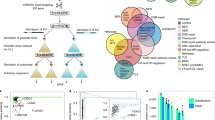
Abstract
DNA replication−associated mutations are repaired by two components: polymerase proofreading and mismatch repair. The mutation consequences of disruption to both repair components in humans are not well studied. We sequenced cancer genomes from children with inherited biallelic mismatch repair deficiency (bMMRD). High-grade bMMRD brain tumors exhibited massive numbers of substitution mutations (>250/Mb), which was greater than all childhood and most cancers (>7,000 analyzed). All ultra-hypermutated bMMRD cancers acquired early somatic driver mutations in DNA polymerase ɛ or δ. The ensuing mutation signatures and numbers are unique and diagnostic of childhood germ-line bMMRD (P < 10−13). Sequential tumor biopsy analysis revealed that bMMRD/polymerase-mutant cancers rapidly amass an excess of simultaneous mutations (∼600 mutations/cell division), reaching but not exceeding ∼20,000 exonic mutations in <6 months. This implies a threshold compatible with cancer-cell survival. We suggest a new mechanism of cancer progression in which mutations develop in a rapid burst after ablation of replication repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wimmer, K. et al. Diagnostic criteria for constitutional mismatch repair deficiency syndrome: suggestions of the European consortium 'care for CMMRD' (C4CMMRD). J. Med. Genet. 51, 355–365 (2014).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Liu, B. et al. Mismatch repair gene defects in sporadic colorectal cancers with microsatellite instability. Nat. Genet. 9, 48–55 (1995).
Wu, G. et al. The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat. Genet. 46, 444–450 (2014).
Narayanan, L., Fritzell, J.A., Baker, S.M., Liskay, R.M. & Glazer, P.M. Elevated levels of mutation in multiple tissues of mice deficient in the DNA mismatch repair gene Pms2. Proc. Natl. Acad. Sci. USA 94, 3122–3127 (1997).
Thomas, D.C., Roberts, J.D. & Kunkel, T.A. Heteroduplex repair in extracts of human HeLa cells. J. Biol. Chem. 266, 3744–3751 (1991).
Panigrahi, G.B., Slean, M.M., Simard, J.P., Gileadi, O. & Pearson, C.E. Isolated short CTG/CAG DNA slip-outs are repaired efficiently by hMutSbeta, but clustered slip-outs are poorly repaired. Proc. Natl. Acad. Sci. USA 107, 12593–12598 (2010).
Henninger, E.E. & Pursell, Z.F. DNA polymerase epsilon and its roles in genome stability. IUBMB Life 66, 339–351 (2014).
Preston, B.D., Albertson, T.M. & Herr, A.J. DNA replication fidelity and cancer. Semin. Cancer Biol. 20, 281–293 (2010).
Korona, D.A., Lecompte, K.G. & Pursell, Z.F. The high fidelity and unique error signature of human DNA polymerase epsilon. Nucleic Acids Res. 39, 1763–1773 (2011).
Bebenek, K. & Kunkel, T.A. Analyzing fidelity of DNA polymerases. Methods Enzymol. 262, 217–232 (1995).
Ghodgaonkar, M.M. et al. Phenotypic characterization of missense polymerase-delta mutations using an inducible protein-replacement system. Nat. Commun. 5, 4990 (2014).
Venkatesan, R.N. et al. Mutation at the polymerase active site of mouse DNA polymerase delta increases genomic instability and accelerates tumorigenesis. Mol. Cell. Biol. 27, 7669–7682 (2007).
Schmitt, M.W. et al. Active site mutations in mammalian DNA polymerase delta alter accuracy and replication fork progression. J. Biol. Chem. 285, 32264–32272 (2010).
Nick McElhinny, S.A., Gordenin, D.A., Stith, C.M., Burgers, P.M. & Kunkel, T.A. Division of labor at the eukaryotic replication fork. Mol. Cell 30, 137–144 (2008).
Lujan, S.A. et al. Heterogeneous polymerase fidelity and mismatch repair bias genome variation and composition. Genome Res. 24, 1751–1764 (2014).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Kandoth, C. et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Durno, C.A. et al. Oncologic surveillance for subjects with biallelic mismatch repair gene mutations: 10 year follow-up of a kindred. Pediatr. Blood Cancer 59, 652–656 (2012).
Nikolaev, S.I. et al. A single-nucleotide substitution mutator phenotype revealed by exome sequencing of human colon adenomas. Cancer Res. 72, 6279–6289 (2012).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl. Acad. Sci. USA 105, 4283–4288 (2008).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Bakry, D. et al. Genetic and clinical determinants of constitutional mismatch repair deficiency syndrome: report from the constitutional mismatch repair deficiency consortium. Eur. J. Cancer 50, 987–996 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Homer, N. & Nelson, S.F. Improved variant discovery through local re-alignment of short-read next-generation sequencing data using SRMA. Genome Biol. 11, R99 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Tomé, S. et al. Tissue-specific mismatch repair protein expression: MSH3 is higher than MSH6 in multiple mouse tissues. DNA Repair (Amst.) 12, 46–52 (2013).
