Abstract
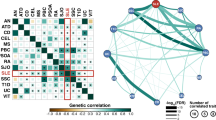
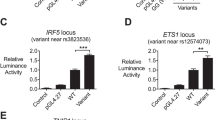
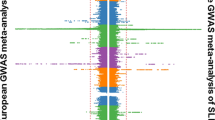
Systemic lupus erythematosus (SLE) has a strong but incompletely understood genetic architecture. We conducted an association study with replication in 4,478 SLE cases and 12,656 controls from six East Asian cohorts to identify new SLE susceptibility loci and better localize known loci. We identified ten new loci and confirmed 20 known loci with genome-wide significance. Among the new loci, the most significant locus was GTF2IRD1-GTF2I at 7q11.23 (rs73366469, Pmeta = 3.75 × 10−117, odds ratio (OR) = 2.38), followed by DEF6, IL12B, TCF7, TERT, CD226, PCNXL3, RASGRP1, SYNGR1 and SIGLEC6. We identified the most likely functional variants at each locus by analyzing epigenetic marks and gene expression data. Ten candidate variants are known to alter gene expression in cis or in trans. Enrichment analysis highlights the importance of these loci in B cell and T cell biology. The new loci, together with previously known loci, increase the explained heritability of SLE to 24%. The new loci share functional and ontological characteristics with previously reported loci and are possible drug targets for SLE therapeutics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Jakes, R.W. et al. Systematic review of the epidemiology of systemic lupus erythematosus in the Asia-Pacific region: prevalence, incidence, clinical features, and mortality. Arthritis Care Res. (Hoboken) 64, 159–168 (2012).
Danchenko, N., Satia, J.A. & Anthony, M.S. Epidemiology of systemic lupus erythematosus: a comparison of worldwide disease burden. Lupus 15, 308–318 (2006).
Wandstrat, A. & Wakeland, E. The genetics of complex autoimmune diseases: non-MHC susceptibility genes. Nat. Immunol. 2, 802–809 (2001).
Harley, I.T., Kaufman, K.M., Langefeld, C.D., Harley, J.B. & Kelly, J.A. Genetic susceptibility to SLE: new insights from fine mapping and genome-wide association studies. Nat. Rev. Genet. 10, 285–290 (2009).
Boackle, S.A. Advances in lupus genetics. Curr. Opin. Rheumatol. 25, 561–568 (2013).
Yang, W. et al. Meta-analysis followed by replication identifies loci in or near CDKN1B, TET3, CD80, DRAM1, and ARID5B as associated with systemic lupus erythematosus in Asians. Am. J. Hum. Genet. 92, 41–51 (2013).
So, H.C., Gui, A.H.S., Cherny, S.S. & Sham, P.C. Evaluating the heritability explained by known susceptibility variants: a survey of ten complex diseases. Genet. Epidemiol. 35, 310–317 (2011).
Gateva, V. et al. A large-scale replication study identifies TNIP1, PRDM1, JAZF1, UHRF1BP1 and IL10 as risk loci for systemic lupus erythematosus. Nat. Genet. 41, 1228–1233 (2009).
Cortes, A. & Brown, M.A. Promise and pitfalls of the Immunochip. Arthritis Res. Ther. 13, 101 (2011).
Wellcome Trust Case Control Consortium. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Farh, K.K. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Westra, H.J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Tantin, D., Tussie-Luna, M.I., Roy, A.L. & Sharp, P.A. Regulation of immunoglobulin promoter activity by TFII-I class transcription factors. J. Biol. Chem. 279, 5460–5469 (2004).
Li, Y. et al. A genome-wide association study in Han Chinese identifies a susceptibility locus for primary Sjögren's syndrome at 7q11.23. Nat. Genet. 45, 1361–1365 (2013).
Zheng, J. et al. The GTF2I rs117026326 polymorphism is associated with anti-SSA-positive primary Sjögren's syndrome. Rheumatology (Oxford) 54, 562–564 (2015).
Lessard, C.J. et al. Variants at multiple loci implicated in both innate and adaptive immune responses are associated with Sjögren's syndrome. Nat. Genet. 45, 1284–1292 (2013).
Perl, A. Emerging new pathways of pathogenesis and targets for treatment in systemic lupus erythematosus and Sjogren's syndrome. Curr. Opin. Rheumatol. 21, 443–447 (2009).
Johnatty, S.E. et al. Evaluation of candidate stromal epithelial cross-talk genes identifies association between risk of serous ovarian cancer and TERT, a cancer susceptibility “hot-spot”. PLoS Genet. 6, e1001016 (2010).
Berndt, S.I. et al. Genome-wide association study identifies multiple risk loci for chronic lymphocytic leukemia. Nat. Genet. 45, 868–876 (2013).
