Abstract
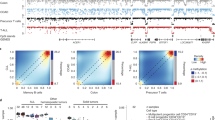
Charting differences between tumors and normal tissue is a mainstay of cancer research. However, clonal tumor expansion from complex normal tissue architectures potentially obscures cancer-specific events, including divergent epigenetic patterns. Using whole-genome bisulfite sequencing of normal B cell subsets, we observed broad epigenetic programming of selective transcription factor binding sites coincident with the degree of B cell maturation. By comparing normal B cells to malignant B cells from 268 patients with chronic lymphocytic leukemia (CLL), we showed that tumors derive largely from a continuum of maturation states reflected in normal developmental stages. Epigenetic maturation in CLL was associated with an indolent gene expression pattern and increasingly favorable clinical outcomes. We further uncovered that most previously reported tumor-specific methylation events are normally present in non-malignant B cells. Instead, we identified a potential pathogenic role for transcription factor dysregulation in CLL, where excess programming by EGR and NFAT with reduced EBF and AP-1 programming imbalances the normal B cell epigenetic program.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Ziller, M.J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013).
Smith, Z.D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013).
Lara-Astiaso, D. et al. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Cabezas-Wallscheid, N. et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell 15, 507–522 (2014).
Schübeler, D. Function and information content of DNA methylation. Nature 517, 321–326 (2015).
Timp, W. & Feinberg, A.P. Cancer as a dysregulated epigenome allowing cellular growth advantage at the expense of the host. Nat. Rev. Cancer 13, 497–510 (2013).
Kulis, M. et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat. Genet. 44, 1236–1242 (2012).
Queiros, A.C. et al. A B-cell epigenetic signature defines three biologic subgroups of chronic lymphocytic leukemia with clinical impact. Leukemia 29, 598–605 (2015).
Cahill, N. et al. 450K-array analysis of chronic lymphocytic leukemia cells reveals global DNA methylation to be relatively stable over time and similar in resting and proliferative compartments. Leukemia 27, 150–158 (2013).
Oakes, C.C. et al. Evolution of DNA methylation is linked to genetic aberrations in chronic lymphocytic leukemia. Cancer Discov. 4, 348–361 (2014).
Landau, D.A. et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell 26, 813–825 (2014).
Kulis, M. et al. Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat. Genet. 47, 746–756 (2015).
Lebecque, S., de Bouteiller, O., Arpin, C., Banchereau, J. & Liu, Y.J. Germinal center founder cells display propensity for apoptosis before onset of somatic mutation. J. Exp. Med. 185, 563–571 (1997).
Wang, Q. et al. Tagmentation-based whole-genome bisulfite sequencing. Nat. Protoc. 8, 2022–2032 (2013).
Shaknovich, R. et al. DNA methyltransferase 1 and DNA methylation patterning contribute to germinal center B-cell differentiation. Blood 118, 3559–3569 (2011).
Lai, A.Y. et al. DNA methylation profiling in human B cells reveals immune regulatory elements and epigenetic plasticity at Alu elements during B-cell activation. Genome Res. 23, 2030–2041 (2013).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Dvinge, H. et al. Sample processing obscures cancer-specific alterations in leukemic transcriptomes. Proc. Natl. Acad. Sci. USA 111, 16802–16807 (2014).
Florean, C., Schnekenburger, M., Grandjenette, C., Dicato, M. & Diederich, M. Epigenomics of leukemia: from mechanisms to therapeutic applications. Epigenomics 3, 581–609 (2011).
Teschendorff, A.E. et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 20, 440–446 (2010).
Smith, E.N. et al. Genetic and epigenetic profiling of CLL disease progression reveals limited somatic evolution and suggests a relationship to memory-cell development. Blood Cancer J. 5, e303 (2015).
Claus, R. et al. Validation of ZAP-70 methylation and its relative significance in predicting outcome in chronic lymphocytic leukemia. Blood 124, 42–48 (2014).
Zandi, S. et al. EBF1 is essential for B-lineage priming and establishment of a transcription factor network in common lymphoid progenitors. J. Immunol. 181, 3364–3372 (2008).
Heltemes-Harris, L.M. et al. Ebf1 or Pax5 haploinsufficiency synergizes with STAT5 activation to initiate acute lymphoblastic leukemia. J. Exp. Med. 208, 1135–1149 (2011).