Shinbrot, E. et al. Exonuclease mutations in DNA polymerase epsilon reveal replication strand specific mutation patterns and human origins of replication. Genome Res. 24, 1740–1750 (2014).
Acknowledgements
U.T. received funding from BRAINchild Canada and the Canadian Institute of Health Research (operating grant MOP123268). C.E.P. received funding from the Canadian Institute of Health Research (operating grant FRN131596). P.J.C. and A.G. are personally funded through Wellcome Trust Senior Clinical and Basic Research Fellowships and are members of the Wellcome-funded COMSIG consortium. S.B. is funded through a Wellcome Trust Research Training Fellowship for Clinicians. B.B.C., M.M. and R.A. are supported by a SickKids Restracomp award. We acknowledge J. Costello for his contribution to the manuscript.
Author information
Authors and Affiliations
Consortia
Contributions
A.S., E.B., C.H., P.J.C. and U.T. designed the study. B.B.C., A.P., T.L., A.H., S.D., N.A., B.M., M.G., Y.H., D.M.M., M.R., Ma.R., G.B.P., N.P.T., K.P.H., E.E.H., A.Y.G., D.B., G.S.C., H.D., J.L.E. and M.M. performed experiments. A.S., B.B.C., R.B., L.B.A., Da.M., D.W., P.V.L., P.S.T., P.C., S.B., R.A., C.D., M.A. and U.T collected and analyzed data. R.G., R.D., Ro.G., R.E., R.F., G.P.T., P.C.N., S.A., S.B.-S., S.C.L., S.C., P.D., A.H. and U.T. provided reagents, tissue and clinical data. A.S., M.S.M., M.D.T., Z.F.P., C.E.P., D.M., P.J.C. and U.T. wrote the manuscript. S.G., S.W.S., C.D., M.A., A.G., M.S.M., M.D.T., Z.F.P., C.E.P., D.M., P.A.F., M.R.S., E.B., C.H. and P.J.C. provided technical support and conceptual advice. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of contributing members appears in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 Somatic rearrangements in bMMRD cancers.
Circos plots depicting whole-genome profile of two tumors. Each arc represents a single interchromosomal translocation (purple), large deletion (green), tandem duplication (black) or inversion (orange and blue). Rearrangements were determined by the Breakpoint via assembly algorithm (Brass1).
Supplementary Figure 2 Number of single-nucleotide variations (SNVs) in non-neoplastic bMMRD DNA.
SNVs were determined by standard target capture exome sequencing of blood-derived DNA from 16 bMMRD patients and 35 controls (including individuals unrelated (n = 11) to the patients as well as from parental DNA (n = 24)). Common SNVs found in public databases (dbSNP, 1000 Genomes Project or the NHLBI ESP Project from >6,500 samples) were removed. Shown are the numbers of rare SNVs in patients with bMMRD relative to control samples. Error bars represent s.e.m.
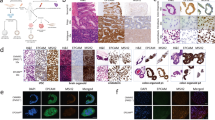
Supplementary Figure 3 MMR protein expression in non-neoplastic biallelic MMR mutant cells by protein blotting.
(a) For mutSα proteins MSH2 and MSH6. (b) For MutLa proteins PMS2 and MLH1. The lymphoblasts used were LoVo as a negative control, MMR10 and MMR8 lymphoblasts from patients with bMMRD, HeLa cells and normal lymphoblasts as a positive control. Each lane was separated by 8% Tris-glycine SDS-PAGE. Protein blotting was simultaneously carried out for mismatch repair proteins and actin.
Supplementary Figure 4 In vitro GT mismatch repair assay
(a) A substrate containing a single GT mismatch was designed keeping a nick several hundred nucleotides away from the mismatch, as shown by the arrow. HindII restriction digestion, which cannot cleave the GT mismatch at its site but can efficiently cleave its repaired AT site, was used for measurement of mismatch repair ability. (b) Reactions were performed as described previously (Panigrahi et al., 2005). GT repaired products were digested by XmnI and HindIII. If repaired correctly, it would be sensitive to HindIII enzyme. The repaired products are marked by the brackets. The digested products were run on a 1% agarose gel and analyzed by Southern hybridization and quantification by Typhoon FLA 9500 Phosphorimager. The bar graphs represent three separate experiments. The last lane contains 50% of LoVo and 50% of MMR10 cell extracts. The protein concentration of the cell extracts was corrected for the purpose of quantification, and the graph was plotted accordingly.
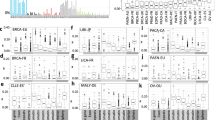
Supplementary Figure 5 Frequency of somatic mutations in DNA replication and repair genes.
