Abstract
We performed fine mapping of 39 established type 2 diabetes (T2D) loci in 27,206 cases and 57,574 controls of European ancestry. We identified 49 distinct association signals at these loci, including five mapping in or near KCNQ1. 'Credible sets' of the variants most likely to drive each distinct signal mapped predominantly to noncoding sequence, implying that association with T2D is mediated through gene regulation. Credible set variants were enriched for overlap with FOXA2 chromatin immunoprecipitation binding sites in human islet and liver cells, including at MTNR1B, where fine mapping implicated rs10830963 as driving T2D association. We confirmed that the T2D risk allele for this SNP increases FOXA2-bound enhancer activity in islet- and liver-derived cells. We observed allele-specific differences in NEUROD1 binding in islet-derived cells, consistent with evidence that the T2D risk allele increases islet MTNR1B expression. Our study demonstrates how integration of genetic and genomic information can define molecular mechanisms through which variants underlying association signals exert their effects on disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others

References
Kooner, J.S. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011).
Cho, Y.S. et al. Meta-analysis of genome-wide association studies identifies eight new loci for type 2 diabetes in East Asians. Nat. Genet. 44, 67–72 (2012).
Voight, B.F. et al. Twelve type 2 diabetes susceptibility loci identified through large scale association analysis. Nat. Genet. 42, 579–589 (2010).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Mahajan, A. et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat. Genet. 46, 234–244 (2014).
Altshuler, D. et al. The common PPARγ Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat. Genet. 26, 76–80 (2000).
Gloyn, A.L. et al. Large-scale association studies of variants in genes encoding the pancreatic-cell KATP channel subunits Kir6.2 (KCNJ11) and SUR1 (ABCC8) conrm that the KCNJ11 E23K variant is associated with type 2 diabetes. Diabetes 52, 568–572 (2003).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes. Nat. Genet. 42, 105–116 (2010).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Elbein, S.C. et al. Genetic risk factors for type 2 diabetes: a trans-regulatory genetic architecture? Am. J. Hum. Genet. 91, 466–477 (2012).
Trynka, G. et al. Chromatin marks identify critical cell types for fine-mapping complex trait variants. Nat. Genet. 45, 124–130 (2013).
Parker, S.C.J. et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbour human disease risk variants. Proc. Natl. Acad. Sci. USA 110, 17921–17926 (2013).
Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat. Genet. 46, 136–143 (2014).
Voight, B.F. et al. The Metabochip, a custom genotyping array for genetic studies of metabolic, cardiovascular, and anthropometric traits. PLoS Genet. 8, e1002793 (2012).
International HapMap Project Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Winkler, T.W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Unoki, H. et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 40, 1098–1102 (2008).
Fitzpatrick, G.V., Soloway, P.D. & Higgins, M.J. Regional loss of imprinting and growth deficiency in mice with a targeted deletion of KvDMR1. Nat. Genet. 32, 426–431 (2002).
Zeggini, E. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316, 1336–1341 (2007).
Shea, J. et al. Comparing strategies to fine-map the association of common SNPs at chromosome 9p21 with type 2 diabetes and myocardial infarction. Nat. Genet. 43, 801–805 (2011).
Maller, J.B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Jafar-Mohammadi, B. et al. A role for coding functional variants in HNF4A in type 2 diabetes susceptibility. Diabetologia 54, 111–119 (2011).
Teslovich, T.M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Florez, J.C. et al. Haplotype structure and genotype-phenotype correlations of the sulfonylurea receptor and the islet ATP-sensitive potassium channel gene. Diabetes 53, 1360–1368 (2004).
Hamming, K.S. et al. Co-expression of the type 2 diabetes susceptibility gene variants KCNJ11 E23K and ABCC8 S1369A alter the ATP and sulfonylurea sensitivities of the ATP-sensitive K+ channel. Diabetes 58, 2419–2424 (2009).
Nicolson, T.J. et al. Insulin storage and glucose homeostasis in mice null for the granule zinc transporter ZnT8 and studies of the type 2 diabetes–associated variants. Diabetes 58, 2070–2083 (2009).
Beer, N.L. et al. The P446L variant in GCKR associated with fasting plasma glucose and triglyceride levels exerts its effect through increased glucokinase activity in liver. Hum. Mol. Genet. 18, 4081–4088 (2009).
