Abstract
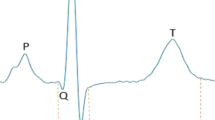
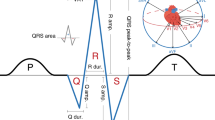
The QT interval, an electrocardiographic measure reflecting myocardial repolarization, is a heritable trait. QT prolongation is a risk factor for ventricular arrhythmias and sudden cardiac death (SCD) and could indicate the presence of the potentially lethal mendelian long-QT syndrome (LQTS). Using a genome-wide association and replication study in up to 100,000 individuals, we identified 35 common variant loci associated with QT interval that collectively explain ∼8–10% of QT-interval variation and highlight the importance of calcium regulation in myocardial repolarization. Rare variant analysis of 6 new QT interval–associated loci in 298 unrelated probands with LQTS identified coding variants not found in controls but of uncertain causality and therefore requiring validation. Several newly identified loci encode proteins that physically interact with other recognized repolarization proteins. Our integration of common variant association, expression and orthogonal protein-protein interaction screens provides new insights into cardiac electrophysiology and identifies new candidate genes for ventricular arrhythmias, LQTS and SCD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
20 July 2014
In the version of this article initially published online, the name of Fabiola del Greco M was misspelled in the author list. The error has been corrected for the print, PDF and HTML versions of this article.
References
Schwartz, P.J., Crotti, L. & Insolia, R. Long-QT syndrome: from genetics to management. Circ. Arrhythm. Electrophysiol. 5, 868–877 (2012).
Newton-Cheh, C. et al. QT interval is a heritable quantitative trait with evidence of linkage to chromosome 3 in a genome-wide linkage analysis: The Framingham Heart Study. Heart Rhythm 2, 277–284 (2005).
Newton-Cheh, C. et al. Common variants at ten loci influence QT interval duration in the QTGEN Study. Nat. Genet. 41, 399–406 (2009).
Pfeufer, A. et al. Common variants at ten loci modulate the QT interval duration in the QTSCD Study. Nat. Genet. 41, 407–414 (2009).
Arking, D.E. et al. A common genetic variant in the NOS1 regulator NOS1AP modulates cardiac repolarization. Nat. Genet. 38, 644–651 (2006).
Nolte, I.M. et al. Common genetic variation near the phospholamban gene is associated with cardiac repolarisation: meta-analysis of three genome-wide association studies. PLoS ONE 4, e6138 (2009).
Holm, H. et al. Several common variants modulate heart rate, PR interval and QRS duration. Nat. Genet. 42, 117–122 (2010).
Noseworthy, P.A. et al. Common genetic variants, QT interval, and sudden cardiac death in a Finnish population-based study. Circ. Cardiovasc. Genet. 4, 305–311 (2011).
Kim, J.W. et al. A common variant in SLC8A1 is associated with the duration of the electrocardiographic QT interval. Am. J. Hum. Genet. 91, 180–184 (2012).
Yang, J. et al. Genome partitioning of genetic variation for complex traits using common SNPs. Nat. Genet. 43, 519–525 (2011).
Voight, B.F. et al. The metabochip, a custom genotyping array for genetic studies of metabolic, cardiovascular, and anthropometric traits. PLoS Genet. 8, e1002793 (2012).
Smith, J.G. et al. Impact of ancestry and common genetic variants on QT interval in African Americans. Circ. Cardiovasc. Genet. 5, 647–655 (2012).
den Hoed, M. et al. Identification of heart rate–associated loci and their effects on cardiac conduction and rhythm disorders. Nat. Genet. 45, 621–631 (2013).
Sotoodehnia, N. et al. Common variants in 22 loci are associated with QRS duration and cardiac ventricular conduction. Nat. Genet. 42, 1068–1076 (2010).
Pfeufer, A. et al. Genome-wide association study of PR interval. Nat. Genet. 42, 153–159 (2010).
