Abstract
Shoot meristems of plants are composed of stem cells that are continuously replenished through a classical feedback circuit involving the homeobox WUSCHEL (WUS) gene and the CLAVATA (CLV) gene signaling pathway. In CLV signaling, the CLV1 receptor complex is bound by CLV3, a secreted peptide modified with sugars. However, the pathway responsible for modifying CLV3 and its relevance for CLV signaling are unknown. Here we show that tomato inflorescence branching mutants with extra flower and fruit organs due to enlarged meristems are defective in arabinosyltransferase genes. The most extreme mutant is disrupted in a hydroxyproline O-arabinosyltransferase and can be rescued with arabinosylated CLV3. Weaker mutants are defective in arabinosyltransferases that extend arabinose chains, indicating that CLV3 must be fully arabinosylated to maintain meristem size. Finally, we show that a mutation in CLV3 increased fruit size during domestication. Our findings uncover a new layer of complexity in the control of plant stem cell proliferation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Rickett, H.W. The classification of inflorescences. Bot. Rev. 10, 187–231 (1944).
Prusinkiewicz, P., Erasmus, Y., Lane, B., Harder, L.D. & Coen, E. Evolution and development of inflorescence architectures. Science 316, 1452–1456 (2007).
Park, S.J., Eshed, Y. & Lippman, Z.B. Meristem maturation and inflorescence architecture—lessons from the Solanaceae. Curr. Opin. Plant Biol. 17, 70–77 (2014).
Bommert, P., Nagasawa, N.S. & Jackson, D. Quantitative variation in maize kernel row number is controlled by the FASCIATED EAR2 locus. Nat. Genet. 45, 334–337 (2013).
Doebley, J. The genetics of maize evolution. Annu. Rev. Genet. 38, 37–59 (2004).
Barton, M.K. Twenty years on: the inner workings of the shoot apical meristem, a developmental dynamo. Dev. Biol. 341, 95–113 (2010).
Clark, S.E., Williams, R.W. & Meyerowitz, E.M. The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 89, 575–585 (1997).
Fletcher, J.C., Brand, U., Running, M.P., Simon, R. & Meyerowitz, E.M. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 283, 1911–1914 (1999).
Ohyama, K., Shinohara, H., Ogawa-Ohnishi, M. & Matsubayashi, Y. A glycopeptide regulating stem cell fate in Arabidopsis thaliana. Nat. Chem. Biol. 5, 578–580 (2009).
Stahl, Y. & Simon, R. Plant primary meristems: shared functions and regulatory mechanisms. Curr. Opin. Plant Biol. 13, 53–58 (2010).
Schoof, H. et al. The stem cell population of Arabidopsis shoot meristems in maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 100, 635–644 (2000).
Pautler, M., Tanaka, W., Hirano, H.Y. & Jackson, D. Grass meristems I: shoot apical meristem maintenance, axillary meristem determinacy and the floral transition. Plant Cell Physiol. 54, 302–312 (2013).
Menda, N., Semel, Y., Peled, D., Eshed, Y. & Zamir, D. In silico screening of a saturated mutation library of tomato. Plant J. 38, 861–872 (2004).
Lippman, Z.B. et al. The making of a compound inflorescence in tomato and related nightshades. PLoS Biol. 6, e288 (2008).
Jeong, S., Trotochaud, A.E. & Clark, S.E. The Arabidopsis CLAVATA2 gene encodes a receptor-like protein required for the stability of the CLAVATA1 receptor-like kinase. Plant Cell 11, 1925–1934 (1999).
Muños, S. et al. Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL. Plant Physiol. 156, 2244–2254 (2011).
Reinhardt, D., Frenz, M., Mandel, T. & Kuhlemeier, C. Microsurgical and laser ablation analysis of interactions between the zones and layers of the tomato shoot apical meristem. Development 130, 4073–4083 (2003).
Zhang, Y., Yang, S., Song, Y. & Wang, J. Genome-wide characterization, expression and functional analysis of CLV3/ESR gene family in tomato. BMC Genomics 15, 827 (2014).
