Abstract
Loss-of-function mutations protective against human disease provide in vivo validation of therapeutic targets1,2,3, but none have yet been described for type 2 diabetes (T2D). Through sequencing or genotyping of ∼150,000 individuals across 5 ancestry groups, we identified 12 rare protein-truncating variants in SLC30A8, which encodes an islet zinc transporter (ZnT8)4 and harbors a common variant (p.Trp325Arg) associated with T2D risk and glucose and proinsulin levels5,6,7. Collectively, carriers of protein-truncating variants had 65% reduced T2D risk (P = 1.7 × 10−6), and non-diabetic Icelandic carriers of a frameshift variant (p.Lys34Serfs*50) demonstrated reduced glucose levels (−0.17 s.d., P = 4.6 × 10−4). The two most common protein-truncating variants (p.Arg138* and p.Lys34Serfs*50) individually associate with T2D protection and encode unstable ZnT8 proteins. Previous functional study of SLC30A8 suggested that reduced zinc transport increases T2D risk8,9, and phenotypic heterogeneity was observed in mouse Slc30a8 knockouts10,11,12,13,14,15. In contrast, loss-of-function mutations in humans provide strong evidence that SLC30A8 haploinsufficiency protects against T2D, suggesting ZnT8 inhibition as a therapeutic strategy in T2D prevention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Nassar, M.A. et al. Nociceptor-specific gene deletion reveals a major role for Nav1.7 (PN1) in acute and inflammatory pain. Proc. Natl. Acad. Sci. USA 101, 12706–12711 (2004).
Cohen, J. et al. Low LDL cholesterol in individuals of African descent resulting from frequent nonsense mutations in PCSK9. Nat. Genet. 37, 161–165 (2005).
Sullivan, D. et al. Effect of a monoclonal antibody to PCSK9 on low-density lipoprotein cholesterol levels in statin-intolerant patients: the GAUSS randomized trial. J. Am. Med. Assoc. 308, 2497–2506 (2012).
Chimienti, F. et al. In vivo expression and functional characterization of the zinc transporter ZnT8 in glucose-induced insulin secretion. J. Cell Sci. 119, 4199–4206 (2006).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Strawbridge, R.J. et al. Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes. Diabetes 60, 2624–2634 (2011).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Nicolson, T.J. et al. Insulin storage and glucose homeostasis in mice null for the granule zinc transporter ZnT8 and studies of the type 2 diabetes–associated variants. Diabetes 58, 2070–2083 (2009).
Rutter, G.A. Think zinc: new roles for zinc in the control of insulin secretion. Islets 2, 49–50 (2010).
Lemaire, K. et al. Insulin crystallization depends on zinc transporter ZnT8 expression, but is not required for normal glucose homeostasis in mice. Proc. Natl. Acad. Sci. USA 106, 14872–14877 (2009).
Pound, L.D. et al. Deletion of the mouse Slc30a8 gene encoding zinc transporter-8 results in impaired insulin secretion. Biochem. J. 421, 371–376 (2009).
Wijesekara, N. et al. Beta cell–specific Znt8 deletion in mice causes marked defects in insulin processing, crystallisation and secretion. Diabetologia 53, 1656–1668 (2010).
Pound, L.D. et al. The physiological effects of deleting the mouse Slc30a8 gene encoding zinc transporter-8 are influenced by gender and genetic background. PLoS ONE 7, e40972 (2012).
Hardy, A.B. et al. Effects of high-fat diet feeding on Znt8-null mice: differences between β-cell and global knockout of Znt8. Am. J. Physiol. Endocrinol. Metab. 302, E1084–E1096 (2012).
da Silva Xavier, G., Bellomo, E.A., McGinty, J.A., French, P.M. & Rutter, G.A. Animal models of GWAS-identified type 2 diabetes genes. J. Diabetes Res. 2013, 906590 (2013).
van de Bunt, M. & Gloyn, A.L. From genetic association to molecular mechanism. Curr. Diab. Rep. 10, 452–466 (2010).
Plenge, R.M., Scolnick, E.M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J.A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009).
Guey, L.T. et al. Power in the phenotypic extremes: a simulation study of power in discovery and replication of rare variants. Genet. Epidemiol. doi:10.1002/gepi.20572 (9 February 2011).
Chimienti, F., Devergnas, S., Favier, A. & Seve, M. Identification and cloning of a β-cell-specific zinc transporter, ZnT-8, localized into insulin secretory granules. Diabetes 53, 2330–2337 (2004).
Tamaki, M. et al. The diabetes-susceptible gene SLC30A8/ZnT8 regulates hepatic insulin clearance. J. Clin. Invest. 123, 4513–4524 (2013).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Gudmundsson, J. et al. A study based on whole-genome sequencing yields a rare variant at 8q24 associated with prostate cancer. Nat. Genet. 44, 1326–1329 (2012).
Bloom, J. & Pagano, M. Experimental tests to definitively determine ubiquitylation of a substrate. Methods Enzymol. 399, 249–266 (2005).
Waters, P.J. Degradation of mutant proteins, underlying “loss of function” phenotypes, plays a major role in genetic disease. Curr. Issues Mol. Biol. 3, 57–65 (2001).
Mathieson, I. & McVean, G. Differential confounding of rare and common variants in spatially structured populations. Nat. Genet. 44, 243–246 (2012).