Elks, C.E. et al. Thirty new loci for age at menarche identified by a meta-analysis of genome-wide association studies. Nat. Genet. 42, 1077–1085 (2010).
Zabaneh, D. & Balding, D.J. A genome-wide association study of the metabolic syndrome in Indian Asian men. PLoS ONE 5, e11961 (2010).
Lemaitre, R.N. et al. Genetic loci associated with plasma phospholipid n-3 fatty acids: a meta-analysis of genome-wide association studies from the CHARGE Consortium. PLoS Genet. 7, e1002193 (2011).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Chambers, J.C. et al. Genome-wide association study identifies loci influencing concentrations of liver enzymes in plasma. Nat. Genet. 43, 1131–1138 (2011).
Hu, X. et al. Integrating autoimmune risk loci with gene-expression data identifies specific pathogenic immune cell subsets. Am. J. Hum. Genet. 89, 496–506 (2011).
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Corradin, O. et al. Combinatorial effects of multiple enhancer variants in linkage disequilibrium dictate levels of gene expression to confer susceptibility to common traits. Genome Res. 24, 1–13 (2014).
Segrè, A.V. et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Lage, K. et al. A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat. Biotechnol. 25, 309–316 (2007).
Rossin, E.J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Lundby, A. et al. Annotation of loci from genome-wide association studies using tissue-specific quantitative interaction proteomics. Nat. Methods 10.1038/nmeth.2997 (22 June 2014).
Yoshida, M. et al. Impaired Ca2+ store functions in skeletal and cardiac muscle cells from sarcalumenin-deficient mice. J. Biol. Chem. 280, 3500–3506 (2005).
Shimura, M. et al. Sarcalumenin alleviates stress-induced cardiac dysfunction by improving Ca2+ handling of the sarcoplasmic reticulum. Cardiovasc. Res. 77, 362–370 (2008).
Jiao, Q. et al. Sarcalumenin is essential for maintaining cardiac function during endurance exercise training. Am. J. Physiol. Heart Circ. Physiol. 297, H576–H582 (2009).
Splawski, I. et al. Severe arrhythmia disorder caused by cardiac L-type calcium channel mutations. Proc. Natl. Acad. Sci. USA 102, 8089–8096, discussion 8086–8088 (2005).
Milberg, P. et al. Inhibition of the Na+/Ca2+ exchanger suppresses torsades de pointes in an intact heart model of long QT syndrome-2 and long QT syndrome-3. Heart Rhythm 5, 1444–1452 (2008).
Milberg, P. et al. Acute inhibition of the Na+/Ca2+ exchanger reduces proarrhythmia in an experimental model of chronic heart failure. Heart Rhythm 9, 570–578 (2012).
Pott, C. et al. Proarrhythmia in a non-failing murine model of cardiac-specific Na+/Ca 2+ exchanger overexpression: whole heart and cellular mechanisms. Basic Res. Cardiol. 107, 247 (2012).
Braz, J.C. et al. PKC-α regulates cardiac contractility and propensity toward heart failure. Nat. Med. 10, 248–254 (2004).
Sakuntabhai, A. et al. Mutations in ATP2A2, encoding a Ca2+ pump, cause Darier disease. Nat. Genet. 21, 271–277 (1999).
Ji, Y. et al. Disruption of a single copy of the SERCA2 gene results in altered Ca2+ homeostasis and cardiomyocyte function. J. Biol. Chem. 275, 38073–38080 (2000).
Pani, B. et al. Up-regulation of transient receptor potential canonical 1 (TRPC1) following sarco(endo)plasmic reticulum Ca2+ ATPase 2 gene silencing promotes cell survival: a potential role for TRPC1 in Darier's disease. Mol. Biol. Cell 17, 4446–4458 (2006).
Lyon, A.R. et al. SERCA2a gene transfer decreases sarcoplasmic reticulum calcium leak and reduces ventricular arrhythmias in a model of chronic heart failure. Circ. Arrhythm. Electrophysiol. 4, 362–372 (2011).