Diévart, A. et al. CLAVATA1 dominant-negative alleles reveal functional overlap between multiple receptor kinases that regulate meristem and organ development. Plant Cell 15, 1198–1211 (2003).
Ogawa-Ohnishi, M., Matsushita, W. & Matsubayashi, Y. Identification of three hydroxyproline O-arabinosyltransferases in Arabidopsis thaliana. Nat. Chem. Biol. 9, 726–730 (2013).
Kieliszewski, M.J. & Lamport, D.T. Extensin: repetitive motifs, functional sites, post-translational codes, and phylogeny. Plant J. 5, 157–172 (1994).
Lamport, D.T., Kieliszewski, M.J., Chen, Y. & Cannon, M.C. Role of the extensin superfamily in primary cell wall architecture. Plant Physiol. 156, 11–19 (2011).
Cannon, M.C. et al. Self-assembly of the plant cell wall requires an extensin scaffold. Proc. Natl. Acad. Sci. USA 105, 2226–2231 (2008).
Matsubayashi, Y. Small post-translationally modified peptide signals in. Arabidopsis. Arabidopsis Book 9, e0150 (2011).
Xu, T.T., Song, X.F., Ren, S.C. & Liu, C.M. The sequence flanking the N-terminus of the CLV3 peptide is critical for its cleavage and activity in stem cell regulation in Arabidopsis. BMC Plant Biol. 13, 225 (2013).
Shinohara, H. & Matsubayashi, Y. Chemical synthesis of Arabidopsis CLV3 glycopeptide reveals the impact of hydroxyproline arabinosylation on peptide conformation and activity. Plant Cell Physiol. 54, 369–374 (2013).
Doudna, J.A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Belhaj, K., Chaparro-Garcia, A., Kamoun, S., Patron, N.J. & Nekrasov, V. Editing plant genomes with CRISPR/Cas9. Curr. Opin. Biotechnol. 32, 76–84 (2015).
Brooks, C., Nekrasov, V., Lippman, Z.B. & Van Eck, J. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 166, 1292–1297 (2014).
Nimchuk, Z.L., Tarr, P.T. & Meyerowitz, E.M. An evolutionarily conserved pseudokinase mediates stem cell production in plants. Plant Cell 23, 851–854 (2011).
Gille, S., Hansel, U., Ziemann, M. & Pauly, M. Identification of plant cell wall mutants by means of a forward chemical genetic approach using hydrolases. Proc. Natl. Acad. Sci. USA 106, 14699–14704 (2009).
Velasquez, S.M. et al. O-glycosylated cell wall proteins are essential in root hair growth. Science 332, 1401–1403 (2011).
Egelund, J. et al. Molecular characterization of two Arabidopsis thaliana glycosyltransferase mutants, rra1 and rra2, which have a reduced residual arabinose content in a polymer tightly associated with the cellulosic wall residue. Plant Mol. Biol. 64, 439–451 (2007).
Tanksley, S.D. The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 16 (suppl.), S181–S189 (2004).
Barrero, L.S. & Tanksley, S.D. Evaluating the genetic basis of multiple-locule fruit in a broad cross section of tomato cultivars. Theor. Appl. Genet. 109, 669–679 (2004).
Lippman, Z. & Tanksley, S.D. Dissecting the genetic pathway to extreme fruit size in tomato using a cross between the small-fruited wild species Lycopersicon pimpinellifolium and L. esculentum var. Giant Heirloom. Genetics 158, 413–422 (2001).
van der Knaap, E. et al. What lies beyond the eye: the molecular mechanisms regulating tomato fruit weight and shape. Front. Plant Sci. 5, 227 (2014).
Cong, B., Barrero, L.S. & Tanksley, S.D. Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication. Nat. Genet. 40, 800–804 (2008).
Huang, Z. & van der Knaap, E. Tomato fruit weight 11.3 maps close to fasciated on the bottom of chromosome 11. Theor. Appl. Genet. 123, 465–474 (2011).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P.M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Yin, Y., Mohhen, D., Gelineo-Albersheim, I., Xu, Y. & Hahn, M. in Annual Plant Reviews: Plant Polysaccharides: Biosynthesis and Bioengineering (ed. Ulvskov, P.) (Wiley-Black, 2011).