Kim, K. et al. High-density genotyping of immune loci in Koreans and Europeans identifies eight new rheumatoid arthritis risk loci. Ann. Rheum. Dis. 74, e13 (2015).
Kim, K. et al. The HLA-DRβ1 amino acid positions 11-13-26 explain the majority of SLE-MHC associations. Nat. Commun. 5, 5902 (2014).
Lessard, C.J. et al. Identification of IRF8, TMEM39A, and IKZF3-ZPBP2 as susceptibility loci for systemic lupus erythematosus in a large-scale multiracial replication study. Am. J. Hum. Genet. 90, 648–660 (2012).
Chu, Q., Liu, L. & Wang, W. Overexpression of hCLP46 enhances Notch activation and regulates cell proliferation in a cell type–dependent manner. Cell Prolif. 46, 254–262 (2013).
Chen, E.Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Xiong, W. & Lahita, R.G. Pragmatic approaches to therapy for systemic lupus erythematosus. Nat. Rev. Rheumatol. 10, 97–107 (2014).
Trost, B., Arsenault, R., Griebel, P., Napper, S. & Kusalik, A. DAPPLE: a pipeline for the homology-based prediction of phosphorylation sites. Bioinformatics 29, 1693–1695 (2013).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 41, D793–D800 (2013).
Wren, J.D., Bekeredjian, R., Stewart, J.A., Shohet, R.V. & Garner, H.R. Knowledge discovery by automated identification and ranking of implicit relationships. Bioinformatics 20, 389–398 (2004).
Hu, X. et al. Integrating autoimmune risk loci with gene-expression data identifies specific pathogenic immune cell subsets. Am. J. Hum. Genet. 89, 496–506 (2011).
Molineros, J.E. et al. Admixture mapping in lupus identifies multiple functional variants within IFIH1 associated with apoptosis, inflammation, and autoantibody production. PLoS Genet. 9, e1003222 (2013).
Maiti, A.K. et al. Combined protein- and nucleic acid–level effects of rs1143679 (R77H), a lupus-predisposing variant within ITGAM. Hum. Mol. Genet. 23, 4161–4176 (2014).
Guthridge, J.M. et al. Two functional lupus-associated BLK promoter variants control cell-type- and developmental-stage-specific transcription. Am. J. Hum. Genet. 94, 586–598 (2014).
Vandeweyer, G., Van der Aa, N., Reyniers, E. & Kooy, R.F. The contribution of CLIP2 haploinsufficiency to the clinical manifestations of the Williams-Beuren syndrome. Am. J. Hum. Genet. 90, 1071–1078 (2012).
Howard, M.L. et al. Mutation of Gtf2ird1 from the Williams-Beuren syndrome critical region results in facial dysplasia, motor dysfunction, and altered vocalisations. Neurobiol. Dis. 45, 913–922 (2012).
Antonell, A. et al. Partial 7q11.23 deletions further implicate GTF2I and GTF2IRD1 as the main genes responsible for the Williams-Beuren syndrome neurocognitive profile. J. Med. Genet. 47, 312–320 (2010).
Roy, A.L. Biochemistry and biology of the inducible multifunctional transcription factor TFII-I: 10 years later. Gene 492, 32–41 (2012).
Malcolm, T., Kam, J., Pour, P.S. & Sadowski, I. Specific interaction of TFII-I with an upstream element on the HIV-1 LTR regulates induction of latent provirus. FEBS Lett. 582, 3903–3908 (2008).
Gupta, S. et al. T cell receptor engagement leads to the recruitment of IBP, a novel guanine nucleotide exchange factor, to the immunological synapse. J. Biol. Chem. 278, 43541–43549 (2003).
Biswas, P.S. et al. Dual regulation of IRF4 function in T and B cells is required for the coordination of T-B cell interactions and the prevention of autoimmunity. J. Exp. Med. 209, 581–596 (2012).
Noble, J.A. et al. A polymorphism in the TCF7 gene, C883A, is associated with type 1 diabetes. Diabetes 52, 1579–1582 (2003).
Klapper, W. et al. Telomerase activity in B and T lymphocytes of patients with systemic lupus erythematosus. Ann. Rheum. Dis. 63, 1681–1683 (2004).
Iguchi-Manaka, A. et al. Accelerated tumor growth in mice deficient in DNAM-1 receptor. J. Exp. Med. 205, 2959–2964 (2008).
Alcina, A. et al. The autoimmune disease–associated KIF5A, CD226 and SH2B3 gene variants confer susceptibility for multiple sclerosis. Genes Immun. 11, 439–445 (2010).
Deshmukh, H.A. et al. Evaluation of 19 autoimmune disease–associated loci with rheumatoid arthritis in a Colombian population: evidence for replication and gene-gene interaction. J. Rheumatol. 38, 1866–1870 (2011).
