Abstract
DNA methylation is a key epigenetic modification involved in regulating gene expression and maintaining genomic integrity. Here we inactivated all three catalytically active DNA methyltransferases (DNMTs) in human embryonic stem cells (ESCs) using CRISPR/Cas9 genome editing to further investigate the roles and genomic targets of these enzymes. Disruption of DNMT3A or DNMT3B individually as well as of both enzymes in tandem results in viable, pluripotent cell lines with distinct effects on the DNA methylation landscape, as assessed by whole-genome bisulfite sequencing. Surprisingly, in contrast to findings in mouse, deletion of DNMT1 resulted in rapid cell death in human ESCs. To overcome this immediate lethality, we generated a doxycycline-responsive tTA-DNMT1* rescue line and readily obtained homozygous DNMT1-mutant lines. However, doxycycline-mediated repression of exogenous DNMT1* initiates rapid, global loss of DNA methylation, followed by extensive cell death. Our data provide a comprehensive characterization of DNMT-mutant ESCs, including single-base genome-wide maps of the targets of these enzymes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Jackson, M. et al. Severe global DNA hypomethylation blocks differentiation and induces histone hyperacetylation in embryonic stem cells. Mol. Cell. Biol. 24, 8862–8871 (2004).
Li, E., Bestor, T.H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Okano, M., Bell, D.W., Haber, D.A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Jin, B. et al. DNA methyltransferase 3B (DNMT3B) mutations in ICF syndrome lead to altered epigenetic modifications and aberrant expression of genes regulating development, neurogenesis and immune function. Hum. Mol. Genet. 17, 690–709 (2008).
Klein, C.J. et al. Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat. Genet. 43, 595–600 (2011).
Shah, M.Y. & Licht, J.D. DNMT3A mutations in acute myeloid leukemia. Nat. Genet. 43, 289–290 (2011).
Winkelmann, J. et al. Mutations in DNMT1 cause autosomal dominant cerebellar ataxia, deafness and narcolepsy. Hum. Mol. Genet. 21, 2205–2210 (2012).
Gifford, C.A. et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell 153, 1149–1163 (2013).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Smith, Z.D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012).
Ziller, M.J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013).
Dodge, J.E., Ramsahoye, B.H., Wo, Z.G., Okano, M. & Li, E. De novo methylation of MMLV provirus in embryonic stem cells: CpG versus non-CpG methylation. Gene 289, 41–48 (2002).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Ziller, M.J. et al. Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet. 7, e1002389 (2011).
Fan, G. et al. DNA hypomethylation perturbs the function and survival of CNS neurons in postnatal animals. J. Neurosci. 21, 788–797 (2001).
Jackson-Grusby, L. et al. Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation. Nat. Genet. 27, 31–39 (2001).
Sen, G.L., Reuter, J.A., Webster, D.E., Zhu, L. & Khavari, P.A. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463, 563–567 (2010).
Trowbridge, J.J., Snow, J.W., Kim, J. & Orkin, S.H. DNA methyltransferase 1 is essential for and uniquely regulates hematopoietic stem and progenitor cells. Cell Stem Cell 5, 442–449 (2009).
Tsumura, A. et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11, 805–814 (2006).
Smith, Z.D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Chen, T. et al. Complete inactivation of DNMT1 leads to mitotic catastrophe in human cancer cells. Nat. Genet. 39, 391–396 (2007).
Egger, G. et al. Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc. Natl. Acad. Sci. USA 103, 14080–14085 (2006).
Rhee, I. et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416, 552–556 (2002).
Rhee, I. et al. CpG methylation is maintained in human cancer cells lacking DNMT1. Nature 404, 1003–1007 (2000).
Martins-Taylor, K., Schroeder, D.I., LaSalle, J.M., Lalande, M. & Xu, R.H. Role of DNMT3B in the regulation of early neural and neural crest specifiers. Epigenetics 7, 71–82 (2012).