Petersen, B.L., Faber, K. & Ulvskov, P. in Annual Plant Reviews: Plant Polysaccharides: Biosynthesis and Bioengineering (ed. Ulvskov, P.) (Wiley-Black, 2011).
Schnabel, E.L. et al. The ROOT DETERMINED NODULATION1 gene regulates nodule number in roots of Medicago truncatula and defines a highly conserved, uncharacterized plant gene family. Plant Physiol. 157, 328–340 (2011).
Müller, R., Borghi, L., Kwiatkowska, D., Laufs, P. & Simon, R. Dynamic and compensatory responses of Arabidopsis shoot and floral meristems to CLV3 signaling. Plant Cell 18, 1188–1198 (2006).
Betsuyaku, S. et al. Mitogen-activated protein kinase regulated by the CLAVATA receptors contributes to shoot apical meristem homeostasis. Plant Cell Physiol. 52, 14–29 (2011).
DeYoung, B.J. et al. The CLAVATA1-related BAM1, BAM2 and BAM3 receptor kinase-like proteins are required for meristem function in. Arabidopsis. Plant J. 45, 1–16 (2006).
DeYoung, B.J. & Clark, S.E. BAM receptors regulate stem cell specification and organ development through complex interactions with CLAVATA signaling. Genetics 180, 895–904 (2008).
Kinoshita, A. et al. RPK2 is an essential receptor-like kinase that transmits the CLV3 signal in Arabidopsis. Development 137, 3911–3920 (2010).
Müller, R., Bleckmann, A. & Simon, R. The receptor kinase CORYNE of Arabidopsis transmits the stem cell–limiting signal CLAVATA3 independently of CLAVATA1. Plant Cell 20, 934–946 (2008).
Nimchuk, Z.L., Zhou, Y., Tarr, P.T., Petersen, B.A. & Meyerowitz, E.M. Plant stem cell maintenance by transcriptional cross-regulation of related receptor kinases. Development 142, 1043–1049 (2015).
Fan, C. et al. A novel single-nucleotide mutation in a CLAVATA3 gene homolog controls a multilocular silique trait in Brassica rapa L. Mol. Plant 7, 1788–1792 (2014).
Park, S.J., Jiang, K., Schatz, M.C. & Lippman, Z.B. Rate of meristem maturation determines inflorescence architecture in tomato. Proc. Natl. Acad. Sci. USA 109, 639–644 (2012).
Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Jackson, D.P. in Molecular Plant Pathology: A Practical Approach (eds. Gurr, S.J., Bowles, D.J. & McPherson, M.J.) 163–174 (Oxford University Press, 1992).
Saint-Jore-Dupas, C. et al. Plant N-glycan processing enzymes employ different targeting mechanisms for their spatial arrangement along the secretory pathway. Plant Cell 18, 3182–3200 (2006).
Chevalier, L. et al. Subcompartment localization of the side chain xyloglucan-synthesizing enzymes within Golgi stacks of tobacco suspension-cultured cells. Plant J. 64, 977–989 (2010).
Nelson, B.K., Cai, X. & Nebenfuhr, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Xu, C. et al. Degradation of MONOCULM 1 by APC/CTAD1 regulates rice tillering. Nat. Commun. 3, 750 (2012).
Yoo, S.D., Cho, Y.H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Xiong, G., Cheng, K. & Pauly, M. Xylan O-acetylation impacts xylem development and enzymatic recalcitrance as indicated by the Arabidopsis mutant tbl29. Mol. Plant 6, 1373–1375 (2013).
York, W.S., Darvill, A.G., McNeil, M. & Albersheim, P. 3-deoxy-D-manno-2-octulosonic acid (KDO) is a component of rhamnogalacturonan II, a pectic polysaccharide in the primary cell walls of plants. Carbohydr. Res. 138, 109–126 (1985).
Fiers, M. et al. The CLAVATA3/ESR motif of CLAVATA3 is functionally independent from the nonconserved flanking sequences. Plant Physiol. 141, 1284–1292 (2006).
