Abstract
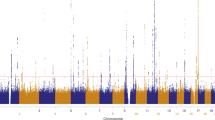
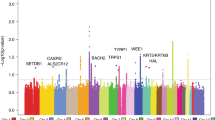
Thirteen common susceptibility loci have been reproducibly associated with cutaneous malignant melanoma (CMM). We report the results of an international 2-stage meta-analysis of CMM genome-wide association studies (GWAS). This meta-analysis combines 11 GWAS (5 previously unpublished) and a further three stage 2 data sets, totaling 15,990 CMM cases and 26,409 controls. Five loci not previously associated with CMM risk reached genome-wide significance (P < 5 × 10−8), as did 2 previously reported but unreplicated loci and all 13 established loci. Newly associated SNPs fall within putative melanocyte regulatory elements, and bioinformatic and expression quantitative trait locus (eQTL) data highlight candidate genes in the associated regions, including one involved in telomere biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Holly, E.A., Aston, D.A., Cress, R.D., Ahn, D.K. & Kristiansen, J.J. Cutaneous melanoma in women. II. Phenotypic characteristics and other host-related factors. Am. J. Epidemiol. 141, 934–942 (1995).
Holly, E.A., Aston, D.A., Cress, R.D., Ahn, D.K. & Kristiansen, J.J. Cutaneous melanoma in women. I. Exposure to sunlight, ability to tan, and other risk factors related to ultraviolet light. Am. J. Epidemiol. 141, 923–933 (1995).
Naldi, L. et al. Cutaneous malignant melanoma in women. Phenotypic characteristics, sun exposure, and hormonal factors: a case-control study from Italy. Ann. Epidemiol. 15, 545–550 (2005).
Titus-Ernstoff, L. et al. Pigmentary characteristics and moles in relation to melanoma risk. Int. J. Cancer 116, 144–149 (2005).
Bataille, V. et al. Risk of cutaneous melanoma in relation to the numbers, types and sites of naevi: a case-control study. Br. J. Cancer 73, 1605–1611 (1996).
Chang, Y.M. et al. A pooled analysis of melanocytic nevus phenotype and the risk of cutaneous melanoma at different latitudes. Int. J. Cancer 124, 420–428 (2009).
Cannon-Albright, L.A., Bishop, D.T., Goldgar, C. & Skolnick, M.H. Genetic predisposition to cancer. Important Adv. Oncol. 1991, 39–55 (1991).
Brown, K.M. et al. Common sequence variants on 20q11.22 confer melanoma susceptibility. Nat. Genet. 40, 838–840 (2008).
Bishop, D.T. et al. Genome-wide association study identifies three loci associated with melanoma risk. Nat. Genet. 41, 920–925 (2009).
Amos, C.I. et al. Genome-wide association study identifies novel loci predisposing to cutaneous melanoma. Hum. Mol. Genet. 20, 5012–5023 (2011).
Barrett, J.H. et al. Genome-wide association study identifies three new melanoma susceptibility loci. Nat. Genet. 43, 1108–1113 (2011).
Macgregor, S. et al. Genome-wide association study identifies a new melanoma susceptibility locus at 1q21.3. Nat. Genet. 43, 1114–1118 (2011).
Iles, M.M. et al. A variant in FTO shows association with melanoma risk not due to BMI. Nat. Genet. 45, 428–432 (2013).
Gudbjartsson, D.F. et al. ASIP and TYR pigmentation variants associate with cutaneous melanoma and basal cell carcinoma. Nat. Genet. 40, 886–891 (2008).
Antonopoulou, K. et al. Updated field synopsis and systematic meta-analyses of genetic association studies in cutaneous melanoma: the MelGene database. J. Invest. Dermatol. 135, 1074–1079 (2015).
Peña-Chilet, M. et al. Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish population. BMC Cancer 13, 160 (2013).
Falchi, M. et al. Genome-wide association study identifies variants at 9p21 and 22q13 associated with development of cutaneous nevi. Nat. Genet. 41, 915–919 (2009).
Rafnar, T. et al. Sequence variants at the TERT-CLPTM1L locus associate with many cancer types. Nat. Genet. 41, 221–227 (2009).
Pooley, K.A. et al. No association between TERT-CLPTM1L single nucleotide polymorphism rs401681 and mean telomere length or cancer risk. Cancer Epidemiol. Biomarkers Prev. 19, 1862–1865 (2010).
Nan, H., Qureshi, A.A., Prescott, J., De Vivo, I. & Han, J. Genetic variants in telomere-maintaining genes and skin cancer risk. Hum. Genet. 129, 247–253 (2011).
Law, M.H. et al. Meta-analysis combining new and existing data sets confirms that the TERT-CLPTM1L locus influences melanoma risk. J. Invest. Dermatol. 132, 485–487 (2012).