Jin, J. et al. Deletion of Trpm7 disrupts embryonic development and thymopoiesis without altering Mg2+ homeostasis. Science 322, 756–760 (2008).
Runnels, L.W., Yue, L. & Clapham, D.E. TRP-PLIK, a bifunctional protein with kinase and ion channel activities. Science 291, 1043–1047 (2001).
Elizondo, M.R. et al. Defective skeletogenesis with kidney stone formation in dwarf zebrafish mutant for trpm7. Curr. Biol. 15, 667–671 (2005).
Arduini, B.L. & Henion, P.D. Melanophore sublineage-specific requirement for zebrafish touchtone during neural crest development. Mech. Dev. 121, 1353–1364 (2004).
Sah, R. et al. Ion channel–kinase TRPM7 is required for maintaining cardiac automaticity. Proc. Natl. Acad. Sci. USA 110, E3037–E3046 (2013).
Wei, C. et al. Calcium flickers steer cell migration. Nature 457, 901–905 (2009).
Du, J. et al. TRPM7-mediated Ca2+ signals confer fibrogenesis in human atrial fibrillation. Circ. Res. 106, 992–1003 (2010).
Sah, R. et al. Timing of myocardial trpm7 deletion during cardiogenesis variably disrupts adult ventricular function, conduction, and repolarization. Circulation 128, 101–114 (2013).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825 (2010).
Schwartz, P.J., Moss, A.J., Vincent, G.M. & Crampton, R.S. Diagnostic criteria for the long QT syndrome. An update. Circulation 88, 782–784 (1993).
Acknowledgements
A full listing of acknowledgments is provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Author contributions are indicated by cohort and group. All coauthors revised and approved the manuscript.
Writing group. C.N.-C. takes overall responsibility for the QT-IGC study. The study design was developed by M.J.A., D.E.A., A. Chakravarti, L.C., P.I.W.d.B., T.T.K., P.B.M., C.N.-C., A. Pfeufer, S.L.P., P.J.S. and N.S. in consultation with the respective study groups. The manuscript was written by C.N.-C. The manuscript was critically revised in detail by members of the writing team before circulation to all coauthors.
GWAS cohorts. AGES: Phenotyping: V.G. Data analysis: A.V.S. Oversight: T.B.H., L.J.L., V.G. Amish studies: Clinical data collection, genotyping and oversight: A.R.S. EKG data collection: W.S.P. Analysis: A. Parsa, J.R.O. Interpretation: A. Parsa, W.S.P. ARIC: Study design: A.A., D.E.A., A. Chakravarti, W.H.L.K. Analyses: D.E.A., J.S.B., A. Chakravarti, G.E., H. Huang. Steering: D.E.A., A. Chakravarti. Writing: D.E.A., A. Chakravarti. BLSA: Analysis: T.T. Phenotype collection: J.B.S. Overall project supervision: L. Ferrucci. BRIGHT: Phenotyping: M. Brown, M.J.C., P.W.M., P.B.M., N.J.S. Genotyping: P.B.M., S.J.N. Analysis: S.J.N. Overall study supervision: M. Brown, M.J.C., P.B.M., N.J.S. Carlantino: Sample and/or data collection: M.C., L.Z. Overall study supervision: P.G. Data collection and/or statistical analysis: S.U. CHS: Study design: J.C.B., S.R.H., B.M.P., N.S. Data collection: S.R.H., B.M.P., D.S.S. Genotyping: J.I.R. Analysis, interpretation: J.C.B., N.S. Supervision of analyses: B.M.P. Funding for GWAS: B.M.P. Croatia-Korcula and Croatia-Split: GWAS analysis: C.H. Data collection, phenotype measurement, data entry and field work supervision: I.K., O.P. Study design and funding: I.R., A.F.W. DCCT/EDIC: Analyses: D.W. Supervision of analyses: A.D.P. deCODE: Data collection: D.O.A., H. Hólm. Study design: K. Stefansson, U.T., H. Hólm, D.F.G., D.O.A. Data alignment, imputation and statistical analysis: D.F.G. Additional analysis and interpretation of results: K. Stefansson, U.T., H. Hólm, D.F.G., D.O.A. eMERGE: Data curation and GWAS analysis: R.L.Z., Y.B. Supervision of quality control and analysis of data set: M.D.R. Study conception and analysis framework: D.M.R. Algorithm for case ascertainment: J.C.D. ERF: Analysis: A. Isaacs. Data acquisition: C.M.v.D., A. Isaacs, B.A.O., J.A.K., A.G.U. Overall study principal investigators: C.M.v.D., B.A.O. FHS: Analysis plan development: C.N.