Acknowledgements
We thank members of the Lippman laboratory, especially S. Thomain for her initial finding of FAB2 and for invaluable conversations that helped shape this work. We thank D. Zamir (Hebrew University of Jerusalem) for providing mutants and also Y. Eshed (Weizmann Institute of Science) for providing mutants and comments on the manuscript. We thank S. Hearn at the Cold Spring Harbor Laboratory St. Giles Advanced Microscopy Center for providing technical service for transmission electron microscopy. We thank W. Wang for assistance with tomato transformation, X. Song for advice on peptide assays, DuPont Pioneer for research support, and T. Mulligan, A. Krainer and staff from Cornell University's Long Island Horticultural Research and Extension Center in Riverhead, New York, for assistance with plant care. This research was supported by the Energy Biosciences Institute and the Fred Dickinson Chair for M.P., a National Science Foundation Graduate Research Fellowship (DGE-0914548) to K.L.L., a Gordon and Betty Moore Foundation Fellowship from the Life Sciences Research Foundation to C.A.M., grants from the National Science Foundation Plant Genome Research Program to E.v.d.K. (0922661) and to J.V.E. and Z.B.L. (1237880), and an Agriculture and Food Research Initiative competitive grant (2015-67013-22823) of the US Department of Agriculture National Institute of Food and Agriculture to Z.B.L.
Author information
Authors and Affiliations
Contributions
C.X., K.L.L., C.A.M., Z.H., Y.-H.C., M.P., J.V.E., Y.M., E.v.d.K. and Z.B.L. designed and planned experiments. C.X., K.L.L., C.A.M., Z.H., Y.-H.C., C.B., M.O.-O., G.X. and Z.B.L. performed experiments and collected the data. C.X., K.L.L., C.A.M., Z.H., Y.-H.C., K.J., C.B., M.O.-O., G.X., M.P., Y.M., E.v.d.K. and Z.B.L. analyzed the data. K.L.L., C.X., Y.M., E.v.d.K. and Z.B.L. designed the research. C.X., K.L.L. and Z.B.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors (C.A.M., K.L.L. and Z.B.L., on behalf of Cold Spring Harbor Laboratory and DuPont Pioneer) have filed a PCT patent application based in part on this work with the US Patent and Trademark Office.
Integrated supplementary information
Supplementary Figure 1 The FIN and FAB genes act separately from the meristem maturation pathway.
(a) Representative inflorescence from the compound inflorescence (s, defective in the homolog of Arabidopsis WUSCHEL HOMEOBOX 9)14 meristem maturation mutant showing extreme branching. (b,c) Double mutants of fab s (b) and fin s (c) show additive genetic relationships. Double mutants exhibit branching like the s single mutant and fasciation of the flowers like the fin and fab single mutants. Red arrows in b show greater branching in fab s double mutants compared to fab mutants alone. The inset shows a fasciated flower. (d) Representative highly branched inflorescences with cauliflower-like flowers from the anantha (an, defective in the homolog of the Arabidopsis F-box gene UNUSUAL FLORAL ORGANS)14 meristem maturation mutant. (e,f) fab an (e) and fin an (f) double mutants show additive phenotypes. Yellow arrows point to the base of each inflorescence. Insets with red circles show normal-sized meristems in an single mutants (d) and fasciated meristems in fab an (e) and fin an (f) double mutants. Scale bars, 1 cm.
Supplementary Figure 2 Stereoscope and scanning electron micrograph (SEM) imaging showing enlarged meristems of the fab and fin mutants at different stages.