Mocellin, S. et al. Telomerase reverse transcriptase locus polymorphisms and cancer risk: a field synopsis and meta-analysis. J. Natl. Cancer Inst. 104, 840–854 (2012).
Gomez, M. et al. PARP1 is a TRF2-associated poly(ADP-ribose)polymerase and protects eroded telomeres. Mol. Biol. Cell 17, 1686–1696 (2006).
Derheimer, F.A. & Kastan, M.B. Multiple roles of ATM in monitoring and maintaining DNA integrity. FEBS Lett. 584, 3675–3681 (2010).
Bataille, V. et al. Nevus size and number are associated with telomere length and represent potential markers of a decreased senescence in vivo. Cancer Epidemiol. Biomarkers Prev. 16, 1499–1502 (2007).
Han, J. et al. A prospective study of telomere length and the risk of skin cancer. J. Invest. Dermatol. 129, 415–421 (2009).
Burke, L.S. et al. Telomere length and the risk of cutaneous malignant melanoma in melanoma-prone families with and without CDKN2A mutations. PLoS ONE 8, e71121 (2013).
Iles, M.M. et al. The effect on melanoma risk of genes previously associated with telomere length. J. Natl. Cancer Inst. 106, dju267 (2014).
Barrett, J.H. et al. Fine mapping of genetic susceptibility loci for melanoma reveals a mixture of single variant and multiple variant regions. Int. J. Cancer 136, 1351–1360 (2015).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Miyake, Y. et al. RPA-like mammalian Ctc1-Stn1-Ten1 complex binds to single-stranded DNA and protects telomeres independently of the Pot1 pathway. Mol. Cell 36, 193–206 (2009).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Robles-Espinoza, C.D. et al. POT1 loss-of-function variants predispose to familial melanoma. Nat. Genet. 46, 478–481 (2014).
Shi, J. et al. Rare missense variants in POT1 predispose to familial cutaneous malignant melanoma. Nat. Genet. 46, 482–486 (2014).
Codd, V. et al. Identification of seven loci affecting mean telomere length and their association with disease. Nat. Genet. 45, 422–427 (2013).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Innocenti, F. et al. Identification, replication, and functional fine-mapping of expression quantitative trait loci in primary human liver tissue. PLoS Genet. 7, e1002078 (2011).
Grundberg, E. et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat. Genet. 44, 1084–1089 (2012).
Gajjar, K., Martin-Hirsch, P.L. & Martin, F.L. CYP1B1 and hormone-induced cancer. Cancer Lett. 324, 13–30 (2012).
Muthusamy, V. et al. Epigenetic silencing of novel tumor suppressors in malignant melanoma. Cancer Res. 66, 11187–11193 (2006).
Shen, M. et al. Quantitative assessment of the influence of CYP1B1 polymorphisms and head and neck squamous cell carcinoma risk. Tumour Biol. 35, 3891–3897 (2014).
Stoilov, I., Akarsu, A.N. & Sarfarazi, M. Identification of three different truncating mutations in cytochrome P4501B1 (CYP1B1) as the principal cause of primary congenital glaucoma (Buphthalmos) in families linked to the GLC3A locus on chromosome 2p21. Hum. Mol. Genet. 6, 641–647 (1997).
Arragain, S. et al. Identification of eukaryotic and prokaryotic methylthiotransferase for biosynthesis of 2-methylthio-N6-threonylcarbamoyladenosine in tRNA. J. Biol. Chem. 285, 28425–28433 (2010).
Brambillasca, S. et al. CDK5 regulatory subunit–associated protein 1–like 1 (CDKAL1) is a tail-anchored protein in the endoplasmic reticulum (ER) of insulinoma cells. J. Biol. Chem. 287, 41808–41819 (2012).
Praetorius, C. et al. A polymorphism in IRF4 affects human pigmentation through a tyrosinase-dependent MITF/TFAP2A pathway. Cell 155, 1022–1033 (2013).
Sulem, P. et al. Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat. Genet. 39, 1443–1452 (2007).
Han, J. et al. A genome-wide association study identifies novel alleles associated with hair color and skin pigmentation. PLoS Genet. 4, e1000074 (2008).
Duffy, D.L. et al. IRF4 variants have age-specific effects on nevus count and predispose to melanoma. Am. J. Hum. Genet. 87, 6–16 (2010).
Zhang, M. et al. Genome-wide association studies identify several new loci associated with pigmentation traits and skin cancer risk in European Americans. Hum. Mol. Genet. 22, 2948–2959 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Persson, S. et al. Diversity of the protein disulfide isomerase family: identification of breast tumor induced Hag2 and Hag3 as novel members of the protein family. Mol. Phylogenet. Evol. 36, 734–740 (2005).