Seifert, M. et al. Cellular origin and pathophysiology of chronic lymphocytic leukemia. J. Exp. Med. 209, 2183–2198 (2012).
Ferreira, P.G. et al. Transcriptome characterization by RNA sequencing identifies a major molecular and clinical subdivision in chronic lymphocytic leukemia. Genome Res. 24, 212–226 (2014).
Puente, X.S. et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475, 101–105 (2011).
Landau, D.A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
Herishanu, Y. et al. The lymph node microenvironment promotes B-cell receptor signaling, NF-κB activation, and tumor proliferation in chronic lymphocytic leukemia. Blood 117, 563–574 (2011).
Mockridge, C.I. et al. Reversible anergy of sIgM-mediated signaling in the two subsets of CLL defined by VH-gene mutational status. Blood 109, 4424–4431 (2007).
Reindl, L. et al. Biological and clinical characterization of recurrent 14q deletions in CLL and other mature B-cell neoplasms. Br. J. Haematol. 151, 25–36 (2010).
Edelmann, J. et al. High-resolution genomic profiling of chronic lymphocytic leukemia reveals new recurrent genomic alterations. Blood 120, 4783–4794 (2012).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Pham, T.H. et al. Dynamic epigenetic enhancer signatures reveal key transcription factors associated with monocytic differentiation states. Blood 119, e161–e171 (2012).
Damm, F. et al. Acquired initiating mutations in early hematopoietic cells of CLL patients. Cancer Discov. 4, 1088–1101 (2014).
Klein, U. et al. Gene expression profiling of B cell chronic lymphocytic leukemia reveals a homogeneous phenotype related to memory B cells. J. Exp. Med. 194, 1625–1638 (2001).
Klein, U., Rajewsky, K. & Küppers, R. Human immunoglobulin (Ig)M+IgD+ peripheral blood B cells expressing the CD27 cell surface antigen carry somatically mutated variable region genes: CD27 as a general marker for somatically mutated (memory) B cells. J. Exp. Med. 188, 1679–1689 (1998).
Chen, L. et al. ZAP-70 enhances IgM signaling independent of its kinase activity in chronic lymphocytic leukemia. Blood 111, 2685–2692 (2008).
Prange, K.H., Singh, A.A. & Martens, J.H. The genome-wide molecular signature of transcription factors in leukemia. Exp. Hematol. 42, 637–650 (2014).
Kretzmer, H. et al. DNA methylome analysis in Burkitt and follicular lymphomas identifies differentially methylated regions linked to somatic mutation and transcriptional control. Nat. Genet. 47, 1316–1325 (2015).
Zhuang, Y., Soriano, P. & Weintraub, H. The helix-loop-helix gene E2A is required for B cell formation. Cell 79, 875–884 (1994).
Gururajan, M. et al. Early growth response genes regulate B cell development, proliferation, and immune response. J. Immunol. 181, 4590–4602 (2008).
Peng, S.L., Gerth, A.J., Ranger, A.M. & Glimcher, L.H. NFATc1 and NFATc2 together control both T and B cell activation and differentiation. Immunity 14, 13–20 (2001).
Garrett-Sinha, L.A. et al. PU.1 and Spi-B are required for normal B cell receptor–mediated signal transduction. Immunity 10, 399–408 (1999).
Gustems, M. et al. c-Jun/c-Fos heterodimers regulate cellular genes via a newly identified class of methylated DNA sequence motifs. Nucleic Acids Res. 42, 3059–3072 (2014).
Rosén, A., Murray, F., Evaldsson, C. & Rosenquist, R. Antigens in chronic lymphocytic leukemia—implications for cell origin and leukemogenesis. Semin. Cancer Biol. 20, 400–409 (2010).
Pekarsky, Y. et al. Tcl1 functions as a transcriptional regulator and is directly involved in the pathogenesis of CLL. Proc. Natl. Acad. Sci. USA 105, 19643–19648 (2008).
Kröber, A. et al. VH mutation status, CD38 expression level, genomic aberrations, and survival in chronic lymphocytic leukemia. Blood 100, 1410–1416 (2002).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Bibikova, M. et al. Genome-wide DNA methylation profiling using Infinium® assay. Epigenomics 1, 177–200 (2009).
