Abstract
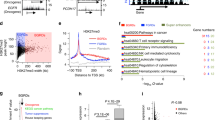
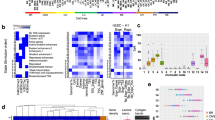
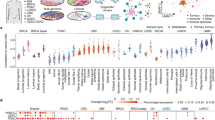
Tumor suppressors are mostly defined by inactivating mutations in tumors, yet little is known about their epigenetic features in normal cells. Through integrative analysis of 1,134 genome-wide epigenetic profiles, mutations from >8,200 tumor-normal pairs and our experimental data from clinical samples, we discovered broad peaks for trimethylation of histone H3 at lysine 4 (H3K4me3; wider than 4 kb) as the first epigenetic signature for tumor suppressors in normal cells. Broad H3K4me3 is associated with increased transcription elongation and enhancer activity, which together lead to exceptionally high gene expression, and is distinct from other broad epigenetic features, such as super-enhancers. Genes with broad H3K4me3 peaks conserved across normal cells may represent pan-cancer tumor suppressors, such as TP53 and PTEN, whereas genes with cell type–specific broad H3K4me3 peaks may represent cell identity genes and cell type–specific tumor suppressors. Furthermore, widespread shortening of broad H3K4me3 peaks in cancers is associated with repression of tumor suppressors. Thus, the broad H3K4me3 epigenetic signature provides mutation-independent information for the discovery and characterization of new tumor suppressors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Weinstein, J.N. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
De, S. & Michor, F. DNA replication timing and long-range DNA interactions predict mutational landscapes of cancer genomes. Nat. Biotechnol. 29, 1103–1108 (2011).
Schuster-Böckler, B. & Lehner, B. Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488, 504–507 (2012).
Lawrence, M.S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Lauberth, S.M. et al. H3K4me3 interactions with TAF3 regulate preinitiation complex assembly and selective gene activation. Cell 152, 1021–1036 (2013).
Zhao, X.D. et al. Whole-genome mapping of histone H3 Lys4 and 27 trimethylations reveals distinct genomic compartments in human embryonic stem cells. Cell Stem Cell 1, 286–298 (2007).
Sims, R.J. et al. Recognition of trimethylated histone H3 lysine 4 facilitates the recruitment of transcription postinitiation factors and pre-mRNA splicing. Mol. Cell 28, 665–676 (2007).
Borde, V. et al. Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J. 28, 99–111 (2009).
Pena, P.V., Hom, R.A., Hung, T., Lin, H. & Kuo, A.J. Histone H3K4me3 binding is required for the DNA repair and apoptotic activities of ING1 tumor suppressor. J. Mol. Biol. 380, 303–312 (2008).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why? Mol. Cell 49, 825–837 (2013).
Whyte, W.A. et al. Master transcription factors and Mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Virely, C. et al. Haploinsufficiency of the IKZF1 (IKAROS) tumor suppressor gene cooperates with BCR-ABL in a transgenic model of acute lymphoblastic leukemia. Leukemia 24, 1200–1204 (2010).
Georgopoulos, K., Winandy, S. & Avitahl, N. The role of the Ikaros gene in lymphocyte development and homeostasis. Annu. Rev. Immunol. 15, 155–176 (1997).
Dail, M. et al. Mutant Ikzf1, KrasG12D, and Notch1 cooperate in T lineage leukemogenesis and modulate responses to targeted agents. Proc. Natl. Acad. Sci. USA 107, 5106–5111 (2010).
Gutierrez, A. et al. High frequency of PTEN, PI3K, and AKT abnormalities in T-cell acute lymphoblastic leukemia. Blood 114, 647–650 (2009).
Mendes, R.D. et al. PTEN microdeletions in T-cell acute lymphoblastic leukemia are caused by illegitimate RAG-mediated recombination events. Blood 124, 567–578 (2014).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Eisenberg, E. & Levanon, E.Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Yasuo, K. & Shigeo, O. Regulation of mitochondrial ATP synthesis in mammalian cells by transcriptional control. Int. J. Biochem. 22, 219–229 (1990).
Smolle, M. & Workman, J.L. Transcription-associated histone modifications and cryptic transcription. Biochim. Biophys. Acta 1829, 84–97 (2013).
Rahl, P.B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Glover-Cutter, K., Kim, S., Espinosa, J. & Bentley, D.L. RNA polymerase II pauses and associates with pre-mRNA processing factors at both ends of genes. Nat. Struct. Mol. Biol. 15, 71–78 (2008).
Jonkers, I., Kwak, H. & Lis, J.T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Veloso, A. et al. Rate of elongation by RNA polymerase II is associated with specific gene features and epigenetic modifications. Genome Res. 24, 896–905 (2014).
Nie, Z. et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 151, 68–79 (2012).
Lin, C.Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012).
Chao, S.H. & Price, D.H. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J. Biol. Chem. 276, 31793–31799 (2001).