-C., C.J.O., P.A.N., M.G.L. GWAS analysis: X.Y., M.G.L. Secured funding: C.N.-C., C.J.O. FVG: Data collection: M. Bobbo. Primary analysis: A. Iorio. Statistical analysis: A.P.D.A. Overall study supervision: G.S. Health2000: Data analysis, replication genotyping and quality control: A.M.L. Primary data analysis: A. Marjamaa. Phenotyping, including ECGs: A.J. Electrocardiographic measurements: K.P. GWAS and replication genotyping: M.P. Design of ECG study, analysis and interpretation: L.O. Genetic data collection and analysis: K.K.K. Principal investigator and supervision: V.S. HealthABC: Data collection and supervision: S.R.C., Y.L. Data analysis: D.S.E., M.A.N. HNR: Data collection: H.K. Data generation: H.K., T.W.M. Genetic data generation: M.M.N., P.H., T.W.M. Data analysis: L.E., P.H., T.W.M., M.M.N. Overall study design and principal investigators: R.E., K.-H.J. KORA-F3/S4: Overall QT project supervision: A. Pfeufer. Genotyping oversight: T.M. ECG collection, measurement and interpretation: M.F.S., S. Perz, B.M.B., E.M. Primary genetic analysis: C.G., M.M.-N. Interpretation of results: C.G., A. Pfeufer, H.P., S. Kääb, T.M., M.W. Overall study principal investigator: A. Peters. LifeLines: Phenotyping: R.A.d.B., P.A.v.d.V. Genotyping: L. Franke. Analyses: I.M.N. and L. Franke. MICROS: Sample recruitment and overall study principal investigator: P.P.P. Study supervision, genotyping and data coordination: A.A.H. Data analysis: F.D.G.M., C.F. ORCADES: Phenotype collection: S.H.W. Genotype generation: H.C., J.F.W. GWAS analysis: P.N. Raised funding: J.F.W. Overall study supervision: J.F.W. PopGen: Recruitment and phenotyping: N.E.E.M., N.F. Genotyping and data preparation: A.F. Data preparation and analysis: D.E. PREVEND: Phenotyping: M.P.v.d.B., D.J.v.V., G.N. Genotyping and data analysis: F.W.A., I.M.L., P.v.d.H. Obtained funding: G.N., D.J.v.V., F.W.A., P.v.d.H. Rotterdam Study I and II: Study concept and design: M.E., B.H.S. Data acquisition: M.E., O.H.F., B.P.K., J.A.K., A.H., J.C.M.W., B.H.S., A.G.U. Statistical analysis: M.E. Interpretation: M.E., B.H.S. Obtained funding: A.H., J.C.M.W., A.G.U., B.H.S. Study supervision: B.H.S. SardiNIA: Phenotyping: M.O. Genotyping and data analysis: G.R.A., E.G.L., A. Mulas, M.O., S.S., D.S., K.V.T., M.U. Overall study supervision and principal investigators: D.S., M.U. SHIP: Data acquisition: M.D., M.R.P.M., U.V., S.B.F. Statistical analysis: U.V., M.D., M.R.P.M. Interpretation: U.V., M.D., S.B.F. Obtained funding: S.B.F., U.V. TwinsUK: Study concept and design: H.S., Y.J. Data acquisition: T.D.S. Statistical analysis and interpretation: I.M.N., H.S., Y.J. Obtained funding: Y.J., T.D.S. Young Finns Study: Data collection: T.J.L., O.T.R., M. Kähönen, J.S.V. Genotyping: T.J.L., N.M. Genotyping: T.J.L. Phenotype preparation: O.T.R., M. Kähönen, J.S.V. Analysis: T.J.L., O.T.R., M. Kähönen, J.S.V., L.-P.L. Obtained funding: T.J.L., O.T.R., M. Kähönen, J.S.V.
