Abstract
The capacity of microbial pathogens to alter their host tropism leading to epidemics in distinct host species populations is a global public and veterinary health concern. To investigate the molecular basis of a bacterial host-switching event in a tractable host species, we traced the evolutionary trajectory of the common rabbit clone of Staphylococcus aureus. We report that it evolved through a likely human-to-rabbit host jump over 40 years ago and that only a single naturally occurring nucleotide mutation was required and sufficient to convert a human-specific S. aureus strain into one that could infect rabbits. Related mutations were identified at the same locus in other rabbit strains of distinct clonal origin, consistent with convergent evolution. This first report of a single mutation that was sufficient to alter the host tropism of a microorganism during its evolution highlights the capacity of some pathogens to readily expand into new host species populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Marano, N., Arguin, P.M. & Pappaioanou, M. Impact of globalization and animal trade on infectious disease ecology. Emerg. Infect. Dis. 13, 1807–1809 (2007).
Imai, M. et al. Experimental adaptation of an influenza H5 HA confers respiratory droplet transmission to a reassortant H5 HA/H1N1 virus in ferrets. Nature 486, 420–428 (2012).
Russell, C.A. et al. The potential for respiratory droplet–transmissible A/H5N1 influenza virus to evolve in a mammalian host. Science 336, 1541–1547 (2012).
Herfst, S. et al. Airborne transmission of influenza A/H5N1 virus between ferrets. Science 336, 1534–1541 (2012).
Fitzgerald, J.R. Livestock-associated Staphylococcus aureus: origin, evolution and public health threat. Trends Microbiol. 20, 192–198 (2012).
Weinert, L.A. et al. Molecular dating of human-to-bovid host jumps by Staphylococcus aureus reveals an association with the spread of domestication. Biol. Lett. 8, 829–832 (2012).
Guinane, C.M. et al. Evolutionary genomics of Staphylococcus aureus reveals insights into the origin and molecular basis of ruminant host adaptation. Genome Biol. Evol. 2, 454–466 (2010).
Lowder, B.V. et al. Recent human-to-poultry host jump, adaptation, and pandemic spread of Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 106, 19545–19550 (2009).
Price, L.B. et al. Staphylococcus aureus CC398: host adaptation and emergence of methicillin resistance in livestock. MBio. 3, e00305–11 (2012).
Viana, D. et al. Adaptation of Staphylococcus aureus to ruminant and equine hosts involves SaPI-carried variants of von Willebrand factor–binding protein. Mol. Microbiol. 77, 1583–1594 (2010).
van Wamel, W.J.B., Rooijakkers, S.H.M., Ruyken, M., Van Kessel, K.P.M. & Van Strijp, J.A.G. The innate immune modulators staphylococcal complement inhibitor and chemotaxis inhibitory protein of Staphylococcus aureus are located on β-hemolysin–converting bacteriophages. J. Bacteriol. 188, 1310–1315 (2006).
Ubeda, C. et al. Sip, an integrase protein with excision, circularization and integration activities, defines a new family of mobile Staphylococcus aureus pathogenicity islands. Mol. Microbiol. 49, 193–210 (2003).
Rasigade, J.P. et al. Global distribution and evolution of Panton-Valentine leukocidin-positive methicillin-susceptible Staphylococcus aureus, 1981–2007. J. Infect. Dis. 201, 1589–1597 (2010).
Vancraeynest, D. et al. International dissemination of a high virulence rabbit Staphylococcus aureus clone. J. Vet. Med. B Infect. Dis. Vet. Public Health 53, 418–422 (2006).
Kurt, K. et al. Subpopulations of Staphylococcus aureus clonal complex 121 are associated with distinct clinical entities. PLoS One 8, e58155 (2013).
Drummond, A.J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Spaan, A.N. et al. The staphylococcal toxin Panton-Valentine Leukocidin targets human C5a receptors. Cell Host Microbe 13, 584–594 (2013).
Burts, M.L., Williams, W.A., DeBord, K. & Missiakas, D.M. EsxA and EsxB are secreted by an ESAT-6–like system that is required for the pathogenesis of Staphylococcus aureus infections. Proc. Natl. Acad. Sci. USA 102, 1169–1174 (2005).
Hu, C., Xiong, N., Zhang, Y., Rayner, S. & Chen, S. Functional characterization of lipase in the pathogenesis of Staphylococcus aureus. Biochem. Biophys. Res. Commun. 419, 617–620 (2012).
Saïd-Salim, B. et al. Global regulation of Staphylococcus aureus genes by Rot. J. Bacteriol. 185, 610–619 (2003).
Peschel, A. et al. Inactivation of the dlt operon in Staphylococcus aureus confers sensitivity to defensins, protegrins, and other antimicrobial peptides. J. Biol. Chem. 274, 8405–8410 (1999).