Holmkvist, J. et al. Common variants in HNF-1α and risk of type 2 diabetes. Diabetologia 49, 2882–2891 (2006).
Yamagata, K. et al. Mutations in the hepatocyte nuclear factor-1α gene in maturity-onset diabetes of the young (MODY3). Nature 384, 455–458 (1996).
Yamagata, K. et al. Mutations in the hepatocyte nuclear factor-4α gene in maturity-onset diabetes of the young (MODY1). Nature 384, 458–460 (1996).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Soccio, R.E. et al. Species-specific strategies underlying conserved functions of metabolic transcription factors. Mol. Endocrinol. 25, 694–706 (2011).
Gaulton, K.J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet. 42, 255–259 (2010).
Fogarty, M.P., Cannon, M.E., Vadlamudi, S., Gaulton, K.J. & Mohlke, K.L. Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS Genet. 10, e1004633 (2014).
Manning, A.K. et al. A genome-wide approach accounting for body-mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat. Genet. 44, 659–669 (2012).
Dimas, A.S. et al. Impact of type 2 diabetes susceptibility variants on quantitative glycemic traits reveals mechanistic heterogeneity. Diabetes 63, 2158–2171 (2014).
Ravassard, P. et al. A genetically engineered human pancreatic β cell line exhibiting glucose-inducible insulin secretion. J. Clin. Invest. 121, 3589–3597 (2011).
Fadista, J. et al. Global genomic and transcriptomic analysis of human pancreatic islets reveals novel genes influencing glucose metabolism. Proc. Natl. Acad. Sci. USA 111, 13924–13929 (2014).
Lyssenko, V. et al. Common variant in MTNR1B associated with increased risk of type 2 diabetes and impaired early insulin secretion. Nat. Genet. 41, 82–88 (2009).
Gao, N. et al. Foxa1 and Foxa2 maintain the metabolic and secretory features of the mature β-cell. Mol. Endocrinol. 24, 1594–1604 (2010).
Zhou, Y. et al. TCF7L2 is a master regulator of insulin production and processing. Hum. Mol. Genet. 23, 6419–6431 (2014).
Lyssenko, V. et al. Mechanisms by which common variants in the TCF7L2 gene increase risk of type 2 diabetes. J. Clin. Invest. 117, 2155–2163 (2007).
Gloyn, A.L. et al. Activating mutations in the gene encoding the ATP-sensitive potassium-channel subunit Kir6.2 and permanent neonatal diabetes. N. Engl. J. Med. 350, 1838–1849 (2004).
Flannick, J. et al. Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nat. Genet. 46, 357–363 (2014).
Dickson, S.P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D.B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010).
Bonnefond, A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat. Genet. 44, 297–301 (2012).
Zaret, K.S. & Carroll, J.S. Pioneer transcription factors: establishing a competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Gao, N. et al. Dynamic regulation of Pdx1 enhancers by Foxa1 and Foxa2 is essential for pancreas development. Genes Dev. 22, 3435–3448 (2008).
Lee, C.S., Friedman, J.R., Fulmer, J.T. & Kaestner, K.H. The initiation of liver development is dependent on Foxa transcription factors. Nature 435, 944–947 (2005).
Scott, R.A. et al. Large-scale association analyses identify new loci influencing glycaemic traits and provide insight into the underlying biological pathways. Nat. Genet. 44, 991–1005 (2012).
Tabassum, R., Chavali, S., Dwivedi, O.P., Tandon, N. & Bharadwaj, D. Genetic variants of FOXA2: risk of type 2 diabetes and effect on metabolic traits in North Indians. J. Hum. Genet. 53, 957–965 (2008).
Johnson, M.E., Schug, J., Wells, A.D., Kaestner, K.H. & Grant, S.F. Genome-wide analyses of ChIP-Seq derived FOXA2 DNA occupancy in liver points to genetic networks underpinning multiple complex traits. J. Clin. Endocrinol. Metab. 99, E1580–E1585 (2014).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Wakefield, J. Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kharchenko, P.V., Tolstorukov, M.Y. & Park, P.J. Design and analysis of ChIP-seq experiments for DNA-binding proteins. Nat. Biotechnol. 26, 1351–1359 (2008).