Kheradpour, P. & Kellis, M. Systematic discovery and characterization of regulatory motifs in ENCODE TF binding experiments. Nucleic Acids Res. 42, 2976–2987 (2014).
Wang, J. et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 22, 1798–1812 (2012).
Barrett, C.W. et al. Tumor suppressor function of the plasma glutathione peroxidase Gpx3 in colitis-associated carcinoma. Cancer Res. 73, 1245–1255 (2013).
Matthew, E.M. et al. The p53 target Plk2 interacts with TSC proteins impacting mTOR signaling, tumor growth and chemosensitivity under hypoxic conditions. Cell Cycle 8, 4168–4175 (2009).
Coley, H.M. et al. Polo like kinase 2 tumour suppressor and cancer biomarker: new perspectives on drug sensitivity/resistance in ovarian cancer. Oncotarget 3, 78–83 (2012).
Smith, P., Syed, N. & Crook, T. Epigenetic inactivation implies a tumor suppressor function in hematologic malignancies for Polo-like kinase 2 but not Polo-like kinase 3. Cell Cycle 5, 1262–1264 (2006).
Rincon-Arano, H., Halow, J., Delrow, J.J., Parkhurst, S.M. & Groudine, M. UpSET recruits HDAC complexes and restricts chromatin accessibility and acetylation at promoter regions. Cell 151, 1214–1228 (2012).
Ali, M. et al. Molecular basis for chromatin binding and regulation of MLL5. Proc. Natl. Acad. Sci. USA 110, 11296–11301 (2013).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nat. Genet. 46, 17–23 (2014).
Pekowska, A., Benoukraf, T., Ferrier, P. & Spicuglia, S. A unique H3K4me2 profile marks tissue-specific gene regulation. Genome Res. 20, 1493–1502 (2010).
Shulha, H.P. et al. Epigenetic signatures of autism: trimethylated H3K4 landscapes in prefrontal neurons. Arch. Gen. Psychiatry 69, 314–324 (2012).
Benayoun, B.A. et al. H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell 158, 673–688 (2014).
Chen, Z. et al. Agonist and antagonist switch DNA motifs recognized by human androgen receptor in prostate cancer. EMBO J. 34, 502–516 (2015).
Wang, Q. et al. Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell 138, 245–256 (2009).
Chen, K. et al. DANPOS: Dynamic Analysis of Nucleosome Position and Occupancy by Sequencing. Genome Res. 23, 341–351 (2013).
Acknowledgements
We are grateful to M. Luo, M. Goodell, L. Donehower, T. Westbrook and A. Brunet for helpful discussions. This work was supported by the US National Institutes of Health (NIH) grants R01HG007538 and R01CA193466, Cancer Prevention Research Institute of Texas (CPRIT) grants RP110471 and RP150292 (W.L.), and US NIH grant R01CA151979, US Department of Defense grant W81XWH-12-1-0615 and US NIH grant U54CA113001 (Q.W.). X.S. is an inaugural MD Anderson Cancer Center R. Lee Clark Fellow.
Author information
Authors and Affiliations
Contributions
K.C. and W.L. conceived the project, designed the experiments and performed the data analysis. Z.C., D.W. and Q.W. designed and performed the experiments. L.Z. designed the experiments and performed the data analysis. X.L., J.S., Y.X. and Z.X. analyzed the data. K.C., Q.W. and W.L. interpreted the data and wrote the manuscript with comments from B.R., X.C. and X.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–24. (PDF 2813 kb)
Supplementary Table 1
A list of public data sets used in this study. (XLS 530 kb)
Supplementary Table 2
Number of genes assigned with broad H3K4me3 peaks in each sample. (XLS 67 kb)
Supplementary Table 3
H3K4me3 peak width at each gene in each sample from ENCODE and Roadmap Epigenomics. (XLS 28472 kb)
Supplementary Table 4
Broad H3K4me3 peaks at tumor suppressors that are shortened, lengthened or stable between 105 normal and 63 cancer samples. (XLS 29 kb)
Rights and permissions
About this article
Cite this article
Chen, K., Chen, Z., Wu, D. et al. Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor-suppressor genes. Nat Genet 47, 1149–1157 (2015). https://doi.org/10.1038/ng.3385
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3385
This article is cited by
-
Multimodal epigenetic sequencing analysis (MESA) of cell-free DNA for non-invasive colorectal cancer detection
Genome Medicine (2024)
-
Dynamic and distinct histone modifications facilitate human trophoblast lineage differentiation
Scientific Reports (2024)
-
An overview of artificial intelligence in the field of genomics
Discover Artificial Intelligence (2024)
-
Inhibition of fatty acid oxidation enables heart regeneration in adult mice
Nature (2023)
-
Transcription and FACT facilitate the restoration of replication-coupled chromatin assembly defects
Scientific Reports (2023)