Abstract
Juvenile myelomonocytic leukemia (JMML) is a rare and severe myelodysplastic and myeloproliferative neoplasm of early childhood initiated by germline or somatic RAS-activating mutations1,2,3. Genetic profiling and whole-exome sequencing of a large JMML cohort (118 and 30 cases, respectively) uncovered additional genetic abnormalities in 56 cases (47%). Somatic events were rare (0.38 events/Mb/case) and restricted to sporadic (49/78; 63%) or neurofibromatosis type 1 (NF1)-associated (8/8; 100%) JMML cases. Multiple concomitant genetic hits targeting the RAS pathway were identified in 13 of 78 cases (17%), disproving the concept of mutually exclusive RAS pathway mutations and defining new pathways activated in JMML involving phosphoinositide 3-kinase (PI3K) and the mTORC2 complex through RAC2 mutation. Furthermore, this study highlights PRC2 loss (26/78; 33% of sporadic JMML cases) that switches the methylation/acetylation status of lysine 27 of histone H3 in JMML cases with altered RAS and PRC2 pathways. Finally, the association between JMML outcome and mutational profile suggests a dose-dependent effect for RAS pathway activation, distinguishing very aggressive JMML rapidly progressing to acute myeloid leukemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Referenced accessions
NCBI Reference Sequence
Protein Data Bank
References
Chang, T.Y., Dvorak, C.C. & Loh, M.L. Bedside to bench in juvenile myelomonocytic leukemia: insights into leukemogenesis from a rare pediatric leukemia. Blood 124, 2487–2497 (2014).
Niemeyer, C.M. et al. Chronic myelomonocytic leukemia in childhood: a retrospective analysis of 110 cases. European Working Group on Myelodysplastic Syndromes in Childhood (EWOG-MDS). Blood 89, 3534–3543 (1997).
Locatelli, F. & Niemeyer, C.M. How I treat juvenile myelomonocytic leukemia. Blood 125, 1083–1090 (2015).
Niemeyer, C.M. RAS diseases in children. Haematologica 99, 1653–1662 (2014).
Shlien, A. et al. Combined hereditary and somatic mutations of replication error repair genes result in rapid onset of ultra-hypermutated cancers. Nat. Genet. 47, 257–262 (2015).
Sakaguchi, H. et al. Exome sequencing identifies secondary mutations of SETBP1 and JAK3 in juvenile myelomonocytic leukemia. Nat. Genet. 45, 937–941 (2013).
Strullu, M. et al. Juvenile myelomonocytic leukaemia and Noonan syndrome. J. Med. Genet. 51, 689–697 (2014).
Niemeyer, C.M. et al. Germline CBL mutations cause developmental abnormalities and predispose to juvenile myelomonocytic leukemia. Nat. Genet. 42, 794–800 (2010).
Pérez, B. et al. Germline mutations of the CBL gene define a new genetic syndrome with predisposition to juvenile myelomonocytic leukaemia. J. Med. Genet. 47, 686–691 (2010).
Makishima, H. et al. Somatic SETBP1 mutations in myeloid malignancies. Nat. Genet. 45, 942–946 (2013).
Brown, K.M. et al. Phosphodiesterase-8A binds to and regulates Raf-1 kinase. Proc. Natl. Acad. Sci. USA 110, E1533–E1542 (2013).
Flex, E. et al. Activating mutations in RRAS underlie a phenotype within the RASopathy spectrum and contribute to leukaemogenesis. Hum. Mol. Genet. 23, 4315–4327 (2014).
Shang, X. et al. R-Ras and Rac GTPase cross-talk regulates hematopoietic progenitor cell migration, homing, and mobilization. J. Biol. Chem. 286, 24068–24078 (2011).
Kawazu, M. et al. Transforming mutations of RAC guanosine triphosphatases in human cancers. Proc. Natl. Acad. Sci. USA 110, 3029–3034 (2013).
Emanuel, P.D. Hallway gossip between Ras and PI3K pathways. Blood 123, 2751–2753 (2014).
Goodwin, C.B. et al. PI3K p110δ uniquely promotes gain-of-function Shp2-induced GM-CSF hypersensitivity in a model of JMML. Blood 123, 2838–2842 (2014).