Li, Q., Brown, J.B., Huang, H. & Bickel, P.J. Measuring reproducibility of high-throughput experiments. Ann. App. Stat. 5, 1752–1779 (2011).
Mikkelsen, T.S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169 (2010).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825 (2010).
Morán, I. et al. Human β cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 16, 435–448 (2012).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Machanick, P. & Bailey, T.L. MEME-ChiP. Motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Mathelier, A. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B-cell identifies. Mol. Cell 38, 576–589 (2010).
Bailey, T.L. et al. MEME Suite: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Grant, C.E., Bailey, T.L. & Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Pugh, C.W., Tan, C.C., Jones, R.W. & Ratcliffe, P.J. Functional analysis of an oxygen-regulated transcriptional enhancer lying 3′ to the mouse erythropoietin gene. Proc. Natl. Acad. Sci. USA 88, 10553–10557 (1991).
Acknowledgements
Funding for the research undertaken in this study has been received from the Academy of Finland (including grants 77299, 102318, 10493, 118065, 123885, 124243, 129293, 129680, 136895, 139635, 211119, 213506, 251217 and 263836); Agence National de la Recherche; Association de Langue Française pour l'Etude du Diabète et des Maladies Métaboliques; Association Diabèete Risque Vasculaire; Association Française des Diabétiques; the Association of Danish Pharmacies; the Augustinus Foundation; the Becket Foundation; the British Diabetes Association (BDA) Research; the British Heart Foundation; the Central Norway Health Authority; the Central Finland Hospital District; the Center for Inherited Disease Research (CIDR); the City of Kuopio; the City of Leutkirch; Copenhagen County; the Danish Centre for Evaluation and Health Technology Assessment; the Danish Council for Independent Research; the Danish Heart Foundation; the Danish Research Councils; Deutsche Forschungsgemeinschaft (including project ER 155/6-2); the Diabetes Research Foundation; Diabetes UK; the Doris Duke Charitable Foundation; Erasmus Medical Center; Erasmus University; the Estonian government (SF0180142s08); the European Commission (including ENGAGE HEALTH-F4-2007-201413, FP7-201413, FP7-245536, EXGENESIS LSHM-CT-2004-005272, FP6 LSHM_CT_2006_037197, LSHM-CT-2007-037273, Directorate C-Public Health 2004310, DG XII); the European Regional Development Fund; the Federal Ministry of Education and Research, Germany (including FKZ 01GI1128 and FKZ 01EO1001); the Federal Ministry of Health, Germany; the Finnish Diabetes Association; the Finnish Diabetes Research Foundation; the Finnish Foundation for Cardiovascular Research; the Finnish Medical Society; the Folkhalsan Research Foundation; the Foundation for Life and Health in Finland; the Foundation for Old Servants; the Fredrick och Ingrid Thuring Foundation; the French region of Nord-Pas-de-Calais (Contrat de Projets Etat-Région); the German Center for Diabetes Research; the German Research Council (including grant GRK1041); the German National Genome Research Network; Groupe d'Etude des Maladies Métaboliques et Systémiques; the Health Care Centers in Vasa, Närpes and Korsholm, Finland; the Health Foundation; the Heinz Nixdorf Foundation; Helmholtz Zentrum München; the Helsinki University Central Hospital Research Foundation; the Hospital District of Southwest Finland; the Ib Henriksens Foundation; IngaBritt and Arne Lundberg's Research Foundation (including grant 359); Karolinska Institutet; the Knut and Alice Wallenberg Foundation (including grant KAW 2009.0243); Kuopio University Hospital; the