(a–c) Stereoscope images from the primary-shoot apex comparing the SAM at the early vegetative meristem (EVM, fifth leaf initiated) stage from WT (a), fab (b) and fin (c). Dashed lines mark the height and width dimensions used for meristem size measurements. (d) Quantification and statistical comparisons of EVM size from WT, fab and fin. Data are means ± s.d., n = 8–21. A two-tailed, two-sample t test was performed, and significant differences are represented by black asterisks: *P < 0.01. (e–h) SEM images showing the enlarged transition meristem (TM) stage of WT (e) compared to fab (f) and fin (g,h). (i,j) Two stages of sympodial inflorescence development in WT plants. Multi-flowered unbranched inflorescences arise from sympodial inflorescence meristems (SIM), each of which initiates one SIM before terminating (i)14. A sympodial shoot meristem (SYM) from the last PSM leaf produces a sympodial shoot with three leaves and the next inflorescence, and initiation of a new SYM from each prior sympodial shoot perpetuates growth (j)14. (k,l) The enlarged SAM of fin at the TM stage allowed additional leaf primordia to develop at its periphery, resulting in multiple SYMs (k). Likewise, multiple SIMs developed as the SAM terminated in a large fasciated flower (l). Orange arrowheads indicate extra SYMs and SIMs. L, leaf. Scale bars, 100 μm.
Supplementary Figure 3 fab fin double mutant plants show enhanced fasciation.
(a) Image of a fab fin double mutant plant. The red arrow points to the primary inflorescence. White arrows mark axillary shoots. Note the absence of inflorescences and flowers. (b) Close-up image of the primary shoot and primary inflorescence showing enhanced stem and meristem fasciation (red arrow). (c,d) Stereoscopic images showing extreme enlargement of the primary shoot meristem at the transition stage (c) and after the supposed transition to flowering (d). Note the proliferation of ectopic meristems at the meristem flanks. After several weeks, fab fin double mutants eventually produce a few inflorescences and flowers that are more fasciated than those of fin alone.
Supplementary Figure 4 The tomato CLV3/EMBRYO-SURROUNDING REGION (CLE) peptide family.
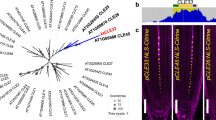
(a) Alignment of the dodecapeptide motifs from 15 tomato SlCLE proteins18. Residues are colored on the basis of their physiochemical properties, and consensus symbols are shown at the top of the sequences indicating positions with strongly similar properties (:) and weakly similar properties (.) with the Arabidopsis thaliana CLV3 dodecapeptide sequence shown for reference. Ten SlCLE peptides start with arginine (R), characteristic of the CLV3 peptide class, and five others start with histidine (H), characteristic of the TDIF peptide class64. (b) Neighbor-joining phylogenetic tree of the tomato and Arabidopsis CLE peptide family. CLE dodecapeptides from tomato and Arabidopsis were aligned using ClustalW, and a tree was constructed using the FigTree v1.3 program. The tree is shown with 100 replicate bootstrap values indicated at each node. Red color highlights tomato CLEs; black arrows point to SlCLV3, SlCLE3 and SlCLE9, also in bold. CLE family genes were retrieved from AGRIS (http://arabidopsis.med.ohio-state.edu/) and from the Solanaceae Genomics Network (SGN; http://solgenomics.net). (c–e) Normalized RNA-seq read counts (RPKM) from our tomato gene expression atlas (http://tomatolab/~lippmanlab2/allexp_query.html)52 showing that SlCLV3 (c) and SlCLE9 (e) are expressed primarily in meristems, unlike SlCLE3 (d), which is expressed broadly. Data from the EVM and floral meristem (FM) stages are shown52.
Supplementary Figure 5 Phylogenetic trees of the FIN/HPAT and FAB/CLV1 protein families in tomato and Arabidopsis.
(a) An unrooted maximum-parsimony phylogenetic tree from a subset of leucine-rich repeat (LRR) receptor–like proteins most similar to FAB/CLV1 from tomato and Arabidopsis. Red font indicates tomato, and blue font indicates Arabidopsis. (b) mRNA in situ hybridization on a WT tomato vegetative meristem showing accumulation of FAB transcripts in the L3 meristem layer. Scale bar, 100 μm. L5, fifth leaf. Maximum-parsimony consensus trees are shown with 100 replicate bootstrap values indicated at each node. (c) An unrooted maximum-parsimony phylogenetic tree of the FIN/HPAT family based on complete predicted protein-coding sequences. Red font indicates tomato, and blue font indicates Arabidopsis. FIN is most similar to HPAT3. (d) RT-PCR showing that FIN transcripts are absent in three deletion alleles of the fin mutant (Fig. 2e). The UBIQUITIN (UBI) gene served as an internal control.