Worzfeld, T. et al. Genetic dissection of plexin signaling in vivo. Proc. Natl. Acad. Sci. USA 111, 2194–2199 (2014).
Innocenti, M. et al. Phosphoinositide 3-kinase activates Rac by entering in a complex with Eps8, Abi1, and Sos-1. J. Cell Biol. 160, 17–23 (2003).
Offenhäuser, N. et al. The eps8 family of proteins links growth factor stimulation to actin reorganization generating functional redundancy in the Ras/Rac pathway. Mol. Biol. Cell 15, 91–98 (2004).
Cutts, B.A. et al. Nf1 deficiency cooperates with oncogenic K-RAS to induce acute myeloid leukemia in mice. Blood 114, 3629–3632 (2009).
Xu, J. et al. Dominant role of oncogene dosage and absence of tumor suppressor activity in Nras-driven hematopoietic transformation. Cancer Discov. 3, 993–1001 (2013).
Wang, J. et al. Endogenous oncogenic Nras mutation initiates hematopoietic malignancies in a dose- and cell type–dependent manner. Blood 118, 368–379 (2011).
Khan, S.N. et al. Multiple mechanisms deregulate EZH2 and histone H3 lysine 27 epigenetic changes in myeloid malignancies. Leukemia 27, 1301–1309 (2013).
Son, J., Shen, S.S., Margueron, R. & Reinberg, D. Nucleosome-binding activities within JARID2 and EZH1 regulate the function of PRC2 on chromatin. Genes Dev. 27, 2663–2677 (2013).
Kinkel, S.A. et al. Jarid2 regulates hematopoietic stem cell function by acting with Polycomb repressive complex 2. Blood 125, 1890–1900 (2015).
Ciferri, C. et al. Molecular architecture of human Polycomb repressive complex 2. eLife 1, e00005 (2012).
Puda, A. et al. Frequent deletions of JARID2 in leukemic transformation of chronic myeloid malignancies. Am. J. Hematol. 87, 245–250 (2012).
Zhang, Y. et al. Corepressor protein CDYL functions as a molecular bridge between Polycomb repressor complex 2 and repressive chromatin mark trimethylated histone lysine 27. J. Biol. Chem. 286, 42414–42425 (2011).
Abdel-Wahab, O. et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 22, 180–193 (2012).
Abdel-Wahab, O. & Levine, R.L. Mutations in epigenetic modifiers in the pathogenesis and therapy of acute myeloid leukemia. Blood 121, 3563–3572 (2013).
Kim, E. et al. SRSF2 mutations contribute to myelodysplasia by mutant-specific effects on exon recognition. Cancer Cell 27, 617–630 (2015).
Huether, R. et al. The landscape of somatic mutations in epigenetic regulators across 1,000 paediatric cancer genomes. Nat. Commun. 5, 3630 (2014).
Yang, F. PRC2 dysfunction through multiple mechanisms in myeloid malignancies. Epigenomics 5, 481 (2013).
Pasini, D. et al. Characterization of an antagonistic switch between histone H3 lysine 27 methylation and acetylation in the transcriptional regulation of Polycomb group target genes. Nucleic Acids Res. 38, 4958–4969 (2010).
Leiserson, M.D. et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genet. 47, 106–114 (2015).
Lee, W. et al. PRC2 is recurrently inactivated through EED or SUZ12 loss in malignant peripheral nerve sheath tumors. Nat. Genet. 46, 1227–1232 (2014).
Zhang, M. et al. Somatic mutations of SUZ12 in malignant peripheral nerve sheath tumors. Nat. Genet. 46, 1170–1172 (2014).
De Raedt, T. et al. PRC2 loss amplifies Ras-driven transcription and confers sensitivity to BRD4-based therapies. Nature 514, 247–251 (2014).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Jacobs, K.B. et al. Detectable clonal mosaicism and its relationship to aging and cancer. Nat. Genet. 44, 651–658 (2012).
Laurie, C.C. et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat. Genet. 44, 642–650 (2012).
Locatelli, F. et al. Hematopoietic stem cell transplantation (HSCT) in children with juvenile myelomonocytic leukemia (JMML): results of the EWOG-MDS/EBMT trial. Blood 105, 410–419 (2005).