Horii, T., Tamura, D., Morita, S., Kimura, M. & Hatada, I. Generation of an ICF syndrome model by efficient genome editing of human induced pluripotent stem cells using the CRISPR system. Int. J. Mol. Sci. 14, 19774–19781 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 29, 731–734 (2011).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Miller, J.C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Ding, Q. et al. Enhanced efficiency of human pluripotent stem cell genome editing through replacing TALENs with CRISPRs. Cell Stem Cell 12, 393–394 (2013).
Gordon, C.A., Hartono, S.R. & Chedin, F. Inactive DNMT3B splice variants modulate de novo DNA methylation. PLoS ONE 8, e69486 (2013).
Tsankov, A.M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011).
Chen, T., Ueda, Y., Dodge, J.E., Wang, Z. & Li, E. Establishment and maintenance of genomic methylation patterns in mouse embryonic stem cells by Dnmt3a and Dnmt3b. Mol. Cell. Biol. 23, 5594–5605 (2003).
Smith, Z.D. et al. DNA methylation dynamics of the human preimplantation embryo. Nature 511, 611–615 (2014).
Illingworth, R. et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 6, e22 (2008).
Landau, D.A. et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell 26, 813–825 (2014).
Gerstein, M.B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Stadler, M.B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Barra, V., Schillaci, T., Lentini, L., Costa, G. & Di Leonardo, A. Bypass of cell cycle arrest induced by transient DNMT1 post-transcriptional silencing triggers aneuploidy in human cells. Cell Div. 7, 2 (2012).
Clements, E.G. et al. DNMT1 modulates gene expression without its catalytic activity partially through its interactions with histone-modifying enzymes. Nucleic Acids Res. 40, 4334–4346 (2012).
Shipony, Z. et al. Dynamic and static maintenance of epigenetic memory in pluripotent and somatic cells. Nature 513, 115–119 (2014).
Orford, K.W. & Scadden, D.T. Deconstructing stem cell self-renewal: genetic insights into cell-cycle regulation. Nat. Rev. Genet. 9, 115–128 (2008).
Li, L. et al. Individual cell movement, asymmetric colony expansion, Rho-associated kinase, and E-cadherin impact the clonogenicity of human embryonic stem cells. Biophys. J. 98, 2442–2451 (2010).
Hutnick, L.K., Huang, X., Loo, T.C., Ma, Z. & Fan, G. Repression of retrotransposal elements in mouse embryonic stem cells is primarily mediated by a DNA methylation–independent mechanism. J. Biol. Chem. 285, 21082–21091 (2010).
Gafni, O. et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 504, 282–286 (2013).
Chan, Y.S. et al. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast. Cell Stem Cell 13, 663–675 (2013).
Theunissen, T.W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Di Giaimo, R. et al. The expression of de novo DNA methylase DNMT3b, of the methyl-CpG binding protein MBD2b and of 5-MCDG glycosylase shows two waves of induction during CaCO-2 cell differentiation. Gene 351, 73–81 (2005).
Chen, A.E. et al. Optimal timing of inner cell mass isolation increases the efficiency of human embryonic stem cell derivation and allows generation of sibling cell lines. Cell Stem Cell 4, 103–106 (2009).
Hsiao, E.C. et al. Constitutive Gs activation using a single-construct tetracycline-inducible expression system in embryonic stem cells and mice. Stem Cell Res. Ther. 2, 11 (2011).
Papapetrou, E.P. et al. Stoichiometric and temporal requirements of Oct4, Sox2, Klf4, and c-Myc expression for efficient human iPSC induction and differentiation. Proc. Natl. Acad. Sci. USA 106, 12759–12764 (2009).
Li, H. et al. The histone methyltransferase SETDB1 and the DNA methyltransferase DNMT3A interact directly and localize to promoters silenced in cancer cells. J. Biol. Chem. 281, 19489–19500 (2006).
Rao, L. et al. Highly efficient derivation of skeletal myotubes from human embryonic stem cells. Stem Cell Rev. 8, 1109–1119 (2012).