Supplementary Figure 6 Quantification of floral organ numbers from fab/+ heterozygous plants.
fab/+ heterozygotes show weak fasciation due to a weak dominant-negative effect similar to the classical Arabidopsis clv1-9 mutant allele, which is based on the same amino acid change as fab (Fig. 2d)19. Data are presented as means ± s.d, n = 29–70. Significant differences were calculated using a Tukey-Kramer HSD test. ***P < 0.0001, **P < 0.001. *P < 0.01; n.s., not significant.
Supplementary Figure 7 Transmission electron microscopy of meristems and cell walls from WT, fin and fab meristems.
Progressive zooming in on meristems, cells and cell walls from WT (a,d,g), fin (b,e,f) and fab (c,f,h) plants. No differences in cell wall thickness (red arrowheads) were observed between the three genotypes. Scale bars, 10 μm (a–c), 2 μm (d–f), 0.2 μm (g,h).
Supplementary Figure 8 CRISPR/Cas9-engineered mutations in SlCLV3 result in fasciated plants resembling fin.
(a) Schematic illustrating two guide RNAs (sgRNAs; red arrows) targeting the SlCLV3 coding sequence. Cas9/sgRNA1/sgRNA2 were expressed from the same construct28,29. Black arrows indicate the PCR primers used to evaluate mutation type and efficiency. (b) PCR genotyping of seven first-generation (T0) CRISPR/Cas9-slclv3 (CR-slclv3) plants showing the presence of indel mutations. PCR for the presence of Cas9 is shown along with evaluation for fasciation. All four T0 CR-slclv3 fasciated plants exhibited similar severity and resembled fin. (c) Representative primary inflorescence from a CRISPR/Cas9-slclv3 plant (CR-slclv3-7) showing branching (red arrowheads) and a large fasciated first flower and fruit with extra locules (white arrowheads) similar to fin mutants. Scale bar, 1 cm. (d) Quantification and statistical comparisons of floral organ numbers from WT and CR-slclv3 flowers. Data were collected from three plants with confirmed mutations in all sequenced alleles (e; see also Online Methods). Data are means ± s.d., n ≥ 10. A two-tailed, two-sample t test was performed, and significant differences are represented by black asterisks: ***P < 0.0001. (e) CR-slclv3 alleles identified by cloning and sequencing PCR products from the SlCLV3 targeted region from three fasciated T0 plants. All alleles carried indel mutations (blue dashed lines and letters), and one of each representative allele identified from sequencing at least eight cloned PCR products is shown. CR-clv3-7 is biallelic, and the other two plants are chimeric. Red font highlights sgRNA target sites, and black rectangles indicate protospacer-adjacent motif (PAM) sequences; a, allele.
Supplementary Figure 9 CRISPR/Cas9-engineered mutations in FAB result in weakly fasciated plants resembling fab mutants.
(a) Schematic illustrating two guide RNAs (sgRNAs; red arrows) targeting the FAB (SlCLV1) coding sequence. Cas9/sgRNA1/sgRNA2 were expressed from the same construct. Black arrows indicate the PCR primers used to evaluate mutation type and efficiency. (b) PCR genotyping of eight T0 CR-fab plants showing predominantly chimeric plants with a range of indel mutations, including the desired deletion. One plant (CR-fab-8) was homozygous for the desired deletion. PCR for the presence of Cas9 is shown along with evaluation for fasciation. All eight CR-fab plants were fasciated and exhibited similar severity. (c) Representative inflorescence from a CR-fab plant (CR-fab-3) showing branching (red arrowheads) and weakly fasciated flowers like fab (Fig. 1b). Images of a representative flower and fruit are shown (insets). Scale bar, 1 cm. (d) Quantification and statistical comparisons of floral organ numbers from WT and CR-fab flowers. Data were collected from three plants with confirmed mutations in all sequenced alleles (e; see also Online Methods). Data are means ± s.d., n ≥ 10. A two-tailed, two-sample t test was performed, and significant differences are represented by black asterisks: ***P < 0.0001. (e) CR-fab alleles identified by cloning and sequencing PCR products from the FAB targeted region from three fasciated T0 plants. All alleles carried indel mutations (blue dashed lines and letters), and one of each representative allele identified from sequencing at least eight cloned PCR products is shown. CR-fab-2 and CR-fab-3 are biallelic, and CR-fab-6 is chimeric. Red font highlights sgRNA target sites, and black rectangles indicate PAM sequences; a, allele.