Directly genotyped SNP replication cohorts. (Author contributions for cohorts that contributed to both GWAS and replication genotyping are shown under the GWAS entry above.) BRHS: Analysis: R.W.M. Custodian of genetic resource: R.W.M., P.H.W. Data collection for genetic resource: P.H.W. Development of genetic resource: A.D.H. Overall study supervision and principal investigators: P.H.W., R.W.M. ECG analyses: P.W.M. Bruneck: Data analysis, interpretation and writing: S. Kiechl. DNA preparation: F. Kronenberg, C.L. ECG measurement and database: M. Knoflach. Supervision, funding, administration and principal investigator: J.W. Carla: Study concept and design: K.H.G., K.W. Genotyping: H.M.z.S. Supervision: K.W. Study design and analysis: A.K. Study concept, supervision and principal investigator: J.H. Cyprus: Study concept, funding, supervision and analysis: A.N.N. Data acquisition, analysis and interpretation: M.G. Genetic, biochemical data acquisition and statistical analysis: A.G.P. Czech Post-MONICA: Data collection and submission: J.A.H., V.A. Galicia: Cohort collection: M. Brion. Study design: M. Brion. Genotyping platform management: A. Carracedo. Genotyping: M.T. Analysis: M.T. Interpretation: M. Brion, A. Carracedo. Financial support: A. Carracedo. Intergene: Genotyping, data analysis and epidemiology expertise: F.N. Genotyping and genetic expertise: Å.T.N. Study design, data collection and disease area knowledge: D.S.T. MIDSPAN Family Study: Data acquisition, statistical analysis and interpretation: S. Padmanabhan. Genotyping: W.K.L. Overall study supervision, data collection and funding: A.F.D., G.C.M.W. PIVUS: Genotyping: A.-C.S. Phenotyping: L.L., J.Ä., J.S. Data analysis: S.G. Overall supervision and principal investigator: E.I. SAPHIR: DNA preparation: F. Kronenberg, C.L. Data collection: L.K., B.P., B.S. Data analysis: L.K., F. Kronenberg, C.L., B.P., B.S. Study design and principal investigator: B.P. ULSAM: Genotyping: A.-C.S. Phenotyping: L.L., J.Ä., J.S. Data analysis: S.G. Overall supervision and principal investigator: E.I. Whitehall II: Data collection and submission: M. Kumari. Overall supervision: M. Kivimaki. Funding: A.D.H.
Meta-analysis of GWAS and replication. D.E.A. and S.L.P. independently performed quality control and meta-analysis of GWAS and replication association results. Polygenic analysis: R.D.K. P.I.W.d.B., A. Pfeufer. and C.N.-C. supervised the analyses.
Non-QT trait lookups. CARe-COGENT: Meta-analysis and lookup: J.G.S. HRGEN: Meta-analysis and lookup: M.d.H. Overall study supervision: R.J.F.L. QRS GWAS: Study supervision: N.S. Meta-analysis: D.E.A., P.I.W.d.B. Results lookup: S.L.P.
Non-cardiac eQTL analyses. Data set acquisition: A.S.P., V.E. Analysis and interpretation: A.D.J. Cell type–specific enrichment tests: S.R., K. Slowikowski.
Left ventricle eQTL analyses. Overall supervision: T.P.C. Recruitment, sample collection: K.M., C.E.M. Sample processing and expression analysis: J. Brandimarto. Statistical analysis: M.M.