Lundbeck Foundation; the Magnus Bergvall Foundation; the Medical Faculty of University Duisburg-Essen; the Medical Research Council, UK (including grants G0000649 and G0601261); the Ministry for Health, Welfare and Sports, the Netherlands; the Ministry of Education and Culture, Finland (including grants 722 and 627; 2004-2011); the Ministry of Education, Culture and Science, the Netherlands; the Ministry of Health and Prevention, Denmark; the Ministry of Social Affairs and Health, Finland; the Ministry of Innovation, Science, Research and Technology of North Rhine-Westphalia, Germany; the Munich Center of Health Sciences; the Municipal Health Care Center and Hospital in Jakobstad, Finland; the municipality of Rotterdam, the Netherlands; the Närpes Health Care Foundation; the National Health Screening Service of Norway; the National Heart, Lung, and Blood Institute, USA (including grant numbers/contracts HHSN268201100005C, HHSN268201100006C, HHSN268201100007C, HHSN268201100008C, HHSN268201100009C, HHSN268201100010C, HHSN268201100011C, HHSN268201100012C, N01HC25195, N02HL64278, R01HL087641, R01HL59367 and R01HL086694); the National Human Genome Research Institute, USA (including grant numbers/contracts U01HG004402 and N01HG65403); the National Institute for Diabetes and Digestive and Kidney Diseases, USA (including grants R01DK078616, U01DK085526, K24DK080140 and R01DK073490); the National Institute for Health and Welfare, Finland; the National Institutes of Health, USA (including grant numbers/contracts HHSN268200625226C, UL1RR025005, R01DK062370, R01DK072193, 1Z01HG000024, AG028555, AG08724, AG04563, AG10175, AG08861, U01HG004399, DK58845, CA055075, DK085545 and DK098032); the Netherlands Genomics Initiative; the Netherlands Organisation for Health Research and Development; the Netherlands Organisation of Scientific Research NOW Investments (including grants 175.010.2005.011, 911-03-012 and 050-060-810); the Nord-Trondelag County Council; the Nordic Center of Excellence in Disease Genetics; the Norwegian Institute of Public Health; the Norwegian Research Council; the Novo Nordisk Foundation; the Ollquist Foundation; the Oxford National Institute for Health Research (NIHR) Biomedical Research Centre; the Paavo Nurmi Foundation; the Paivikki and Sakari Sohlberg Foundation; the Perklen Foundation; the Pirkanmaa Hospital District, Finland; Programme Hospitalier de Recherche Clinique; Programme National de Recherche sur la Diabète; the Research Institute for Diseases in the Elderly (including grant 014-93-015); the Robert Dawson Evans Endowment, Department of Medicine, Boston University School of Medicine and Boston Medical Center; the Royal Swedish Academy of Sciences; Sarstedt, Germany; the Signe and Ane Gyllenberg Foundation; the Sigrid Juselius Foundation; the Slottery Machine Association, Finland; the Social Insurance Institution of Finland; the South OstroBothnia Hospital District; the state of Baden-Württemberg, Germany; the Stockholm County Council (including grant 560183); the Swedish Cultural Foundation, Finland; the Swedish Diabetes Foundation; the Swedish e-science Research Center; the Swedish Foundation for Strategic Research; the Swedish Heart-Lung Foundation; the Swedish Research Council (including grants SFO EXODIAB 2009-1039, 521-2010-3490, 521-2007-4037, 521-2008-2974, ANDIS 825-2010-5983, LUDC 349-2008-6589 and 8691); the Swedish Society of Medicine; the Tore Nilsson Foundation; the Torsten and Ragnar Soderbergs Stiftelser (including grant MT33/09); University Hospital Essen; University of Tromsø; the University College London NIHR Biomedical Research Centre; the UK NIHR Cambridge Biomedical Research