ENCODE Project Consortium. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640 (2004).
Teschendorff, A.E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Assenov, Y. et al. Comprehensive analysis of DNA methylation data with RnBeads. Nat. Methods 11, 1138–1140 (2014).
Brocks, D. et al. Epigenetic intratumor heterogeneity and clonal evolution in aggressive prostate cancer. Cell Rep. 8, 798–806 (2014).
Desper, R. & Gascuel, O. Fast and accurate phylogeny reconstruction algorithms based on the minimum-evolution principle. J. Comput. Biol. 9, 687–705 (2002).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Bohle, V., Döring, C., Hansmann, M.L. & Küppers, R. Role of early B-cell factor 1 (EBF1) in Hodgkin lymphoma. Leukemia 27, 671–679 (2013).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Acknowledgements
We would like to thank the Genomics and Proteomics Core Facility at the German Cancer Research Center, in particular R. Fisher and M. Schick for their excellent technical support and expertise. We thank Imaging Center Essen (IMCES) for support in B cell sorting. We are grateful to M. Bähr, O. Mücke, M. Helf and S. Ohl for technical support. We also thank J. Edelmann, M. Seiffert, L. Sellner, B. Wu, V. Hovestadt, A. Kundaje and J.I. Martín-Subero for providing samples, data and/or analytical tools. S.S. is supported by the Else Kröner Fresenius Stiftung (2012_A146), the Virtual Helmholtz Institute (VH-VI-404) and the Deutsche Forschungsgemeinschaft (SFB 1074 projects B1 and B2). This work was supported in part by the Helmholtz Association, from the DKFZ–Heidelberg Center for Personalized Oncology (DKFZ-HIPO), the German Cancer Consortium (DKTK), the CLL Research Consortium (CRC), the German Federal Ministry of Education and Research CancerEpiSys network (BMBF 031 6049C), the Virtual Helmholtz Institute (VH-VI-404), the Deutsche Forschungsgemeinschaft (GKR1431 and SE1885/2-1), the Leukemia and Lymphoma Society (P01 CA081534), the Four Winds Foundation, the European Union's Seventh Framework Programme through the Blueprint Consortium, the German Ministry of Education and Research (BMBF) through the ICGC MMML-Seq Project (01KU1002A-J) and the US National Institutes of Health (PO1-CA81534).
Author information
Authors and Affiliations
Contributions
C.C.O., M.S., M.P., A.S., D.W. and C.P. designed and performed experimental work. C.C.O., Y.A., L.G., A.S.R., Q.W., C.D.I., S.D.K., D.B., D.B.L. and O.B. performed data analysis. M.S., L.R., T.J.K., H.D., R.K., T.Z., S.S. and J.C.B. provided clinical samples or data. C.C.O., M.S. and C.P. prepared the manuscript and figures. B.B., D.M., M.Z., P.L., T.Z., S.S., J.C.B. and C.P. provided project leadership. All authors contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Note and Supplementary Tables 2–11. (PDF 17289 kb)
Supplementary Table 1
DNA methylation levels of the top 5,000 hypermethylated windows. (XLSX 1089 kb)
Supplementary Table 12
TWGBS windows overlapping a TFBS with the corresponding motif. (XLSX 2403 kb)
Supplementary Table 13
450K probes overlapping a TFBS with the corresponding motif. (XLSX 74 kb)
Rights and permissions
About this article
Cite this article
Oakes, C., Seifert, M., Assenov, Y. et al. DNA methylation dynamics during B cell maturation underlie a continuum of disease phenotypes in chronic lymphocytic leukemia. Nat Genet 48, 253–264 (2016). https://doi.org/10.1038/ng.3488
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3488
This article is cited by
-
The importance of personalized medicine in chronic myeloid leukemia management: a narrative review
Egyptian Journal of Medical Human Genetics (2023)
-
Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Nature Communications (2023)
-
A leukemia-protective germline variant mediates chromatin module formation via transcription factor nucleation
Nature Communications (2022)
-
A pan-cancer analysis of the oncogenic role of dual-specificity tyrosine (Y)-phosphorylation- regulated kinase 2 (DYRK2) in human tumors
Scientific Reports (2022)
-
Molecular map of chronic lymphocytic leukemia and its impact on outcome
Nature Genetics (2022)