Supplementary Figure 10 CRISPR/Cas9-engineered mutations in SlCLV2 result in weakly fasciated plants.
(a) Schematic illustrating two guide RNAs (sgRNAs; red arrows) targeting the SlCLV2 coding sequence. Cas9/sgRNA1/sgRNA2 were expressed from the same construct. Black arrows indicate the PCR primers used to evaluate mutation type and efficiency. (b) PCR genotyping of six T0 CR-slclv2 plants showing the presence of indel mutations from predominantly chimeric plants, two of which carry the desired deletion. PCR for the presence of Cas9 is shown along with evaluation for fasciation. All five CR-slclv2 fasciated plants exhibited similar severity. (c) Representative primary inflorescence from an slclv2 plant (CR-slclv2-4) showing weak fasciation like fab and CR-fab mutant plants. Images of a representative flower and fruit are shown (insets). Scale bar, 1 cm. (d) Quantification and statistical comparisons of floral organ numbers from WT and CR-slclv2 flowers. Flower fasciation in CR-slclv2 plants is slightly weaker than in fab and CR-fab mutant plants (Fig. 1h and Supplementary Fig. 9). Data were collected from three plants with confirmed mutations in all sequenced alleles (e; see also Online Methods). Data are means ± s.d., n ≥ 10. A two-tailed, two-sample t test was performed, and significant differences are represented by black asterisks: **P < 0.001. (e) CR-slclv2 alleles identified by cloning and sequencing PCR products from the SlCLV2 targeted region from three fasciated T0 plants. The CR-slclv2-3 and CR-slclv2-5 plants were chimeric, and all alleles carried indels (blue dashed lines and letters), including the desired deletion in CR-slclv2-4. One of each representative allele identified from sequencing at least eight cloned PCR products is shown. Red font highlights sgRNA target sites, and black rectangles indicate PAM sequences; a, allele.
Supplementary Figure 13 Combining fas with lc in the wild species S. pimpinellifolium leads to enhanced fasciation.
(a) An unbranched inflorescence typical of the wild species S. pimpinellifolium. (b,c) Primary inflorescence from fas (b) and lc (c) introgressed into S. pimpinellifolium (Sp) showing stronger branching and fasciation for Sp-fas compared to Sp-lc. Sp-lc alone has little effect on branching or floral organ number. (d) Primary inflorescence from an Sp-fas lc plant showing enhanced fasciation, resembling fin primary inflorescences. White arrowheads indicate the fasciated first flower of the primary inflorescence in Sp-fas and Sp-fas lc. Red arrowheads indicate inflorescence branches. Scale bar, 1 cm. Inflorescences from the side shoots of Sp-fas and Sp-fas lc plants show reduced fasciation compared to primary shoots.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 2–7, and Supplementary Note. (PDF 5947 kb)
Supplementary Table 1
Differentially expressed (DE) gene list for fin and fab vegetative meristem transcriptome profiling compared to WT. (XLSX 4116 kb)
Rights and permissions
About this article
Cite this article
Xu, C., Liberatore, K., MacAlister, C. et al. A cascade of arabinosyltransferases controls shoot meristem size in tomato. Nat Genet 47, 784–792 (2015). https://doi.org/10.1038/ng.3309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3309
This article is cited by
-
OsDPE2 Regulates Rice Panicle Morphogenesis by Modulating the Content of Starch
Rice (2023)
-
Genetic architecture of fresh-market tomato yield
BMC Plant Biology (2023)
-
A network of CLAVATA receptors buffers auxin-dependent meristem maintenance
Nature Plants (2023)
-
Vegetable biology and breeding in the genomics era
Science China Life Sciences (2023)
-
CRISPR/Cas9 gene editing technology: a precise and efficient tool for crop quality improvement
Planta (2023)