Centre; Uppsala University; Uppsala University Hospital; the Vaasa Hospital District; the Velux Foundation; and the Wellcome Trust (including the Biomedical Collections Grant GR072960 and grants 076113, 083948, 090367, 090532, 083270, 086596, 098017, 095101, 098051 and 098381). We are grateful to R. Scharfmann (INSERM U1016, Cochin Institute Paris) for the gift of EndoC βH1 cells and for providing technical support with their maintenance. We thank P. Johnson and the Oxford NIHR Biomedical Research Centre–funded Islet Isolation facility for providing human islets for this study. Detailed acknowledgments are provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Writing group. K.J.G., T.F., Y. Lee, A.R., R.M., M.E.R., A.L.G., D.A., M. Boehnke, T.M.T., M.I.M., A.P.M. Central meta-analysis group. K.J.G., T.F., Y. Lee, R.M., A. Mahajan, A. Locke, N.W.R., N.R., T.M.T., M.I.M., A.P.M. Annotation and functional analysis group. K.J.G., A.R., M.E.R., S.K.T., J.K.R., N.L.B., M.v.d.B., A.C., I.D., E. Birney, L. Pasquali, J. Ferrer, C.A.O'C., A.L.G., M.I.M. Validation meta-analysis group. R.M., R.A.S., I.P., L.J.S., A.P.M. Metabochip cohort-level primary analysis. Y. Lee, T.G., T.S., D.T., L.Y., H.G., S. Wahl, M.F., R.J.S., H. Kestler, H. Chheda, L.E., S.G., T.M.T., A.P.M. Validation cohort-level primary analysis. V. Steinthorsdottir, G.T., L.Q., L.C.K., E.M.v.L., S.M.W., M. Li, H. Chen, C. Fuchsberger, P. Kwan, C.M., M. Linderman, Y. Lu. Metabochip design. H.M.K., B.F.V. Cohort sample collection, genotyping, phenotyping or additional analysis. B.F.V., G.R.A., P.A., D.B., B.B., R.B., M. Blüher, H.B., L.L.B., E.P.B., N.P.B., J.C., G.C., P.S.C., M.C.C., D.J.C., A.T.C., R.M.v.D., A.S.F.D., M.D., S.E., J.G.E., T.E., E.E., J. Fadista, J. Flannick, P. Fontanillas, C. Fox, P.W.F., K.G., C.G., B.G., O.G., G.B.G., N.G., C.J.G., M.H., C.T.H., C.H., O.L.H., A.B.H., S.E.H., D.J.H., A.U.J., A.J., M.E.J., T.J., W.-H.L.K., N.D.K., L.K., N.K., A.K., P. Kovacs, P. Kraft, J. Kravic, C. Langford, K.L., L. Liang, P.L., C.M.L., E.L., A. Linneberg, C.-T.L., S.L., J.L., V.L., S. Männistö, O. McLeod, J.M., E.M., G.M., T.W.M., M.M.-N., C.N., M.M.N., N.N.O., K.R.O., D.P., S.P., L. Peltonen, J.R.B.P., C.G.P.P., M.R., D. Ruderfer, D. Rybin, Y.T.v.d.S., B.S., G. Sigur∂sson, A.S., G. Steinbach, P.S., K. Strauch, H.M.S., Q.S., B.T., E. Tikkanen, A.T., J. Trakalo, E. Tremoli, T.T., R.W., S. Wiltshire, A.R.W., E.Z. Validation cohort principal investigators. R.J.F.L., J.D., J.C.F., E. Boerwinkle, J.S.P., C.v.D., E.S., J.B.M., F.B.H., U.T., K. Stefansson, P.D., P.J.D., T.M.F., A.T.H., I.B., C. Langenberg, N.J.W., M. Boehnke, M.I.M. Metabochip cohort principal investigators. T.A.L., R.R., M.S., N.L.P., L. Lind, S.M.K.-K., E.K.-H., T.E.S., J.S., J. Kuusisto, M. Laakso, A. Metspalu, R.E., K.-H.J., S. Moebus, S.R., V. Salomaa, E.I., B.O.B., R.N.B., F.S.C., K.L.M., H. Koistinen, J. Tuomilehto, K.H., I.N., P.D., P.J.D., T.M.F., A.T.H., U.d.F., A.H., T.I., A.P., S.C., R.S., P. Froguel, O.P., T.H., A.D.M., C.N.A.P., S.K., O. Melander, P.M.N., L.C.G., I.B., C. Langenberg, N.J.W., D.A., M. Boehnke, M.I.M. Project management. K.J.G., A.L.G., D.A., M. Boehnke, T.M.T., M.I.M., A.P.M. DIAGRAM Consortium management. D.A., M. Boehnke, M.I.M.
Corresponding authors
Ethics declarations
Competing interests
V. Steinthorsdottir, G.T., A.K., U.T. and K. Stefansson are employed by deCODE Genetics/Amgen, Inc. I.B. and spouse own stock in GlaxoSmithKline and Incyte.
Additional information
A list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
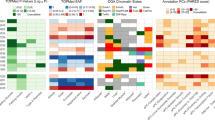
Supplementary Figure 1 Signal plots for association signals achieving locus-wide significance at the KCNQ1 locus.