Hafler, J.P. et al. CD226 Gly307Ser association with multiple autoimmune diseases. Genes Immun. 10, 5–10 (2009).
Maiti, A.K. et al. Non-synonymous variant (Gly307Ser) in CD226 is associated with susceptibility to multiple autoimmune diseases. Rheumatology (Oxford) 49, 1239–1244 (2010).
Qiu, Z.X., Zhang, K., Qiu, X.S., Zhou, M. & Li, W.M. CD226 Gly307Ser association with multiple autoimmune diseases: a meta-analysis. Hum. Immunol. 74, 249–255 (2013).
Wieczorek, S. et al. Novel association of the CD226 (DNAM-1) Gly307Ser polymorphism in Wegener's granulomatosis and confirmation for multiple sclerosis in German patients. Genes Immun. 10, 591–595 (2009).
Du, Y. et al. Association of the CD226 single nucleotide polymorphism with systemic lupus erythematosus in the Chinese Han population. Tissue Antigens 77, 65–67 (2011).
Stoeckman, A.K. et al. A distinct inflammatory gene expression profile in patients with psoriatic arthritis. Genes Immun. 7, 583–591 (2006).
Yasuda, S. et al. Defective expression of Ras guanyl nucleotide–releasing protein 1 in a subset of patients with systemic lupus erythematosus. J. Immunol. 179, 4890–4900 (2007).
He, C.F. et al. TNIP1, SLC15A4, ETS1, RasGRP3 and IKZF1 are associated with clinical features of systemic lupus erythematosus in a Chinese Han population. Lupus 19, 1181–1186 (2010).
Iatropoulos, P. et al. Association study and mutational screening of SYNGR1 as a candidate susceptibility gene for schizophrenia. Psychiatr. Genet. 19, 237–243 (2009).
Liu, J.Z. et al. Dense fine-mapping study identifies new susceptibility loci for primary biliary cirrhosis. Nat. Genet. 44, 1137–1141 (2012).
Gorski, K.S. et al. A set of genes selectively expressed in murine dendritic cells: utility of related cis-acting sequences for lentiviral gene transfer. Mol. Immunol. 40, 35–47 (2003).
Patel, N. et al. OB-BP1/Siglec-6. A leptin- and sialic acid–binding protein of the immunoglobulin superfamily. J. Biol. Chem. 274, 22729–22738 (1999).
Okada, Y. et al. A genome-wide association study identified AFF1 as a susceptibility locus for systemic lupus eyrthematosus in Japanese. PLoS Genet. 8, e1002455 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Liu, E.Y., Li, M., Wang, W. & Li, Y. MaCH-admix: genotype imputation for admixed populations. Genet. Epidemiol. 37, 25–37 (2013).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Liu, H. et al. Discovery of six new susceptibility loci and analysis of pleiotropic effects in leprosy. Nat. Genet. 47, 267–271 (2015).
Verma, S.S. et al. Imputation and quality control steps for combining multiple genome-wide datasets. Front. Genet. 5, 370 (2014).
Li, Y., Willer, C., Sanna, S. & Abecasis, G. Genotype imputation. Annu. Rev. Genomics Hum. Genet. 10, 387–406 (2009).
Li, Y. & Abecasis, G.R. Mach. 1.0: rapid haplotype reconstruction and missing genotype inference. Am. J. Hum. Genet. S79, 2290 (2006).
Song, M., Hao, W. & Storey, J.D. Testing for genetic associations in arbitrarily structured populations. Nat. Genet. 47, 550–554 (2015).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Flicek, P. et al. Ensembl 2014. Nucleic Acids Res. 42, D749–D755 (2014).
Li, M.J., Wang, L.Y., Xia, Z., Sham, P.C. & Wang, J. GWAS3D: detecting human regulatory variants by integrative analysis of genome-wide associations, chromosome interactions and histone modifications. Nucleic Acids Res. 41, W150–W158 (2013).
Guo, L., Du, Y., Chang, S., Zhang, K. & Wang, J. rSNPBase: a database for curated regulatory SNPs. Nucleic Acids Res. 42, D1033–D1039 (2014).
Rouder, J.N. & Morey, R.D. Default Bayes factors for model selection in regression. Multivariate Behav. Res. 47, 877–903 (2012).
Wan, X. et al. BOOST: a fast approach to detecting gene-gene interactions in genome-wide case-control studies. Am. J. Hum. Genet. 87, 325–340 (2010).
Ueki, M. & Cordell, H.J. Improved statistics for genome-wide interaction analysis. PLoS Genet. 8, e1002625 (2012).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Slowikowski, K., Hu, X. & Raychaudhuri, S. SNPsea: an algorithm to identify cell types, tissues and pathways affected by risk loci. Bioinformatics 30, 2496–2497 (2014).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Hyatt, G. et al. Gene expression microarrays: glimpses of the immunological genome. Nat. Immunol. 7, 686–691 (2006).