Shown is the proportion of ultra-hypermutated cancers that contain somatic nonsynonymous mutations in genes known to be involved in DNA replication or DNA repair. POLE, the most frequently mutated gene, is highlighted.
Supplementary Figure 6 Putative polymerase driver mutations found in ultra-hypermutated bMMRD tumors.
A multispecies alignment for POLE is shown, with each putative driver polymerase mutation indicated (red box). The alignment was performed using BLAST and NCBI's CDD database. Amino acids deemed critical for catalysis are highlighted2,3,4 (yellow). (a) Mutated polymerase ɛ residues are shown, and the exonuclease domain is highlighted (horizontal blue line). The Exo motifs are indicated in orange. Asterisks denote recurrent mutations also found in colorectal or endometrial cancers: mutations at residues F104, P436 and S459 were previously found in colorectal cancers5,6, and S297 was previously found in endometrial cancer7. Polymerase active site mutations have also been found in the germ line of individuals predisposed to colorectal adenomas and carcinomas8. Note that only positions 83–127 and 268–472 of POLE are shown. (b) Examples of somatic POLE passenger mutations, which are not conserved.
Supplementary Figure 7 Putative polymerase driver mutations found in ultra-hypermutated bMMRD tumors.
A multispecies alignment for POLD1 is shown, with each putative driver polymerase mutation indicated (red box). The alignment was performed using BLAST and NCBI's CDD database. Amino acids deemed critical for catalysis are highlighted2–4 (yellow). Mutated polymerase δ residues are shown, and the Pol II and Exo I motifs are highlighted (horizontal green and orange lines, respectively). Note that only positions 581–624 and 303–348 of POLD1 are shown.
Supplementary Figure 8 Timing of the polymerase ɛ mutational signature in bMMRD/POLE cancers.
The top panel shows the variant allele fraction of all somatic point mutations in four bMMRD/POLE cancer genomes (similar to Fig. 2d). The percentages of POLE signature mutations are plotted in the lower panel. That is, each point represents the proportion of NpCpT > NpApT (red) or NpTpT > NpApT (teal) at a given allele fraction. Note that the POLE signature is established early in each cancer and that it continues and is relatively unchanged across the life history of the cancer.
Supplementary Figure 9 Correlation of mutation signatures.
Shown is the similarity of mutation type for all ultra-hypermutated brain cancers (n = 10). Mutations were classified according to their 5′ and 3′ flanking bases (forming a trinucleotide). The proportions of each trinucleotide within all possible mutation classes were compared (that is, the proportion of each of 16 trinucleotides within the 6 classes of substitution: C>A, C>G, C>T, T>A, T>C and T>G). Colors represent the Pearson's correlation between samples.
Supplementary Figure 10 Mutation burden in sporadic colorectal and endometrial cancers.
(a) The total number of somatic mutations in sporadic colorectal and endometrial cancers with either MMR mutations, POLE mutations or both. (b) The relationship between mutation burden and copy number. Colon cancers are plotted on the top row, and endometrial cancers are plotted on the bottom row. In all panels, each dot represents one tumor specimen. The red dots represent the genotype of interest. That is, on the left, red dots are samples with mutation in an MMR gene but not in POLE; in the middle, red dots represent POLE-mutated samples, without MMR mutations; and, on the right, red dots represent samples with both MMR and POLE mutations, which contain the highest number of point mutations and fewest copy number alterations. Data are from http://www.cbioportal.org/.
Supplementary Figure 11 Suggested model for the mutation signature of bMMRD malignant cancers.
A multilayer defense for protecting genome stability prevents replication errors in most individuals. An inherited mismatch repair defect leads to the gradual accumulation of mutations and, thus, to increased cancer risk during adulthood. However, the addition of impaired POLE or POLD1 DNA polymerases results in an extremely rapid accumulation of mutations and onset of cancer in young children.
Supplementary Figure 12 Sequencing coverage.
(a) Coverage of exome sequencing data. Shown is a box plot with each tumor's coverage, for all bases in the exome target. The red dotted line shows the minimum coverage of 25× required for downstream analysis. (b) Coverage of genome sequence data. Shown is the average coverage for all tumor genomes sequenced. The red dotted line shows the minimum desired coverage of 30×.
Supplementary Figure 13 Validation of POLE and POLD1 mutations.
Putative driver mutations in (a) POLE and (b) POLD1 were validated by Sanger sequencing using forward and reverse sequencing.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1 and 3, and Supplementary Note. (PDF 3933 kb)
Supplementary Table 2
Nonsynonymous somatic mutations in ultra-hypermutated cancers (VAF >5%). (XLSX 12042 kb)
Rights and permissions
About this article
Cite this article
Shlien, A., Campbell, B., de Borja, R. et al. Combined hereditary and somatic mutations of replication error repair genes result in rapid onset of ultra-hypermutated cancers. Nat Genet 47, 257–262 (2015). https://doi.org/10.1038/ng.3202
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3202