Association summary statistics are based on the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry. Results are presented from exact conditioning after adjusting for all other index variants at the locus. Each point represents a SNP passing quality control in the meta-analysis, plotted with its conditional P value (on a –log10 scale) as a function of genomic position (NCBI Build 37). In each plot, the index variant is represented by the purple symbol. The color coding of all other SNPs indicates LD with the index variant in European-ancestry haplotypes from the 1000 Genomes Project reference panel: red, r2 ≥ 0.8; gold, 0.6 ≤ r2 < 0.8; green, 0.4 ≤ r2 < 0.6; cyan, 0.2 ≤ r2 < 0.4; blue, r2 < 0.2; gray, r2 unknown. The shape of the symbol corresponds to the annotation of the variant: upward triangle, framestop or splice; downward triangle, nonsynonymous; square, synonymous or UTR; and circle, intronic or noncoding. Recombination rates are estimated from HapMap Phase 2, and gene annotations are taken from the UCSC Genome Browser.
Supplementary Figure 2 Signal plots for association signals achieving locus-wide significance at the HNF1A locus.
Association summary statistics are based on the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry. Results are presented from exact conditioning after adjusting for all other index variants at the locus. Each point represents a SNP passing quality control in the meta-analysis, plotted with its conditional P value (on a –log10 scale) as a function of genomic position (NCBI Build 37). In each plot, the index variant is represented by the purple symbol. The color coding of all other SNPs indicates LD with the index variant in European-ancestry haplotypes from the 1000 Genomes Project reference panel: red, r2 ≥ 0.8; gold, 0.6 ≤ r2 < 0.8; green, 0.4 ≤ r2 < 0.6; cyan, 0.2 ≤ r2 < 0.4; blue, r2 < 0.2; gray, r2 unknown. The shape of the symbol corresponds to the annotation of the variant: upward triangle, framestop or splice; downward triangle, nonsynonymous; square, synonymous or UTR; and circle, intronic or noncoding. Recombination rates are estimated from HapMap Phase 2, and gene annotations are taken from the UCSC Genome Browser.
Supplementary Figure 3 Signal plots for association signals achieving locus-wide significance at the DGKB, CDKN2A-CDKN2B, MC4R and GIPR loci.
Association summary statistics are based on the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry. Results are presented from exact conditioning after adjusting for all other index variants at the locus. Each point represents a SNP passing quality control in the meta-analysis, plotted with its conditional P value (on a –log10 scale) as a function of genomic position (NCBI Build 37). In each plot, the index variant is represented by the purple symbol. The color coding of all other SNPs indicates LD with the index variant in European-ancestry haplotypes from the 1000 Genomes Project reference panel: red, r2 ≥ 0.8; gold, 0.6 ≤ r2 < 0.8; green, 0.4 ≤ r2 < 0.6; cyan, 0.2 ≤ r2 < 0.4; blue, r2 < 0.2; gray, r2 unknown. The shape of the symbol corresponds to the annotation of the variant: upward triangle, framestop or splice; downward triangle, nonsynonymous; square, synonymous or UTR; and circle, intronic or noncoding. Recombination rates are estimated from HapMap Phase 2, and gene annotations are taken from the UCSC Genome Browser.
Supplementary Figure 4 Signal plots for association signals at the CDKN2A-CDKN2B locus.
Association summary statistics are based on the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry. (a) Unconditional. (b) After approximate conditioning on the two index SNPs, rs10811660 and rs10757283. Each point represents a SNP passing quality control in the meta-analysis, plotted with its conditional P value (on a –log10 scale) as a function of genomic position (NCBI Build 37). In each plot, the index variant is represented by the purple symbol. The color coding of all other SNPs indicates LD with the index variant in European-ancestry haplotypes from the 1000 Genomes Project reference panel: red, r2 ≥ 0.8; gold, 0.6 ≤ r2 < 0.8; green, 0.4 ≤ r2 < 0.6; cyan, 0.2 ≤ r2 < 0.4; blue, r2 < 0.2; gray, r2 unknown. Recombination rates are estimated from HapMap Phase 2, and gene annotations are taken from the UCSC Genome Browser.
Supplementary Figure 5 Characteristics of index variants for association signals achieving locus-wide significance for T2D susceptibility across fine-mapping regions.
Each association signal was represented by an index variant in the GCTA-COJO joint regression model on the basis of (i) summary statistics from the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry and (ii) reference genotype data from GoDARTS (3,298 cases and 3,708 controls of European ancestry from the UK) to approximate LD across fine-mapping regions. Each variant is plotted according to risk allele frequency on the x axis and allelic log(OR) on the y axis (with error bars representing the corresponding standard error).
