Abstract
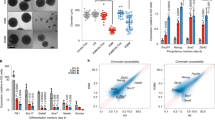
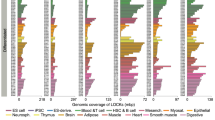
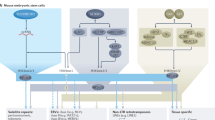
Higher eukaryotes must adapt a totipotent genome to specialized cell types with stable but limited functions. One potential mechanism for lineage restriction is changes in chromatin, and differentiation-related chromatin changes have been observed for individual genes1,2. We have taken a genome-wide view of histone H3 lysine 9 dimethylation (H3K9Me2) and find that differentiated tissues show surprisingly large K9-modified regions (up to 4.9 Mb). These regions are highly conserved between human and mouse and are differentiation specific, covering only ∼4% of the genome in undifferentiated mouse embryonic stem (ES) cells, compared to 31% in differentiated ES cells, ∼46% in liver and ∼10% in brain. These modifications require histone methyltransferase G9a and are inversely related to expression of genes within the regions. We term these regions large organized chromatin K9 modifications (LOCKs). LOCKs are substantially lost in cancer cell lines, and they may provide a cell type–heritable mechanism for phenotypic plasticity in development and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Spivakov, M. & Fisher, A.G. Epigenetic signatures of stem-cell identity. Nat. Rev. Genet. 8, 263–271 (2007).
Atkinson, S.P. et al. Epigenetic marking prepares the human HOXA cluster for activation during differentiation of pluripotent cells. Stem Cells 26, 1174–1185 (2008).
Kim, T.H. et al. A high-resolution map of active promoters in the human genome. Nature 436, 876–880 (2005).
Kim, T.H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007).
Wallace, J.A. & Felsenfeld, G. We gather together: insulators and genome organization. Curr. Opin. Genet. Dev. 17, 400–407 (2007).
Ward, C.M., Barrow, K., Woods, A.M. & Stern, P.L. The 5T4 oncofoetal antigen is an early differentiation marker of mouse ES cells and its absence is a useful means to assess pluripotency. J. Cell Sci. 116, 4533–4542 (2003).
Tachibana, M. et al. G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev. 16, 1779–1791 (2002).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Dennis, G. Jr . et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, 3 (2003).
Heintz, E. Das heterochromatin der moose. Jahresber Wiss Botanik 69, 762–818 (1928).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
O'Geen, H. et al. Genome-wide analysis of KAP1 binding suggests autoregulation of KRAB-ZNFs. PLoS Genet. 3, e89 (2007).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Cai, S., Lee, C.C. & Kohwi-Shigematsu, T. SATB1 packages densely looped, transcriptionally active chromatin for coordinated expression of cytokine genes. Nat. Genet. 38, 1278–1288 (2006).
Fraser, P. & Bickmore, W. Nuclear organization of the genome and the potential for gene regulation. Nature 447, 413–417 (2007).
Pajerowski, J.D., Dahl, K.N., Zhong, F.L., Sammak, P.J. & Discher, D.E. Physical plasticity of the nucleus in stem cell differentiation. Proc. Natl. Acad. Sci. USA 104, 15619–15624 (2007).
Wiblin, A.E., Cui, W., Clark, A.J. & Bickmore, W.A. Distinctive nuclear organisation of centromeres and regions involved in pluripotency in human embryonic stem cells. J. Cell Sci. 118, 3861–3868 (2005).
Efroni, S. et al. Global transcription in pluripotent embryonic stem cells. Cell Stem Cell 2, 437–447 (2008).
Feinberg, A.P. Phenotypic plasticity and the epigenetics of human disease. Nature 447, 433–440 (2007).
Goetze, S. et al. The three-dimensional structure of human interphase chromosomes is related to the transcriptome map. Mol. Cell. Biol. 27, 4475–4487 (2007).
Cremer, M. et al. Inheritance of gene density-related higher order chromatin arrangements in normal and tumor cell nuclei. J. Cell Biol. 162, 809–820 (2003).
Schneider, R. & Grosschedl, R. Dynamics and interplay of nuclear architecture, genome organization, and gene expression. Genes Dev. 21, 3027–3043 (2007).
Reddy, K.L., Zullo, J.M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
Anderson, D.J. & Hetzer, M.W. The life cycle of the metazoan nuclear envelope. Curr. Opin. Cell Biol. 20, 386–392 (2008).
Acknowledgements
We thank S. Taverna, K. Reddy, R. Ohlsson and C. Sapienza for helpful discussions, and S. Taverna and W. Timp for assistance with illustration. This work was supported by US National Institutes of Health grant P50HG003233 to A.P.F.
Author information
Authors and Affiliations
Contributions
B.W. and A.P.F. conceived the project. B.W. performed the experiments. H.W. analyzed the data with the guidance of R.A.I. Y.S. provided the mouse G9a knockout and control ES cell lines. A.P.F. supervised the experiments and wrote the paper with B.W.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Tables 1–8 and Supplementary Figures 1–11 (PDF 478 kb)
Supplementary Data 1
H3K9Me2 LOCKs in human placenta (XLS 24 kb)
Supplementary Data 2
H3K9Me2 LOCKs in mouse (XLS 1258 kb)
Supplementary Data3
H3K9Me2 LOCKs in WT and G9a−/− day 18-differentiated ES cells (XLS 28 kb)
Supplementary Data 4
Individual H3K9Me2 and H3K9Me3 marks in WT and G9a−/− day 18-differentiated ES cells (XLS 75 kb)
Rights and permissions
About this article
Cite this article
Wen, B., Wu, H., Shinkai, Y. et al. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat Genet 41, 246–250 (2009). https://doi.org/10.1038/ng.297
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.297
This article is cited by
-
Diverse silent chromatin states modulate genome compartmentalization and loop extrusion barriers
Nature Structural & Molecular Biology (2023)
-
Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis
Nature Genetics (2023)
-
ZFP462 safeguards neural lineage specification by targeting G9A/GLP-mediated heterochromatin to silence enhancers
Nature Cell Biology (2023)
-
InterLINCing Chromatin Organization and Mechanobiology in Laminopathies
Current Cardiology Reports (2023)
-
The patterns and participants of parental histone recycling during DNA replication in Saccharomyces cerevisiae
Science China Life Sciences (2023)