Si-Tayeb, K. et al. Highly efficient generation of human hepatocyte-like cells from induced pluripotent stem cells. Hepatology 51, 297–305 (2010).
Boyle, P. et al. Gel-free multiplexed reduced representation bisulfite sequencing for large-scale DNA methylation profiling. Genome Biol. 13, R92 (2012).
Bock, C. et al. Quantitative comparison of genome-wide DNA methylation mapping technologies. Nat. Biotechnol. 28, 1106–1114 (2010).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Saxonov, S., Berg, P. & Brutlag, D.L. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc. Natl. Acad. Sci. USA 103, 1412–1417 (2006).
Woodfine, K., Huddleston, J.E. & Murrell, A. Quantitative analysis of DNA methylation at all human imprinted regions reveals preservation of epigenetic stability in adult somatic tissue. Epigenetics Chromatin 4, 1 (2011).
Acknowledgements
We thank all members of the Meissner laboratory, in particular C. Sindhu for helpful discussion and Z.D. Smith for critical feedback on the manuscript. We thank K. Musunuru (Harvard University) for providing the CRIPSR/Cas9 plasmids and thank Q. Ding from the Musunuru laboratory for technical support. J.L. was supported by a postdoctoral fellowship from the Human Frontiers Science Program. J.K.J. is supported by US NIH Director's Pioneer Award DP1 GM105378. A.M. is a New York Stem Cell Foundation Robertson Investigator. The work was funded by a US NIH (National Institute of General Medical Sciences) grant (P01GM099117) and the New York Stem Cell Foundation.
Author information
Authors and Affiliations
Contributions
J.L. and A.M. designed and conceived the study. J.L. generated all the cell lines and performed the experiments. R.K. performed the analysis. H.G. generated the WGBS libraries. A.G. and A.M. supervised the DNA methylation experiments. M.J.Z. and K.C. performed some analysis and assisted in the general data processing. A.M.T. and V.A. performed the experiments with the Scorecard assay. C.A.G. and J.D. assisted in endoderm and hepatocyte differentiation. C.G. performed the dot blot assay. R.P. supported the RNA profiling and FACS analysis. D.R., S.Q.T. and J.K.J. designed and generated the TALENs. W.M. and J.L.R. performed expression analysis. J.L., R.K. and A.M. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.K.J. is a consultant for Horizon Discovery. J.K.J. has financial interests in Editas Medicine and Transposagen Biopharmaceuticals. The interests of J.K.J. were reviewed and are managed by Massachusetts General Hospital and Partners HealthCare in accordance with their conflict of interest policies.
Integrated supplementary information
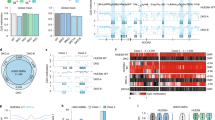
Supplementary Figure 1 Targeted deletion of DNMT1, DNMT3A and DNMT3B in the human ESC line HUES64.
(a) Isoform information relevant to our targeting strategy. The targeted exon is shown in red. Ensembl transcript IDs are shown. (b) Experimental characterization of HUES64 cells. HUES64 cells showed a stable karyotype (46, XY) over the time we have maintained them in culture. Representative bright-field (BR) and OCT4 and NANOG immunostaining images are shown at 10× magnification. Teratoma results from HUES64 cells (P31) demonstrate pluripotency and contribution to all three germ layers: ectoderm (EC), mesoderm (ME) and endoderm (EN). A representative bright-field image of hematoxylin and eosin staining is shown at 4× magnification. Scale bars, 200 μm (4×), 100 μm (10×). (c) Summary of the CRISPR-based targeting efforts. Total refers to the number of total analyzed colonies after selection. Mutations include all validated heterozygous and homozygous mutations. The number of homozygous-mutant clones is shown in the right column for each. (d) Sanger sequencing information for the targeted region of representative clones of DNMT3A–/–, DNMT3B–/– and DNMT3A–/–; DNMT3B–/– cells. The colored lines under the sequence correspond to the lines under sequencing peaks.
Supplementary Figure 2 TaqMan hPSC Scorecard analysis for late-passage wild-type, DNMT3A–/–, DNMT3B–/– and DNMT3A–/–; DNMT3B–/– cells.