Fletcher, G.C. et al. hAG-2 and hAG-3, human homologues of genes involved in differentiation, are associated with oestrogen receptor–positive breast tumours and interact with metastasis gene C4.4a and dystroglycan. Br. J. Cancer 88, 579–585 (2003).
King, E.R. et al. The anterior gradient homolog 3 (AGR3) gene is associated with differentiation and survival in ovarian cancer. Am. J. Surg. Pathol. 35, 904–912 (2011).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Masutani, C. et al. Purification and cloning of a nucleotide excision repair complex involving the xeroderma pigmentosum group C protein and a human homologue of yeast RAD23. EMBO J. 13, 1831–1843 (1994).
Xia, Y. et al. TAL2, a helix-loop-helix gene activated by the (7;9)(q34;q32) translocation in human T-cell leukemia. Proc. Natl. Acad. Sci. USA 88, 11416–11420 (1991).
Wong, C.W. et al. Kruppel-like transcription factor 4 contributes to maintenance of telomerase activity in stem cells. Stem Cells 28, 1510–1517 (2010).
Hoffmeyer, K. et al. Wnt/β-catenin signaling regulates telomerase in stem cells and cancer cells. Science 336, 1549–1554 (2012).
Teerlink, C. et al. A unique genome-wide association analysis in extended Utah high-risk pedigrees identifies a novel melanoma risk variant on chromosome arm 10q. Hum. Genet. 131, 77–85 (2012).
Vízkeleti, L. et al. The role of CCND1 alterations during the progression of cutaneous malignant melanoma. Tumour Biol. 33, 2189–2199 (2012).
Young, R.J. et al. Loss of CDKN2A expression is a frequent event in primary invasive melanoma and correlates with sensitivity to the CDK4/6 inhibitor PD0332991 in melanoma cell lines. Pigment Cell Melanoma Res. 27, 590–600 (2014).
French, J.D. et al. Functional variants at the 11q13 risk locus for breast cancer regulate cyclin D1 expression through long-range enhancers. Am. J. Hum. Genet. 92, 489–503 (2013).
Duffy, D.L. et al. A three–single-nucleotide polymorphism haplotype in intron 1 of OCA2 explains most human eye-color variation. Am. J. Hum. Genet. 80, 241–252 (2007).
Ruiz, Y. et al. Further development of forensic eye color predictive tests. Forensic Sci. Int. Genet. 7, 28–40 (2013).
Li, M.X., Yeung, J.M., Cherny, S.S. & Sham, P.C. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum. Genet. 131, 747–756 (2012).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Li, Y., Willer, C., Sanna, S. & Abecasis, G. Genotype imputation. Annu. Rev. Genomics Hum. Genet. 10, 387–406 (2009).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Higgins, J.P. & Thompson, S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 21, 1539–1558 (2002).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control. Clin. Trials 7, 177–188 (1986).
Mägi, R. & Morris, A.P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Gabriel, S., Ziaugra, L. & Tabbaa, D. SNP genotyping using the Sequenom MassARRAY iPLEX platform. Curr. Protoc. Hum. Genet. Chapter 2, Unit 2.12 (2009).
Cho, E., Rosner, B.A., Feskanich, D. & Colditz, G.A. Risk factors and individual probabilities of melanoma for whites. J. Clin. Oncol. 23, 2669–2675 (2005).
Newton-Bishop, J.A. et al. Melanocytic nevi, nevus genes, and melanoma risk in a large case-control study in the United Kingdom. Cancer Epidemiol. Biomarkers Prev. 19, 2043–2054 (2010).
Newton-Bishop, J.A. et al. Relationship between sun exposure and melanoma risk for tumours in different body sites in a large case-control study in a temperate climate. Eur. J. Cancer 47, 732–741 (2011).
Newton-Bishop, J.A. et al. Serum 25-hydroxyvitamin D3 levels are associated with Breslow thickness at presentation and survival from melanoma. J. Clin. Oncol. 27, 5439–5444 (2009).
Edwards, S.L., Beesley, J., French, J.D. & Dunning, A.M. Beyond GWASs: illuminating the dark road from association to function. Am. J. Hum. Genet. 93, 779–797 (2013).
Nica, A.C. et al. The architecture of gene regulatory variation across multiple human tissues: the MuTHER study. PLoS Genet. 7, e1002003 (2011).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Acknowledgements
Please see the Supplementary Note for acknowledgments.