Supplementary Figure 6 Characteristics of the 99% credible sets of variants for each association signal achieving locus-wide significance for T2D susceptibility across fine-mapping regions.
Each point corresponds to an association signal, plotted according to the highest posterior probability of causality of any variant in the credible set on the x axis and the total number of variants in the credible set on the y axis. Distinct association signals at loci are referenced according to their index variant.
Supplementary Figure 7 Enrichment in chromatin state and noncoding RNA elements.
Variants in regulatory enhancer, promoter and insulator elements from 12 cell types and noncoding RNA elements from 25 cell types were tested for enrichment of average posterior causal probability compared to variants in shifted elements. Variants in islet enhancer elements demonstrated significant enrichment (fold = 1.97; P = 0.00022), and variants in islet and HepG2 promoter elements demonstrated nominally significant enrichment (P < 0.01).
Supplementary Figure 8 Null distribution of enriched transcription factors.
Null distribution of mean posterior probabilities of driving association signals for variants in shifted annotations (blue bars) compared to observed mean probability (dashed line) for the most significantly enriched transcription factors.
Supplementary Figure 9 FOXA2 ChIP-seq sites overlap, genome wide, across cell types.
The number of FOXA2 sites identified in ChIP-seq assays from primary pancreatic islets, HepG2 cells and primary liver was obtained, and sites from each cell type were intersected.
Supplementary Figure 10 Electrophoretic mobility shift assay of rs10830963 in HepG2 cellular extracts.
Nuclear extract (NE) from HepG2 cells was treated with labeled probes of 25 bp of sequence containing each allele of rs10830963. We observed protein bands bound to both sequences (black) as well as bands unique to the non-risk (red) and risk (blue) sequences. We then introduced antibodies against HNF3B (FOXA2 alias) and four factors (YY1, TAL1, NEURDO1 and PTF1A) whose known binding motifs match the de novo motif at rs10830963. We observed no shifting of the allele-specific bands with any antibody. Several bands shared by both alleles were removed in the presence of antibody to YY1.
Supplementary Figure 11 Genes at FOXA2-enriched signals are downregulated in Foxa1/Foxa2 null mice.
Pancreatic islet gene expression data were obtained from wild-type and Foxa1/Foxa2 knockout mice (Gao et al.47). Genes within 500 kb of each FOXA2-enriched T2D signal and closest to each FOXA2-enriched signal (orange) were significantly downregulated in the Foxa1/Foxa2 knockout compared to all genes (gray).
Supplementary Figure 12 Comparison of summary statistics for association signals achieving locus-wide significance from the GCTA-COJO joint regression model using genotype data from two reference studies to approximate LD between variants across each fine-mapping region.
Association summary statistics are based on the meta-analysis of Metabochip studies in 27,206 cases and 57,574 controls of European ancestry. Association summary statistics were obtained using (i) 3,298 T2D cases and 3,708 controls of UK ancestry from GoDARTS (represented on the x axis) and (ii) 4,435 T2D cases and 5,757 controls of Scandinavian ancestry from FUSION (represented on the y axis). Left, comparison of P values (PJ) from the GCTA-COJO joint regression model. Right, comparison of allelic log(OR) (blue diamond) and standard error (gray error bars) from the GCTA-COJO joint regression model.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–17 and Supplementary Note. (PDF 5196 kb)
Rights and permissions
About this article
Cite this article
Gaulton, K., Ferreira, T., Lee, Y. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet 47, 1415–1425 (2015). https://doi.org/10.1038/ng.3437
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3437
This article is cited by
-
MTNR1B genotype and effects of carbohydrate quantity and dietary glycaemic index on glycaemic response to an oral glucose load: the OmniCarb trial
Diabetologia (2024)
-
Genome-wide differential expression profiling of long non-coding RNAs in FOXA2 knockout iPSC-derived pancreatic cells
Cell Communication and Signaling (2023)
-
Genetics of circadian rhythms and sleep in human health and disease
Nature Reviews Genetics (2023)
-
Emerging therapeutic options in the management of diabetes: recent trends, challenges and future directions
International Journal of Obesity (2023)
-
Monogenic diabetes
Nature Reviews Disease Primers (2023)