The gray boxes represent the score ranges of 11 reference human PSC lines. The wild-type and respective knockout cell lines are shown with different colored circles. Output is from our custom analysis.
Supplementary Figure 3 Global methylation trends in mouse Dnmt knockouts and fidelity of maintenance methylation, NANOG promoter methylation changes and satellite clusters in human DNMT knockouts.
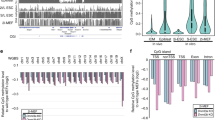
(a) Global analysis of RRBS data from mouse Dnmt3a–/–, Dnmt3b–/–, Dnmt3a–/–; Dnmt3b–/–, and Dnmt1–/– and Dnmt3a–/–; Dnmt3b–/– with Dnmt1 knockdown (TKO). The fractions of CpGs with high (≥ 0.8; red), intermediate (inter, >0.2 and <0.8; green) and low (≤ 0.2; blue) methylation values are shown. Early- and late-passage cells were used for comparison. The “x” in the passage number indicates the passage when we received the cells. (b) Rate of methylation loss in DNMT3A–/–; DNMT3B–/– cells from passage 6 to passage 22 in different genomic features and imputed fidelity of DNMT1 in those regions (RRBS data). Fidelity was calculated by fitting an exponential model (Online Methods). (c) IGV genome browser snapshot of the NANOG promoter showing the complete absence of de novo methylation in DNMT3A–/–; DNMT3B–/– cells. In contrast, the wild-type and single-knockout cells show low levels of methylation that are consistent with the known heterogeneity of NANOG expression. The heat map below shows the DNA methylation values of individual CpGs within the highlighted region. The average DNA methylation value for the entire highlighted region is shown on the right. (d) Distribution of DNA methylation levels across various satellite clusters in the human DNMT knockouts.
Supplementary Figure 4 High-resolution view of CpG island methylation, methylation of imprinted regions and functional enrichment analysis of DMRs.
(a) Venn diagrams showing the overlap between the DMRs detected in each knockout cell line for non-repetitive 1-kb tiles and for CpG islands. (b) Aggregation plots for the Illingworth CpG island (CGI) set showing the loss of methylation in DNMT3A–/–; DNMT3B–/– cells relative to wild-type cells. (c) Methylation levels of known human imprinted control regions in wild-type, DNMT3A–/–, DNMT3B–/– and DNMT3A–/–; DNMT3B–/– cells. (d) Functional annotation enrichment for DNMT3A–/–; DNMT3B–/– DMRs. The test set was non-repetitive 1-kb tiles that were hypomethylated in the DNMT3A–/–; DNMT3B–/– cells, with the background being 1-kb tiles that had methylation of at least 0.4 in wild-type cells. Enrichments were calculated using GREAT, with DMRs being associated with genes within 20 kb.
Supplementary Figure 5 TaqMan hPSC Scorecard analysis for unsorted and sorted endoderm (dEN) cells.
Unsorted and sorted endoderm (dEN) cells are shown as different colored dots. The gray boxes represent the score ranges of 11 reference human PSC lines. Output was taken directly from the Life Technologies analysis website.
Supplementary Figure 6 Alternative strategies/efforts to deplete DNMT1 in human ESCs.