Left ventricle enhancer analyses. Analysis: X.W. Overall supervision: L.A.B., M. Kellis.
Mouse knockout. Enrichment tests: S.R., K. Slowikowski. Candidate gene list: P.v.d.H.
LQTS mutation screening. Amsterdam: SLC8A1 sequencing: T.T.K. Clinical data collection: A.B., N.H., A.A.M.W. Study supervision: C.R.B., A.A.M.W. London: Recruitment, phenotyping and strategy: E.R.B. Screening for mutations in LQT1, LQT2 and LQT3 and sample management: C.D. Mayo Clinic: LQTS cohort characteristic organization: D.J.T. TRPM7 mutation analysis and interpretation: D.J.T., A.M.-D., J.R.G. Patient collection, study design, data review and overall supervision: M.J.A. Munich: Study oversight: S. Kääb, A. Pfeufer. Patient collection: B.M.B., E.M. Genotyping: H.P. Nantes: Scientific management: J.-J.S. Clinical, genetic information collection: S.C. ATP2A2 sequencing: S.C. Screening for mutations in LQT1, LQT2 and LQT3 and clinical data collection: J. Barc. LQTS gene diagnosis management: F. Kyndt. Patient enrollment: V.P. Pavia: Patient collection, patient selection and molecular screening supervision: L.C., P.J.S. SRL mutation screening: A.G., R.I. Toronto: Identification of patients with LQTS free of LQT1, LQT2 and LQT3 mutations: R.M.H. Program codevelopment: S.W.S.
DAPPLE analysis. Concept, design and analysis: E.J.R. Supervision: K.L., M.J.D.
Immunoprecipitation experiments. Proteomic experiments and analysis: A.L. Overall study supervision: J.V.O.
Corresponding author
Ethics declarations
Competing interests
H. Hólm is a former and D.O.A., D.F.G., K. Stefansson and U.T. are current full or part-time employees of deCODE Genetics/Amgen, Inc. A.S.P. was previously an employee of Merck Research Laboratory and is a current employee of Sanofi. F.N. is an employee of AstraZeneca. B.P. serves on the data and safety monitoring board for a clinical trial funded by the manufacturer (Zoll LifeCor) and on the Steering Committee of the Yale Open Data Access Project funded by Johnson & Johnson. J.S. serves on the advisory board of Itrim. M.J.A. is a consultant for Boston Scientific, Gilead Sciences, Medtronic and St. Jude Medical. M.J.A. and the Mayo Clinic receive royalties from Transgenomic with respect to their FAMILION-LQTS and FAMILION-CPVT genetic tests.
Additional information
Full list of members and affiliations appear in the Supplementary Note.
Full list of members and affiliations appear in the Supplementary Note.
Full list of members and affiliations appear in the Supplementary Note.
Full list of members and affiliations appear in the Supplementary Note.
Full list of members and affiliations appear in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–3 and 5–18, and Supplementary Note (PDF 12908 kb)
Supplementary Table 4
Metabochip QT replication + fine-mapping of SNP content. (XLSX 835 kb)
Source data
Rights and permissions
About this article
Cite this article
Arking, D., Pulit, S., Crotti, L. et al. Genetic association study of QT interval highlights role for calcium signaling pathways in myocardial repolarization. Nat Genet 46, 826–836 (2014). https://doi.org/10.1038/ng.3014
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3014
This article is cited by
-
Stem cell models of inherited arrhythmias
Nature Cardiovascular Research (2024)
-
Outlining cardiac ion channel protein interactors and their signature in the human electrocardiogram
Nature Cardiovascular Research (2023)
-
Dosage of the pseudoautosomal gene SLC25A6 is implicated in QTc interval duration
Scientific Reports (2023)
-
Thyroid hormones regulate cardiac repolarization and QT-interval related gene expression in hiPSC cardiomyocytes
Scientific Reports (2022)
-
Genetic insights into cardiac relaxation and filling
Nature Cardiovascular Research (2022)