Collins, L.V. et al. Staphylococcus aureus strains lacking D-alanine modifications of teichoic acids are highly susceptible to human neutrophil killing and are virulence attenuated in mice. J. Infect. Dis. 186, 214–219 (2002).
Hofmann, K. A superfamily of membrane-bound O-acyltransferases with implications for wnt signaling. Trends Biochem. Sci. 25, 111–112 (2000).
Subbarao, E.K., London, W. & Murphy, B.R. A single amino acid in the PB2 gene of influenza A virus is a determinant of host range. J. Virol. 67, 1761–1764 (1993).
Wain, L.V. et al. Adaptation of HIV-1 to its human host. Mol. Biol. Evol. 24, 1853–1860 (2007).
Naffakh, N., Tomoiu, A., Rameix-Welti, M.-A. & van der Werf, S. Host restriction of avian influenza viruses at the level of the ribonucleoproteins. Annu. Rev. Microbiol. 62, 403–424 (2008).
Etienne, L., Hahn, B.H., Sharp, P.M., Matsen, F.A. & Emerman, M. Gene loss and adaptation to hominids underlie the ancient origin of HIV-1. Cell Host Microbe 14, 85–92 (2013).
Mandel, M.J., Wollenberg, M.S., Stabb, E.V., Visick, K.L. & Ruby, E.G. A single regulatory gene is sufficient to alter bacterial host range. Nature 458, 215–218 (2009).
Lecuit, M. et al. A single amino acid in E-cadherin responsible for host specificity towards the human pathogen Listeria monocytogenes. EMBO J. 18, 3956–3963 (1999).
Wollert, T. et al. Extending the host range of Listeria monocytogenes by rational protein design. Cell 129, 891–902 (2007).
Tormo-Más, M.A. et al. Moonlighting bacteriophage proteins derepress staphylococcal pathogenicity islands. Nature 465, 779–782 (2010).
Tormo-Más, M.Á. et al. Phage dUTPases control transfer of virulence genes by a proto-oncogenic G protein–like mechanism. Mol. Cell 49, 947–958 (2013).
Arnaud, M., Chastanet, A. & Débarbouillé, M. New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive bacteria. Appl. Environ. Microbiol. 70, 6887–6891 (2004).
Darling, A.E., Mau, B. & Perna, N.T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5, e11147 (2010).
Marttinen, P. et al. Detection of recombination events in bacterial genomes from large population samples. Nucleic Acids Res. 40, e6 (2012).
Iqbal, Z., Turner, I. & McVean, G. High-throughput microbial population genomics using the Cortex variation assembler. Bioinformatics 29, 275–276 (2013).
Drummond, A.J., Ho, S.Y.W., Phillips, M.J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Zhao, Y. et al. PGAP: pan-genomes analysis pipeline. Bioinformatics 28, 416–418 (2012).
Gründling, A. & Schneewind, O. Synthesis of glycerol phosphate lipoteichoic acid in Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 104, 8478–8483 (2007).
Bertsche, U. et al. Increased cell wall teichoic acid production and D-alanylation are common phenotypes among daptomycin-resistant methicillin-resistant Staphylococcus aureus (MRSA) clinical isolates. PLoS ONE 8, e67398 (2013).
Acknowledgements
We thank J. Etienne for helpful advice, O. Schneewind, M. Woolhouse and Í. Lasa for comments on the manuscript, C. Cervera and E. Blas for their support with the in vivo experiments, and R. Cartwright for excellent technical assistance. We are grateful to Edinburgh Genomics (Roslin Institute) for sequencing services. This work was supported by grants BIO2011-30503-C02-01, Eranet-pathogenomics PIM2010EPA-00606 and Consolider-Ingenio CSD2009-00006 from Ministerio de Ciencia e Innovación (Spain) and strategic grant funding from the University of Glasgow to J.R.P.; by a project grant (BB/I013873/1) and institute strategic grant funding from the Biotechnology and Biological Sciences Research Council (UK) to J.R.F., in addition to a doctoral training grant from the Medical Research Council (UK) to J.R.F.; and by grants AGL2011-30170-CO2-02 (Ministerio de Ciencia e Innovación, Spain) and GV2013-077 (Conselleria d'Educació, Cultura i Esport, Generalitat Valenciana) to D.V.
Author information
Authors and Affiliations
Contributions
J.R.F. and J.R.P. conceived and designed the study. D.V., M.C. and L.S. generated and characterized the different mutant strains. P.R.M., M.J.W. and C.M.G. performed the genomic studies. B.M.G.-M. and S.J.F. measured D-Ala content. A.T. provided human strains. J.R.P., J.R.F. and S.J.F. supervised the research. J.R.F. and J.R.P. wrote the manuscript and obtained funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Alignment of DltB amino acid sequences from S. aureus human, rabbit, ovine and poultry clones, colored according to relative sequence conservation at each position.
