Abstract
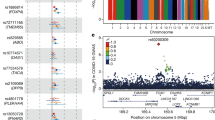
Human genetic factors predispose to tuberculosis (TB). We studied 7.6 million genetic variants in 5,530 people with pulmonary TB and in 5,607 healthy controls. In the combined analysis of these subjects and the follow-up cohort (15,087 TB patients and controls altogether), we found an association between TB and variants located in introns of the ASAP1 gene on chromosome 8q24 (P = 2.6 × 10−11 for rs4733781; P = 1.0 × 10−10 for rs10956514). Dendritic cells (DCs) showed high ASAP1 expression that was reduced after Mycobacterium tuberculosis infection, and rs10956514 was associated with the level of reduction of ASAP1 expression. The ASAP1 protein is involved in actin and membrane remodeling and has been associated with podosomes. The ASAP1-depleted DCs showed impaired matrix degradation and migration. Therefore, genetically determined excessive reduction of ASAP1 expression in M. tuberculosis–infected DCs may lead to their impaired migration, suggesting a potential mechanism of predisposition to TB.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zumla, A., Raviglione, M., Hafner, R. & von Reyn, C.F. Tuberculosis. N. Engl. J. Med. 368, 745–755 (2013).
Möller, M. & Hoal, E.G. Current findings, challenges and novel approaches in human genetic susceptibility to tuberculosis. Tuberculosis (Edinb.) 90, 71–83 (2010).
Apt, A. & Kramnik, I. Man and mouse TB: contradictions and solutions. Tuberculosis (Edinb.) 89, 195–198 (2009).
Thye, T. et al. Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat. Genet. 42, 739–741 (2010).
Thye, T. et al. Common variants at 11p13 are associated with susceptibility to tuberculosis. Nat. Genet. 44, 257–259 (2012).
Chimusa, E.R. et al. Genome-wide association study of ancestry-specific TB risk in the South African Coloured population. Hum. Mol. Genet. 23, 796–809 (2014).
Zhang, F.R. et al. Genome-wide association study of leprosy. N. Engl. J. Med. 361, 2609–2618 (2009).
Zhang, F. et al. Identification of two new loci at IL23R and RAB32 that influence susceptibility to leprosy. Nat. Genet. 43, 1247–1251 (2011).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Nie, Z. & Randazzo, P.A. Arf GAPs and membrane traffic. J. Cell Sci. 119, 1203–1211 (2006).
Randazzo, P.A., Inoue, H. & Bharti, S. Arf GAPs as regulators of the actin cytoskeleton. Biol. Cell 99, 583–600 (2007).
Murphy, D.A. & Courtneidge, S.A. The 'ins' and 'outs' of podosomes and invadopodia: characteristics, formation and function. Nat. Rev. Mol. Cell Biol. 12, 413–426 (2011).
Bharti, S. et al. Src-dependent phosphorylation of ASAP1 regulates podosomes. Mol. Cell. Biol. 27, 8271–8283 (2007).
Lin, D. et al. ASAP1, a gene at 8q24, is associated with prostate cancer metastasis. Cancer Res. 68, 4352–4359 (2008).
Onodera, Y. et al. Expression of AMAP1, an ArfGAP, provides novel targets to inhibit breast cancer invasive activities. EMBO J. 24, 963–973 (2005).
Ehlers, J.P., Worley, L., Onken, M.D. & Harbour, J.W. DDEF1 is located in an amplified region of chromosome 8q and is overexpressed in uveal melanoma. Clin. Cancer Res. 11, 3609–3613 (2005).
Müller, T. et al. ASAP1 promotes tumor cell motility and invasiveness, stimulates metastasis formation in vivo, and correlates with poor survival in colorectal cancer patients. Oncogene 29, 2393–2403 (2010).
Lin, P.L. et al. Sterilization of granulomas is common in active and latent tuberculosis despite within-host variability in bacterial killing. Nat. Med. 20, 75–79 (2014).
Ernst, J.D. The immunological life cycle of tuberculosis. Nat. Rev. Immunol. 12, 581–591 (2012).
Wolf, A.J. et al. Mycobacterium tuberculosis infects dendritic cells with high frequency and impairs their function in vivo. J. Immunol. 179, 2509–2519 (2007).
Wolf, A.J. et al. Initiation of the adaptive immune response to Mycobacterium tuberculosis depends on antigen production in the local lymph node, not the lungs. J. Exp. Med. 205, 105–115 (2008).
Roberts, L.L. & Robinson, C.M. Mycobacterium tuberculosis infection of human dendritic cells decreases integrin expression, adhesion and migration to chemokines. Immunology 141, 39–51 (2014).
Barreiro, L.B. et al. Deciphering the genetic architecture of variation in the immune response to Mycobacterium tuberculosis infection. Proc. Natl. Acad. Sci. USA 109, 1204–1209 (2012).
Szeszko, J.S. et al. Resequencing and association analysis of the SP110 gene in adult pulmonary tuberculosis. Hum. Genet. 121, 155–160 (2007).