Takagi, M. et al. Autoimmunity and persistent RAS-mutated clones long after the spontaneous regression of JMML. Leukemia 27, 1926–1928 (2013).
Matsuda, K. et al. Spontaneous improvement of hematologic abnormalities in patients having juvenile myelomonocytic leukemia with specific RAS mutations. Blood 109, 5477–5480 (2007).
Matsuda, K. et al. Acquisition of loss of the wild-type NRAS locus with aggressive disease progression in a patient with juvenile myelomonocytic leukemia and a heterozygous NRAS mutation. Haematologica 92, 1576–1578 (2007).
Kato, M. et al. Aggressive transformation of juvenile myelomonocytic leukemia associated with duplication of oncogenic KRAS due to acquired uniparental disomy. J. Pediatr. 162, 1285–1288 (2013).
Kalra, R., Paderanga, D.C., Olson, K. & Shannon, K.M. Genetic analysis is consistent with the hypothesis that NF1 limits myeloid cell growth through p21ras. Blood 84, 3435–3439 (1994).
Chan, R.J., Cooper, T., Kratz, C.P., Weiss, B. & Loh, M.L. Juvenile myelomonocytic leukemia: a report from the 2nd International JMML Symposium. Leuk. Res. 33, 355–362 (2009).
Pérez, B. et al. Genetic typing of CBL, ASXL1, RUNX1, TET2 and JAK2 in juvenile myelomonocytic leukaemia reveals a genetic profile distinct from chronic myelomonocytic leukaemia. Br. J. Haematol. 151, 460–468 (2010).
Ahmadian, M.R., Stege, P., Scheffzek, K. & Wittinghofer, A. Confirmation of the arginine-finger hypothesis for the GAP-stimulated GTP-hydrolysis reaction of Ras. Nat. Struct. Biol. 4, 686–689 (1997).
Hemsath, L. & Ahmadian, M.R. Fluorescence approaches for monitoring interactions of Rho GTPases with nucleotides, regulators, and effectors. Methods 37, 173–182 (2005).
Eberth, A. & Ahmadian, M.R. In vitro GEF and GAP assays. Curr. Protoc. Cell Biol. Chapter 14, Unit 14.9 (2009).
Jaiswal, M., Dubey, B.N., Koessmeier, K.T., Gremer, L. & Ahmadian, M.R. Biochemical assays to characterize Rho GTPases. Methods Mol. Biol. 827, 37–58 (2012).
Eberth, A. et al. A BAR domain–mediated autoinhibitory mechanism for RhoGAPs of the GRAF family. Biochem. J. 417, 371–377 (2009).
Jaiswal, M., Dubey, B.N., Koessmeier, K.T., Gremer, L. & Ahmadian, M.R. Biochemical assays to characterize Rho GTPases. Methods Mol. Biol. 827, 37–58 (2012).
Gremer, L. et al. Germline KRAS mutations cause aberrant biochemical and physical properties leading to developmental disorders. Hum. Mutat. 32, 33–43 (2011).
Nakhaei-Rad, S. et al. The function of embryonic stem cell–expressed RAS (E-RAS), a unique RAS family member, correlates with its additional motifs and its structural properties. J. Biol. Chem. 290, 15892–15903 (2015).
Worthylake, D.K., Rossman, K.L. & Sondek, J. Crystal structure of Rac1 in complex with the guanine nucleotide exchange region of Tiam1. Nature 408, 682–688 (2000).
Morreale, A. et al. Structure of Cdc42 bound to the GTPase binding domain of PAK. Nat. Struct. Mol. Biol. 7, 384–388 (2000).
Nassar, N., Hoffman, G.R., Manor, D., Clardy, J.C. & Cerione, R.A. Structures of Cdc42 bound to the active and catalytically compromised forms of Cdc42GAP. Nat. Struct. Biol. 5, 1047–1052 (1998).
Acknowledgements
We thank N. Roodur, M. Keita (Unité de Génétique Moléculaire, Hospital Robert Debré) and L. Coste-Sarguet (INSERM UMR 1131) for expert technical assistance. We thank E. Martin, J.-P. Sarava and M. Letexier (Integragen) for advice and assistance in whole-exome sequencing analysis. SNP array analyses were performed by the Plateforme de Génomique Constitutionnelle–Nord (PfGC-Nord) (J. Soulier, S. Quentin and S. Drunat). We thank the technical team, and more particularly P. Aubin, from the Service de Biologie Cellulaire, Hôpital Saint Louis, for excellent technical help in the management of myeloid progenitor assays. We also thank S. Rasika for careful editing of the manuscript.
