Abstract
The major pathway by which the brain obtains essential omega-3 fatty acids from the circulation is through a sodium-dependent lysophosphatidylcholine (LPC) transporter (MFSD2A), expressed in the endothelium of the blood-brain barrier. Here we show that a homozygous mutation affecting a highly conserved MFSD2A residue (p.Ser339Leu) is associated with a progressive microcephaly syndrome characterized by intellectual disability, spasticity and absent speech. We show that the p.Ser339Leu alteration does not affect protein or cell surface expression but rather significantly reduces, although not completely abolishes, transporter activity. Notably, affected individuals displayed significantly increased plasma concentrations of LPCs containing mono- and polyunsaturated fatty acyl chains, indicative of reduced brain uptake, confirming the specificity of MFSD2A for LPCs having mono- and polyunsaturated fatty acyl chains. Together, these findings indicate an essential role for LPCs in human brain development and function and provide the first description of disease associated with aberrant brain LPC transport in humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Nguyen, L.N. et al. Mfsd2a is a transporter for the essential omega-3 fatty acid docosahexaenoic acid. Nature 509, 503–506 (2014).
Ethayathulla, A.S. et al. Structure-based mechanism for Na+/melibiose symport by MelB. Nat. Commun. 5, 3009 (2014).
Berger, J.H., Charron, M.J. & Silver, D.L. Major facilitator superfamily domain–containing protein 2a (MFSD2A) has roles in body growth, motor function, and lipid metabolism. PLoS ONE 7, e50629 (2012).
Guemez-Gamboa, A. et al. Inactivating mutations in MFSD2A, required for omega-3 fatty acid transport in brain, cause a lethal microcephaly syndrome. Nat. Genet. doi:10.1038/ng.3311 (25 May 2015).
Baisted, D.J., Robinson, B.S. & Vance, D.E. Albumin stimulates the release of lysophosphatidylcholine from cultured rat hepatocytes. Biochem. J. 253, 693–701 (1988).
Croset, M., Brossard, N., Polette, A. & Lagarde, M. Characterization of plasma unsaturated lysophosphatidylcholines in human and rat. Biochem. J. 345, 61–67 (2000).
Innis, S.M. Dietary (n-3) fatty acids and brain development. J. Nutr. 137, 855–859 (2007).
Shindou, H., Hishikawa, D., Harayama, T., Eto, M. & Shimizu, T. Generation of membrane diversity by lysophospholipid acyltransferases. J. Biochem. 154, 21–28 (2013).
Sobel, E., Sengul, H. & Weeks, D.E. Multipoint estimation of identity-by-descent probabilities at arbitrary positions among marker loci on general pedigrees. Hum. Hered. 52, 121–131 (2001).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000).
Ben-Zvi, A. et al. Mfsd2a is critical for the formation and function of the blood-brain barrier. Nature 509, 507–511 (2014).
Zhao, Z. & Xu, Y. An extremely simple method for extraction of lysophospholipids and phospholipids from blood samples. J. Lipid Res. 51, 652–659 (2010).
Acknowledgements
We are grateful to the families for participating in this research. The work was supported by UK Medical Research Council grants G1002279, G0900205 and G1001931 and by the Newlife Foundation for disabled children (to A.H.C., M.A.P. and E.L.B.), the Singapore Ministry of Health's National Medical Research Council grant CBRG/069/2014 (to D.L.S.), Singapore National Research Foundation Competitive Research Program grant 2007-04 (to M.R.W.) and the National University of Singapore's Life Sciences Institute (to M.R.W.).
Author information
Authors and Affiliations
Contributions
A.H.C. and D.L.S. conceived, designed and supervised the project. V.A., D.Q.Y.Q., B.A.C., A.C.-G., L.N.N., M.R.W. and M.N.W. performed experiments. A.H. aided in family recruitment. A.H., A.Q.A., T.T.W., M.A.P., P.R. and E.L.B. aided in assessment of clinical data. A.S.-N. was involved in the genetic studies. A.H.C., D.L.S. and E.L.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Homozygosity map of affected family members.
Homozygosity plots indicate detected autozygous genomic regions shared by affected family members in each family, with the homozygous region containing the MFSD2A region indicated in red.
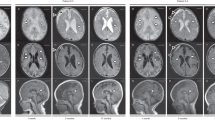
Supplementary Figure 2 Facial features of affected family members.
Features include microcephaly and up-slanting palpebral fissures (a, II:3; b, III:2; c, III:3; d, III:4). Published with consent.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, and Supplementary Tables 1 and 2. (PDF 240 kb)
Rights and permissions
About this article
Cite this article
Alakbarzade, V., Hameed, A., Quek, D. et al. A partially inactivating mutation in the sodium-dependent lysophosphatidylcholine transporter MFSD2A causes a non-lethal microcephaly syndrome. Nat Genet 47, 814–817 (2015). https://doi.org/10.1038/ng.3313
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3313
This article is cited by
-
Substrate binding-induced conformational transitions in the omega-3 fatty acid transporter MFSD2A
Nature Communications (2023)
-
The Role of Major Facilitator Superfamily Domain-Containing 2a in the Central Nervous System
Cellular and Molecular Neurobiology (2023)
-
Brain metastasis in breast cancer: focus on genes and signaling pathways involved, blood–brain barrier and treatment strategies
Clinical and Translational Oncology (2023)
-
Structural insights into the lysophospholipid brain uptake mechanism and its inhibition by syncytin-2
Nature Structural & Molecular Biology (2022)
-
Blood–brain barrier link to human cognitive impairment and Alzheimer’s disease
Nature Cardiovascular Research (2022)