Adapted from an alignment generated by PRALINE. The scoring scheme ranges from 0 for the least conserved alignment position up to 10 (indicated by an asterisk) for the most conserved alignment position.
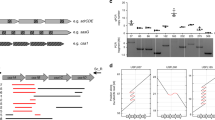
Supplementary Figure 2 Analysis of D-alanylation of wall teichoic acid or lipoteichoic acid and growth inhibition by the cationic peptide nisin.
(a,b) The D-Ala content of the human and rabbit S. aureus clones was tested. (c) Shown is the growth in TSB medium of isogenic strains treated with nisin (10 μg/ml). Cell density was monitored (OD600). J, wild-type rabbit strain; J dlt Bh, derivative J strain expressing the DltB protein from the ST121 human clones; J Δdlt B, J dlt B mutant; F, wild-type human strain; F dlt Br, strain F expressing the DltB protein from the rabbit clones. In both cases, the experiments were performed in triplicate. An ANOVA test was carried out using Bonferroni adjustment. All error bars show s.e.m. Differences that are statistically significant are indicated by an asterisk (P < 0.05); all other comparisons were not significant.
Supplementary Figure 3 Analysis of peptidoglycan structure and composition.
(a) Muropeptide analysis by HPLC. The cell wall from bacteria growing in exponential phase was isolated, digested with a muramidase and analyzed via HPLC. (b) The amino acid composition of the purified peptidoglycan was analyzed by HPLC. In both panels, a representative experiment is shown.
Supplementary Figure 4 Predicted membrane topology of the DltB protein.
DltB topology was predicted using the TMHMM method. Colored in red are amino acid residues that varied in the rabbit ST121 strains, and green indicates amino acid residues that were variant in the other rabbit clones.
Supplementary Figure 5 Survival in rabbit blood.
S. aureus strains were grown to mid-exponential growth phase, washed and resuspended in sterile PBS. 4 × 104 CFU of S. aureus in 100 μl of PBS were pipetted slowly into 3 ml of heparinized rabbit blood, mixed gently for 30 s and incubated at 37 °C for 2 h. To determine survival rates, 100 μl of heparinized blood was plated on TSA for S. aureus detection. Data shown represent the means ± s.e.m. of three separate experiments. An ANOVA test was performed, using Bonferroni adjustment. Differences that are statistically significant are indicated by an asterisk (P < 0.05); all other comparisons were not significant.
Supplementary Figure 6 SDS-PAGE analysis of cell wall–associated protein profiles.
S. aureus strains were grown to mid-exponential or stationary growth phase, washed and resuspended in lysis buffer (50 mM Tris-HCl (pH 7.5), 20 mM MgCl2, supplemented with 30% raffinose) with lysostaphin. Protoplasts were sedimented by centrifugation at 6,000g, and the supernatant fraction, which contained the wall-associated proteins, was analyzed by SDS-PAGE.
Supplementary Figure 7 Alignment of the DltB amino acid sequences from Bacillus amyloliquefaciens and Streptococcus pneumoniae strains, colored according to relative sequence conservation at each position.
Adapted from an alignment generated by PRALINE. The scoring scheme ranges from 0 for the least conserved alignment position up to 10 (indicated by an asterisk) for the most conserved alignment position. (a) B_amyl_FZB42: B. amyloliquefaciens strain FZB42 (plant-associated bacterium). Accession number: YP_001423132. B_amyl_DSM7: B. amyloliquefaciens strain DSM7 (soil adapted). Accession number: YP_003922279. (b) DltB_TIGR4: S. pneumoniae strain TIGR4. Accession number: ZP_01408978. DltB_GA41301: S. pneumoniae strain GA41301. Accession number: ZP_12336902. DltB_GA17227: S. pneumoniae strain GA17227. Accession number: ZP_12797610. DltB_GA47901: S. pneumoniae strain GA47901. Accession number: ZP_12344230.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–6. (PDF 2182 kb)
Rights and permissions
About this article
Cite this article
Viana, D., Comos, M., McAdam, P. et al. A single natural nucleotide mutation alters bacterial pathogen host tropism. Nat Genet 47, 361–366 (2015). https://doi.org/10.1038/ng.3219
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3219
This article is cited by
-
Multi-host infection and phylogenetically diverse lineages shape the recombination and gene pool dynamics of Staphylococcus aureus
BMC Microbiology (2023)
-
Staphylococcus aureus host interactions and adaptation
Nature Reviews Microbiology (2023)
-
Host genotype and genetic diversity shape the evolution of a novel bacterial infection
The ISME Journal (2021)
-
Differences in virulence between the two more prevalent Staphylococcus aureus clonal complexes in rabbitries (CC121 and CC96) using an experimental model of mammary gland infection
Veterinary Research (2020)
-
“Gene accordions” cause genotypic and phenotypic heterogeneity in clonal populations of Staphylococcus aureus
Nature Communications (2020)