This work was supported by the Ligue contre le Cancer (LCC)–Ile-de-France, by the Société Française de Lutte contre les Cancers et Leucémies de l'Enfant (SFCE), the Association pour la Recherche et pour les Etudes dans les Maladies Infantiles Graves (AREMIG, Nancy) and the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG) through the Collaborative Research Center 974 (SFB 974) Communication and Systems Relevance during Liver Injury and Regeneration and the International Graduate School of Protein Science and Technology (iGRASP).
Author information
Authors and Affiliations
Contributions
A. Caye collected subject samples, participated in the study design, performed laboratory assays, analyzed data and wrote the manuscript. M.S. and J.L. performed analyses and collected clinical data. F.G. performed epigenetic studies, and B.C. and E.V. performed colony assays. S.G. performed biostatistical analyses. O.F. and E.L. performed the cytological review. K.N., S.N.-R., R.D., D.H. and M.R.A. performed the functional and biochemical RAC2 studies. J.V. collected clinical data. S.P. performed laboratory assays. D.V. performed NF1 diagnosis. J.-H.D., A.B., C. Paillard, C. Picard, C.G., A.P., Y.R., F.M., B.N., Y.B., M.P., D.A., N.S. and A. Contet contributed subject samples and clinical data. M.S., A.B., C.C., B.C. and M.R.A. reviewed the manuscript. H.C. collected subject samples, designed and coordinated the study, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Spectrum of somatically acquired mutations identified by combining WES and genome-wide DNA array analysis in the discovery cohort of 30 JMML cases.
A total of 85 somatically acquired alterations were found, including 64 nonsynonymous point mutations or small insertion-deletions (indels) identified in the coding regions of these tumors by WES and 21 somatic cytogenetic alterations evidenced by SNP/CGH array, WES and/or metaphase cytogenetics.
Supplementary Figure 2 Graphical representation of the type of data obtained by sample in a cohort of 118 patients with JMML.
Both the discovery cohort (top; n = 30) and validation cohort (bottom; n = 88) are represented.
Supplementary Figure 3 Distribution of RAS-related mutations in a cohort of 118 patients with JMML as detected by routine workup.
Distribution of RAS-related mutations in 118 consecutively diagnosed JMML cases as detected by routine workup and detailed spectrum of KRAS (n = 18) and NRAS (n = 22) mutations.
Supplementary Figure 5 Proportion of mutations predicted to be deleterious versus non-pathogenic substitutions.
The pathogenicity of somatic nonsynonymous exonic missense variants with respect to gene function was predicted using the SIFT, PolyPhen-2 and MutationTaster algorithms (Supplementary Table 6). A total of 91% of all missense mutations were predicted to result in functionally relevant alterations by at least two of the three methods used for functional prediction. This percentage was similar when considering only initiating mutations, known to be deleterious in all cases (92%), as well as secondary mutations (89%).
Supplementary Figure 6 Sequence electrophoregram showing the presence of three concomitant mutations targeting the RAS pathway at diagnosis of JMML_89.
Mutated nucleotides are indicated by a red arrow. The subclonal pattern of NF1 mutation is consistent with late acquisition. NF1 haploinsufficiency was due to a recurrent c.2033delG mutation of a G homopolymer within the NF1 coding region (exon 18). The frequency of this mutation appeared strikingly higher among somatic variants (5/6 cases with a secondary NF1 mutation) than among germline variants (Leiden Open Variation Database, LOVD).
Supplementary Figure 7 Hyperactive RAC2 Asp63Val contributes to AKT activation via both the PI3K-PDK1 and mTORC2 cascades.
