafts and primary ACCs have very similar enhancer profiles.
Spearman correlation between enhancer (H3K27ac) maps of primary ACCs, ACC primagrafts, an HPV-transformed ACC cell line (ACC112), and other normal and malignant tissues and cell lines, based on H3K27ac signal 2 kb outside of the TSS.
Supplementary Figure 3 Several of the translocated enhancers are super-enhancers.
Super-enhancer ‘hockey stick’ plots, showing normalized H3K27ac (top) or BRD4 (bottom) signals across all enhancers genome-wide in ACC primagrafts. Enhancers on the right of the infliction point are considered ‘super-enhancers’. The enhancers translocated near MYB are shown as red, purple or blue dots (color matches the type of MYB rearrangement: MYB-NFIB fusion with 3′ UTR loss in red, MYB-NFIB translocations retaining the 3′ UTR in purple and MYB-TGFBR3 translocation retaining the 3′ UTR in blue).
Supplementary Figure 4 RAD51B locus contains many putative active enhancers.
H3K27ac tracks are shown for the RAD51B locus in five primary tumors and eight primagrafts. H3K27ac signal is shown in fragments per million (fpm).
Supplementary Figure 5 NFIB, TGFBR3 and RAD51B are highly expressed in normal salivary gland.
(a) NFIB and TGFBR3 mRNA are highly expressed in normal salivary gland, as detected by qPCR, normalized to GAPDH expression. OCI-LY-3 (lymphoma cell line) serves as negative control. Error bars, s.e.m. (b) NFIB, TGFBR3 and RAD51B are highly expressed in normal salivary gland, according to RNA-seq data from the Human Protein Atlas20.
Supplementary Figure 6 Translocated MYB-bound enhancers activate transcription.
Putative enhancer sequences of the NFIB enhancers (En3, En6 or En7) or TGFBR3 enhancers (Et3, Et6) or matching controls with the same sequence but scrambled MYB motifs (replacing CNGTT with GTAAG) were cloned upstream of luciferase and a minimal promoter (sequences are listed in Supplementary Table 6). Constructs were delivered by nucleofection into Jurkat cells (Online Methods). Firefly luciferase activity was measured after 36 h and normalized to Renilla luciferase to control for cell number and transfection efficiency. Error bars, s.e.m. for six technical repeats. Four of the five enhancer elements strongly increased activity in the luciferase assay. Two of the NFIB elements (En6 and En7) showed significantly reduced activity with scrambled MYB motifs. Significance was estimated by one-tailed t test. These data support our model that translocated enhancers drive transcription in a MYB-dependent manner.
Supplementary Figure 7 ACCs express the ΔNp63 but not TAp63 isoform of TP63.
(a) H3K27ac, H3K4me3 and MYB tracks are shown for the TP63 locus. MYB binds several intragenic TP63 enhancers in grade 2 primagrafts (X16, X5M1, X2). Only the promoter of the short isoform ΔNp63 is active. Signals are shown in fpm. (b) TP63 is expressed in grade 2 tumors (X2, X5M1) but not in grade 3 tumors (X9, X11). TAp63 is not detected in ACC primagrafts; hence only ΔNp63 is expressed. OCI-LY-3 (lymphoma cell line) serves as a positive control. Error bars, s.e.m. (n = 3).
Supplementary Figure 8 Colocalization of TP63 and MYB in grade 2 ACC.
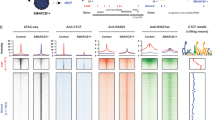
Heat maps of MYB and TP63 binding in 1-kb windows around TP63 peaks, sorted by overall signal strength. 81% of TP63 peaks are co-bound by MYB. Colors are scaled to show a maximum of 3 fpm per 10-bp bin.
Supplementary Figure 9 TP63, MYB and intercellular NOTCH1 expression in grade 2 and grade 3 ACCs.
Representative IHC staining for TP63, MYB and the intracellular domain of NOTCH1 (ICN1) in grade 2 ACCs (top three rows) and grade 3 ACCs (bottom four rows). 40× magnification is shown for ACCD3 and ACCD4; 40× (upper left corner) and 100× magnification is shown for ACCS1, ACCS4, ACCS5, ACCD1 and ACCD2. Scale bar, 100 µm.
Supplementary Figure 10 Heat map of H3K27ac showing similar patterns across primary tumors and primagrafts and distinct patterns between grade 2 and grade 3 tumors.
Heat map of H3K27ac at the enhancers of five primary tumors and eight primagrafts, showing four clusters of enhancers (unsupervised k-means) in a 5-kb window centered around H3K27ac peaks. Notice that the last cluster is preferentially active in grade 2 primagrafts and primary tumors. This cluster is highly enriched for the TP63 motif, consistent with the loss of TP63-expressing myoepithelial cells in grade 3 tumors. Colors are scaled to show a maximum of 2 fpm per 50-bp bin.
Supplementary Figure 11 Most MYB-binding sites are co-bound by BRD4 and show H3K27 acetylation.
Heat maps of H3K27ac, MYB and BRD4 binding in 5-kb windows around MYB peaks. The large majority of MYB peaks have both BRD4 co-binding and acetylation of H3K27. Colors are scaled to show a maximum of 10 fpm per 50-bp bin.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Table 5 (PDF 2148 kb)
Supplementary Table 1
Rearrangements detected in six ACC primagrafts from whole-genome sequencing. (XLSX 27 kb)
Supplementary Table 2
MYB high-confidence peaks and associated nearby genes. (XLSX 531 kb)
Supplementary Table 3
Enriched annotations of MYB targets. (XLSX 787 kb)
Supplementary Table 4
Transcriptional regulators targeted by MYB ranked by MYB binding, with expression levels in ACC and normal gland. (XLSX 67 kb)
Supplementary Table 6
Primer and reporter sequences used in this study. (XLSX 31 kb)
Rights and permissions
About this article
Cite this article
Drier, Y., Cotton, M., Williamson, K. et al. An oncogenic MYB feedback loop drives alternate cell fates in adenoid cystic carcinoma. Nat Genet 48, 265–272 (2016). https://doi.org/10.1038/ng.3502
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3502
This article is cited by
-
Super-enhancers complexes zoom in transcription in cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Pulmonary adenoid cystic carcinoma: molecular characteristics and literature review
Diagnostic Pathology (2023)
-
Increased retinoic acid signaling decreases lung metastasis in salivary adenoid cystic carcinoma by inhibiting the noncanonical Notch1 pathway
Experimental & Molecular Medicine (2023)
-
Functional profiles of curatively treated adenoid cystic carcinoma unveil prognostic features and potentially targetable pathways
Scientific Reports (2023)
-
Top 10 Basaloid Neoplasms of the Sinonasal Tract
Head and Neck Pathology (2023)