FANTOM Consortium and the RIKEN PMI and CLST (DGT). A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Risch, N. & Merikangas, K. The future of genetic studies of complex human diseases. Science 273, 1516–1517 (1996).
International Consortium for Systemic Lupus Erythematosus Genetics (SLEGEN). et al. Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat. Genet. 40, 204–210 (2008).
Zheng, W. et al. Common genetic determinants of breast-cancer risk in East Asian women: a collaborative study of 23 637 breast cancer cases and 25 579 controls. Hum. Mol. Genet. 22, 2539–2550 (2013).
Hughes, T. et al. Analysis of autosomal genes reveals gene-sex interactions and higher total genetic risk in men with systemic lupus erythematosus. Ann. Rheum. Dis. 71, 694–699 (2012).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
DeLong, E.R., DeLong, D.M. & Clarke-Pearson, D.L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Pickrell, J.K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Acknowledgements
We are grateful to the affected and unaffected individuals who participated in this study. We thank the research assistants, coordinators and physicians who helped in the recruitment of subjects, including the individuals in the coordinating projects. A part of the Korean control data was provided from the Korean Biobank Project supported by the Korea Center for Disease Control and Prevention at the Korea National Institute of Health. Genomic DNA from ∼100 Korean patients with SLE was obtained from the Korean National Biobank at Wonkwang University Hospital, which is supported by the Ministry of Health and Welfare, Republic of Korea.
This work was supported by grants from the US National Institutes of Health (AR060366, MD007909, AI103399, AI024717, AI083194, AI107176, TR001425, HG008666 and HG006828), the US Department of Defense (PR094002), the US Department of Veterans Affairs, the National Basic Research Program of China (973 program) (2014CB541902), the Research Fund of Beijing Municipal Science and Technology for the Outstanding PhD Program (20121000110), the National Natural Science Foundation of China (81200524, 81230072) and High-Impact Research Ministry of Education Grant UM.C/625/1/HIR/MoE/E000044-20001, Malaysia. This study was also supported by a grant from the Korea Healthcare Technology R&D Project (HI13C2124), Ministry for Health and Welfare, Republic of Korea.
Author information
Authors and Affiliations
Contributions
S.K.N., J.B.H. and S.-C.B. conceived and initiated the study. S.K.N. designed, coordinated and supervised the overall study. C.S., X.Z., P.M., K.B., A.A. and X.K.-H. prepared samples, performed genotyping, cleaned the data, combined various data sets and maintained the database. C.S., J.E.M., K.K. and Y.O. performed data imputation, association analysis and various statistical analyses on the data. L.L.L., J.E.M., M.D. and J.D.W. performed the bioinformatic analysis. S.-C.B., H.Z., K.H.C., X.Z., K.K., S.-Y.B., H.-S.L., T.-H.K., Y.M.K., C.-H.S., W.T.C., Y.-B.P., J.-Y.C., S.C.S., S.-S.L., Y.J.K., B.-G.H., Y.K., A.S., M.K., T.S., K.Y., J.M., Y.Q., K.M.K. and N.S. recruited and characterized patients with SLE and controls and supplied the demographic and clinical data. C.S., J.E.M., X.K.-H., K.K., S.-C.B., L.L.L. and S.K.N. drafted the manuscript. All authors approved the study, reviewed the manuscript, commented and helped in revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16 and Supplementary Note. (PDF 11732 kb)
Supplementary Tables 1–29
Supplementary Tables 1–29. (XLSX 2075 kb)
Supplementary Data Set
Summary-level association data for the discovery sets. (XLSX 18647 kb)
Rights and permissions
About this article
Cite this article
Sun, C., Molineros, J., Looger, L. et al. High-density genotyping of immune-related loci identifies new SLE risk variants in individuals with Asian ancestry. Nat Genet 48, 323–330 (2016). https://doi.org/10.1038/ng.3496
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3496
This article is cited by
-
Molecular pathways identified from single nucleotide polymorphisms demonstrate mechanistic differences in systemic lupus erythematosus patients of Asian and European ancestry
Scientific Reports (2023)
-
Neuronal Gtf2i deletion alters mitochondrial and autophagic properties
Communications Biology (2023)
-
ChIP-seq analysis found IL21R, a target gene of GTF2I–the susceptibility gene for primary biliary cholangitis in Chinese Han
Hepatology International (2023)
-
Regulation of activated T cell survival in rheumatic autoimmune diseases
Nature Reviews Rheumatology (2022)
-
RasGRP1 promotes the acute inflammatory response and restricts inflammation-associated cancer cell growth
Nature Communications (2022)