Pull-down experiments (a,b) and immunoblot (IB) analysis were conducted using total cell lysates (c–g) derived from transfected COS-7 cells with FLAG-tagged RAC2 and RAS variants. The GTPase-binding domain (GBD) of the RAC effector PAK1 was used as a GST fusion protein for the pulldown experiment. All experiments were performed three times. (a) Pulldown analysis showed that RAC2 Asp63Val largely exists in an active, GTP-bound state as compared to wild-type RAC2 (RAC2 WT), but activation is not as strong as for constitutively active RAC2 Gly12Val. Total RAC2 and RAS proteins were detected using antibody to FLAG and pan-RAS antibody to show the total amounts of the transfected FLAG-tagged RAC2 and RAS variants. (b) The RAC2 protein bands in a were densitometrically quantified (depicted as numbers and bars) as the amount of the GTP-bound RAC2 protein relative to wild-type RAC2. Coexpression of constitutively active NRAS Gly12Val, HRAS Gly12Val and KRAS Gly12Val did not change the level of GTP-bound RAC2 Asp63Val. (c) Total cell lysates were analyzed for the phosphorylation levels of AKT (pAKT 308 and pAKT 473), MEK1/2 (pMEK1/2) and ERK1/2 (pERK1/2). Total amounts of these kinases were used as loading controls. AKT is phosphorylated at Thr308 by the PI3K-PDK1 pathway, whereas the mTORC2 complex phosphorylates AKT at Ser473. (d–g) The protein bands in c were densitometrically quantified (depicted as numbers and bars), clearly showing that the presence of RAC2 Asp63Val resulted in strong AKT phosphorylation and slight MEK phosphorylation but no ERK phosphorylation. Interestingly, a comparison of RAC2 Asp63Val and RAC2 Gly12Val showed that the relative amount of phosphorylated protein was proportional to the amount of the GTP-bound protein.
Supplementary Figure 8 Sequence electrophoregram showing progressive LOH of the KRAS locus with an allelic imbalance in favor of the oncogenic allele in JMML_24.
Wild-type (WT) and mutated nucleotides are indicated by black and red arrows, respectively.
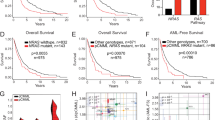
Supplementary Figure 9 Overall survival in sporadic JMML according to initiating RAS-activating lesion.
Kaplan-Meier representation of the overall survival (%) in 96 patients with JMML evaluable for follow-up. Patients with Noonan syndrome were excluded from the analysis.
Supplementary Figure 11 Performances of PCR-based targeted deep sequencing in 75 JMML samples.
(a) The mean coverage of the coding regions is plotted for each gene by 25× descending order. (b) The mean depth of sequencing is plotted for each gene on a logarithmic scale, by descending order.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1, 4 and 5 and Supplementary Note. (PDF 2807 kb)
Supplementary Table 2
List of variants identified by whole-exome sequencing. (XLSX 44 kb)
Supplementary Table 3
List of copy number abnormalities and copy-neutral loss of heterozygosity evidenced by genome-wide DNA arrayanalysis and whole-exome sequencing. (XLSX 22 kb)
Supplementary Table 6
List of all sequence mutations detected in the cohort of 118 JMML cases. (XLSX 55 kb)
Supplementary Table 7
Genetic analysis of myeloid colonies obtained by in vitroculture on methyl-cellulose. (XLSX 30 kb)
Supplementary Table 8
List of PCR primers used for targeted deep sequencing and Sanger targeted sequencing. (XLSX 116 kb)
Supplementary Table 9
Tabular data for survival curves. (XLSX 21 kb)
Rights and permissions
About this article
Cite this article
Caye, A., Strullu, M., Guidez, F. et al. Juvenile myelomonocytic leukemia displays mutations in components of the RAS pathway and the PRC2 network. Nat Genet 47, 1334–1340 (2015). https://doi.org/10.1038/ng.3420
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3420
This article is cited by
-
Epigenetic regulation by ASXL1 in myeloid malignancies
International Journal of Hematology (2023)
-
ASXL1/2 mutations and myeloid malignancies
Journal of Hematology & Oncology (2022)
-
Pediatric Germline Predisposition to Myeloid Neoplasms
Current Hematologic Malignancy Reports (2022)
-
RAS activation induces synthetic lethality of MEK inhibition with mitochondrial oxidative metabolism in acute myeloid leukemia
Leukemia (2022)
-
Integrated in silico MS-based phosphoproteomics and network enrichment analysis of RASopathy proteins
Orphanet Journal of Rare Diseases (2021)