(a) Summary of the TALEN-based targeting efforts. Total refers to the number of colonies after selection. The mutations include all validated heterozygous and homozygous mutations. The number of homozygous clones is shown in the right column for each genotype. (b) RT-qPCR analysis of mRNA expression levels for DNMT1 in DNMT1-knockdown cells. DNMT1-knockdown lines were generated using HUES64 cells. Top, GIPZ DNMT1 shRNA sets from Thermo Scientific. Five individual shRNAs were tested. Bottom, TRIPZ DOX-inducible DNMT1 shRNA sets from Thermo Scientific. Four individual shRNAs were tested with and without DOX induction. (c) Schematic of the DNMT1 rescue constructs. (d) Designed mutations within the exogenous DNMT1* sequence do not alter the protein. The representative amino acid sequence representing the protein remains unchanged. The colored amino acid sequence corresponds to the DNA sequence in Figure 6a. (e) Targeting efficiency of DNMT1 knockout in the tTA+DNMT1* rescue line. The number of mutations includes the number of homozygous clones. (f) Representative sequencing information for DNMT1–/– cells. The colored lines under the sequence correspond to the lines under sequencing peaks. 3′ primer was used for sequencing. Therefore, the reverse-complement sequence is presented in the peaks. (g) RT-qPCR analysis of mRNA expression levels for DNMT3A and DNMT3B in clone 122 validated in f. Primer names and positions are shown in Figure 1d. Error bars were generated from technical triplicates and represent one standard error.
Supplementary Figure 7 Cell death and loss of DNA methylation after DNMT1* withdrawal.
(a) Evidence of acute apoptosis after DNMT1* withdrawal. Representative apoptosis marker Annexin V immunostaining images are shown at 10× magnification. Scale bars, 100 μm. The cell line used here is clone 111. (b) Evidence of DNA damage after DNMT1* withdrawal. Representative DNA damage marker γ-H2A.X (phosphorylated at Ser139) images are shown at 10× magnification. Scale bars, 100 μm. The cell line used here is clone 111. (c) Cell cycle after DNMT1* withdrawal. The cell line used here is clone 111. (d) Schematic of the catalytically inactive DNMT1 rescue constructs. Clone 111 was separately transduced by plenti-ef1α-IRE-nlsEGFP as a non-rescue control, plenti-ef1α-DNMT1C1226W-IRE-nlsEGFP as a catalytically inactive DNMT1 rescue and plenti-ef1α-DNMT1WT-IRE-nlsEGFP as a wild-type DNMT1 rescue control. Stable rescue and control lines were established by sorting for GFP-positive cells. (e) Representative sequencing information for the catalytically inactive DNMT1C1226W construct. The bold letters within the sequence correspond to the lines under sequencing peaks. (f) Western blot analysis of expression of catalytically inactive DNMT1C1226W in 293 cells. There is no apparent degradation of the overexpressed proteins DNMT1WT or DNMT1C1226W. (g) Cell death of the catalytically inactive DNMT1C1226W rescue line after DNMT1* withdrawal. Cells were labeled with nucleus-localized EGFP as the schematic shows in d. Representative images of EGFP-positive cells are shown at 20× magnification. Scale bars, 100 μm. (h) Dot blot assay for DNMT1* withdrawal. The cell line used here is clone 111. (i) Dot blot assay for DNMT1* withdrawal in the catalytically inactive DNMT1C1226W rescue line.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 4 and 5 (PDF 5230 kb)
Supplementary Table 1
List of DMRs in knockout cell lines (1-kb tiles and CpG islands). (XLS 2098 kb)
Supplementary Table 2
GO analysis for enrichment of regions hypomethylated in DNMT3A–/–; DNMT3B–/– cells. (XLS 193 kb)
Supplementary Table 3
Cell cycle analysis after DNMT1* withdrawal. (XLS 27 kb)
Rights and permissions
About this article
Cite this article
Liao, J., Karnik, R., Gu, H. et al. Targeted disruption of DNMT1, DNMT3A and DNMT3B in human embryonic stem cells. Nat Genet 47, 469–478 (2015). https://doi.org/10.1038/ng.3258
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3258
This article is cited by
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
DNMT3B supports meso-endoderm differentiation from mouse embryonic stem cells
Nature Communications (2023)
-
Hybridization-based CpG methylation level detection using methyl-CpG-binding domain–fused luciferase
Analytical and Bioanalytical Chemistry (2023)
-
DNMT3A-coordinated splicing governs the stem state switch towards differentiation in embryonic and haematopoietic stem cells
Nature Cell Biology (2023)
-
Neurogenesis in the neonatal rat hippocampus is regulated by sexually dimorphic epigenetic modifiers
Biology of Sex Differences (2022)