Korn, J.M. et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, 1253–1260 (2008).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 1, 457–470 (2012).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Maller, J.B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G.R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Götz, A. & Jessberger, R. Dendritic cell podosome dynamics does not depend on the F-actin regulator SWAP-70. PLoS ONE 8, e60642 (2013).
Acknowledgements
The study was supported by Wellcome Trust grants 088838/Z/09/Z (to S.N., J.C.B., F.D.) and 095198/Z/10/Z (to S.N.), EU Framework Programme 7 Collaborative grant 201483 (to S.N., F.D., R.D.H., P.N.), European Research Council Starting grant 260477 (to S.N.), and Royal Society grants UF0763346 (to S.N.) and RG090638 (to S.N.). S.N. is a Wellcome Trust Senior Research Fellow in Basic Biomedical Science and is also supported by the National Institute for Health Research (NIHR) Cambridge Biomedical Research Centre. This study makes use of data generated by the Wellcome Trust Case-Control Consortium. A full list of the investigators who contributed to the generation of the data is available from www.wtccc.org.uk. Funding for the project was provided by the Wellcome Trust under award 076113, 085475 and 090355.
Author information
Authors and Affiliations
Contributions
S.N. conceived and supervised the study, participated in sample collection and data analysis, and wrote the first draft of the manuscript. J.C. prepared DNA samples and participated in their genotyping and analysis. Y.L. performed statistical analysis of the GWAS data. J.Z.L. participated in the GWAS data analysis. H.L.Z. and D.C.-L. studied DCs and performed matrix degradation and cell migration experiments. C.W. studied ASAP1 mRNA expression in leukocytes. K.L. performed expression quantitative trait locus (eQTL) analysis in DCs. M.M. and A.A. prepared cells for functional experiments. O.I., Y.B., V.N., R.D.H. and F.D. participated in study design, protocol development and sample collection. E.S. and L.K. participated in DNA sample extraction. I.B. and P.N. participated in genotyping. T.T. and C.G.M. participated in genotyping and analysis of the Ghanaian data. V.P. and J.C.B. participated in and supervised statistical analyses. All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Quality control of the GWAS samples by heterozygosity and missing genotypes rate
Dashed red lines represent QC thresholds of more than 2% missing genotype data or excess ± 3.5 standard deviation of heterozygosity rate. Samples circled in red were excluded from the association study.
Supplementary Figure 2 Principal component analysis (PCA) of the Russian GWAS samples
(a) First and second principal components. Russian samples are plotted with ten HapMap3 populations (ASW – African ancestry in Southwest USA; CEU – Utah residents of European ancestry; CHB – Han Chinese in Beijing, China; CHD – Chinese in Metropolitan Denver, Colorado; JPT – Japanese in Tokyo, Japan; GIH – Gujarati Indians in Houston, Texas; LWK – Luhya in Webuye, Kenya; MKK – Maasai in Kinyawa, Kenya; TSI – Tuscans in Italy; YRI – Yoruba in Ibadan, Nigeria).
(b) All GWAS samples plotted on the first two principal components colored by the city of sample origin.
(c) Russian TB cases and controls projected onto the first two principal components using SNP weights precomputed from European samples in the 1000 Genomes Phase III project (CEU — Utah residents with Northern and Western European ancestry; IBS — Iberian populations in Spain; FIN — Finnish in Finland; GBR — British in England and Scotland; and TSI — Toscani in Italy) using SNPweights (Chen, C.Y. et al. Improved ancestry inference using weights from external reference panels. Bioinformatics 29, 1399-1406 (2013)).
Supplementary Figure 3 The quantile - quantile plot of the observed versus the expected –log10P-values under the null for 7,614,862 SNPs from the GWAS results after genotype imputation
The diagonal black line is y = x, and the grey shapes show 95% confidence interval under the null. The overall distribution has a genomic inflation factor (λGC) of 1.10.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 391 kb)
Supplementary Tables
Supplementary Tables 1–3 and 5–7 (PDF 215 kb)
Supplementary Table 4
TB association in the Russian GWAS for loci that previously were associated with Inflammatory Bowel Disease (XLS 48 kb)
Rights and permissions
About this article
Cite this article
Curtis, J., Luo, Y., Zenner, H. et al. Susceptibility to tuberculosis is associated with variants in the ASAP1 gene encoding a regulator of dendritic cell migration. Nat Genet 47, 523–527 (2015). https://doi.org/10.1038/ng.3248
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3248
This article is cited by
-
The immunogenetics of tuberculosis (TB) susceptibility
Immunogenetics (2023)
-
Decoding the spatial chromatin organization and dynamic epigenetic landscapes of macrophage cells during differentiation and immune activation
Nature Communications (2022)
-
Influence of genetics and the pre-vaccination blood transcriptome on the variability of antibody levels after vaccination against Mycoplasma hyopneumoniae in pigs
Genetics Selection Evolution (2021)
-
An IL7RA exon 5 polymorphism is associated with impaired IL-7Rα splicing and protection against tuberculosis in Ghana
Genes & Immunity (2019)
-
Fine-mapping analysis of a chromosome 2 region linked to resistance to Mycobacterium tuberculosis infection in Uganda reveals potential regulatory variants
Genes & Immunity (2019)