Author information
Authors and Affiliations
Consortia
Contributions
M.M.I. and M.H.L. led, designed and carried out the statistical analyses and wrote the manuscript. M. Harland was involved in the Leeds genotyping design. J.C.T. carried out statistical analyses. J.R.-M. and N.v.d.S. carried out genotyping and contributed to the interpretation of genotyping data. J.A.N.B. led the GenoMEL Consortium and contributed to study design. N.A.G. was deputy lead of the consortium and contributed to study design. S.M., N.K.H., D.T.B. and J.H.B. designed and led the overall study. J. Han supervised and carried out statistical analysis of the Harvard GWAS data. F.S. and A.A.Q. carried out statistical analysis of the Harvard GWAS data. C.I.A. led and carried out statistical analysis of the MD Anderson GWAS data. W.V.C., J.E.L. and S.F. contributed to the analysis and interpretation of the MD Anderson GWAS data. F.D. led, designed and contributed to the sample collection, analysis and interpretation of the French MELARISK GWAS and advised on the overall statistical analysis. M.B. contributed to the analysis and interpretation of the French MELARISK GWAS data. M.-F.A. led, designed and contributed to the sample collection of the French MELARISK GWAS. G.M.L. led and contributed to the genotyping and interpretation in the French MELARISK GWAS. R.K. and D.S. led and contributed to the sample collection and analysis for the Heidelberg data set. H.-J.S. contributed to the sample collection and analysis for the Heidelberg data set. S.V.W. led and contributed to the sample collection for the WAMHS study. E.K.M. provided coordination and oversight for the WAMHS study. D.C.W. led, designed and contributed to the sample collection for the SDH data set. J.E.C. led and designed the Glaucoma study. K.P.B. contributed to the analysis and interpretation of the Glaucoma data set. G.L.R.-S. led and contributed to the analysis and interpretation of the IBD data set. L.A.S. contributed to the analysis and interpretation of the IBD data set. G.J.M. led and contributed to the sample collection, analysis and interpretation of the AMFS study. A.E.C. contributed to the sample collection, analysis and interpretation of the AMFS study. D.R.N. contributed to the sample collection and analysis of the Q-MEGA, Endometriosis and QTWIN data sets. N.G.M. led the sample collection and analysis for the Q-MEGA and QTWIN data sets. G.W.M. led the sample collection and analysis for the Endometriosis data sets and contributed to the sample collection and analysis for the Q-MEGA, Endometriosis and QTWIN data sets. D.L.D. contributed to the sample collection and analysis for the Q-MEGA, Endometriosis and QTWIN data sets. K.M.B. contributed to the sample collection and analysis for the Q-MEGA and QTWIN data sets. A.J. Stratigos and K.P.K. interpreted and contributed genotype data for the Athens stage 2 data set. A.M.G., P.A.K. and E.M.G. advised on statistical analysis. D.E.E. contributed to the design of the GenoMEL GWAS. A.J. Swerdlow and N.O. interpreted and contributed genotype data for the Breakthrough Generations Study. L.A.A., P.A.A., E.A., G.B.S., T.D., E.F., P. Ghiorzo, J. Hansson, P.H., M. Hocˇevar, V.H., C.I., M.T.L., J. Lang, R.M.M., A.M., J. Lubin´ski, S.N., H.O., S.P., J.A.P.-B. and R.v.D. contributed to sample collection, analysis and interpretation for the GenoMEL data sets. K.A.P., A.M.D., P.D.P.P. and D.F.E. interpreted and contributed genotype data for the Cambridge stage 2 data set. P. Galan contributed to the collection, analysis and interpretation of the SU.VI.Max French control data set. All authors provided critical review of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3, 5–8 and 10, and Supplementary Note. (PDF 4182 kb)
Supplementary Table 4
List of SNPs reaching P < 1 × 10–7. (XLSX 115 kb)
Supplementary Table 9
SNPs used in bioinformatics annotation. (XLSX 19 kb)
Rights and permissions
About this article
Cite this article
Law, M., Bishop, D., Lee, J. et al. Genome-wide meta-analysis identifies five new susceptibility loci for cutaneous malignant melanoma. Nat Genet 47, 987–995 (2015). https://doi.org/10.1038/ng.3373
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3373
This article is cited by
-
Early contribution of germline and nevi genetic alterations to a rapidly-progressing cutaneous melanoma patient: a case report
BMC Medical Genomics (2023)
-
Identification of genomic-wide genetic links between cutaneous melanoma and obesity-related physical traits via cFDR
Genes & Genomics (2023)
-
Implementation of individualised polygenic risk score analysis: a test case of a family of four
BMC Medical Genomics (2022)
-
Genome-wide association study of actinic keratosis identifies new susceptibility loci implicated in pigmentation and immune regulation pathways
Communications Biology (2022)
-
First international workshop of the ATM and cancer risk group (4-5 December 2019)
Familial Cancer (2022)