Price, A. L., Zaitlen, N. A., Relch, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Saxena, R. et al. Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nat. Genet. 42, 142–148 (2010).
Acknowledgements
This manuscript is dedicated to the memory of David R. Cox, our dear friend and colleague, who was relentlessly supportive of this work—and more generally, of the use of human genetics to improve human health. He is missed, but his legacy goes on. We gratefully acknowledge the contribution of all ∼150,000 participants from the various population studies that contributed to this work. J.F. was supported in part by US National Institutes of Health (NIH) Training Grant 5-T32-GM007748-33. D.A. was supported by funding from the Doris Duke Charitable Foundation (2006087). N.L.B. was supported by a Fulbright Diabetes UK Fellowship (BDA 11/0004348). This work was supported in part by funding to the Broad Institute (principal investigator D.A.) from Pfizer, Inc. Funding for the GoT2D and T2D-GENES studies was provided by grants 5U01DK085526 (NIH/NIDDK; Multiethnic Study of Type 2 Diabetes Genes), DK088389 (NIH/NIDDK; Low-Pass Sequencing and High-Density SNP Genotyping for Type 2 Diabetes) and U54HG003067 (National Human Genome Research Institute (NHGRI); Large-Scale Sequencing and Analysis of Genomes), as well as by NIH grants U01 DK085501, U01 DK085524, U01 DK085545 and U01 DK085584. The Malmö Preventive Project and the Scania Diabetes Registry were supported by grants from the Swedish Research Council (Dnr 521-2010-3490 to L.G. and Dnr 349-2006-237 to the Lund University Diabetes Centre), as well as by a European Research Council (ERC) grant (GENETARGET T2D, GA269045) and two European Union grants (ENGAGE (2007-201413) and CEED3 (2008-223211)) to L.G. The Botnia study was supported by funding from the Sigrid Juselius Foundation and the Folkhälsan Research Foundation. P.R.N. was funded by the ERC (AdG 293574), the Research Council of Norway (197064/V50), the KG Jebsen Foundation, the University of Bergen, the Western Norway Health Authority, the European Association for the Study of Diabetes Sabbatical Leave Programme and Innovest. The Danish studies were supported by the Lundbeck Foundation (Lundbeck Foundation Centre for Applied Medical Genomics in Personalised Disease Prediction, Prevention and Care (LuCamp); http://www.lucamp.org/) and the Danish Council for Independent Research. The Novo Nordisk Foundation Center for Basic Metabolic Research is an independent Research Center at the University of Copenhagen, partially funded by an unrestricted donation from the Novo Nordisk Foundation (http://www.metabol.ku.dk/). The PIVUS/ULSAM cohort was supported by Wellcome Trust grants WT098017, WT064890 and WT090532, Uppsala University, the Uppsala University Hospital, the Swedish Research Council and the Swedish Heart-Lung Foundation. The METSIM study was supported by the Academy of Finland (contract 124243), the Finnish Heart Foundation, the Finnish Diabetes Foundation, Tekes (contract 1510/31/06), the Commission of the European Community (HEALTH-F2-2007-201681) and grants R01DK062370 and R01DK072193 (NIH/NIDDK) and grant Z01HG000024 (NHGRI). The FUSION study was supported by grants R01DK062370 and R01DK072193 (NIH/NIDDK) and grant Z01HG000024 (NHGRI). The DR's EXTRA Study was supported by the Ministry of Education and Culture of Finland (627; 2004-2011), the Academy of Finland (102318; 123885), Kuopio University Hospital, the Finnish Diabetes Association, the Finnish Heart Association and the Päivikki and Sakari Sohlberg Foundation and by grants from the European Commission Framework Programme 6 Integrated Project (EXGENESIS); LSHM-CT-2004-005272, City of Kuopio and Social Insurance Institution of Finland (4/26/2010). V. Salomaa is funded by the Academy of Finland, grant 139635, and by the Finnish Foundation for Cardiovascular Disease. Sequencing and genotyping of British individuals was supported by Wellcome Trust grants WT090367, WT090532 and WT098381 and by NIH/NIDDK grant U01-DK085545. Funding for the Jackson Heart Study (JHS) was provided by the National Heart, Lung, and Blood Institute (NHLBI) and by the National Institute on Minority Health and Health Disparities (N01 HC-95170, N01 HC-95171 and N01 HC-95172). A.P.M. acknowledges support from Wellcome Trust grants WT098017, WT090532 and WT064890. F.V.-S. and H.S. were supported by the European Union Seventh Framework Programme, DIAPREPP (Diabetes Type 1 Prediction, Early Pathogenesis and Prevention, grant agreement 202013), and by the Swedish Child Diabetes Foundation (Barndiabetesfonden).
Author information
Authors and Affiliations
Consortia
Contributions
This manuscript describes an analysis spanning four initially distinct sequencing studies—a collaborative project between Pfizer, Massachusetts General Hospital, the Broad Institute and Lund University entitled “Towards Therapeutic Targets for Type 2 Diabetes and Myocardial Infarction in the Background of Type 2 Diabetes” (PMBL), an effort by deCODE Genetics to use whole-genome sequencing and imputation to identify and genotype over 35 million variants in up to 370,000 Icelanders23, the Genetics of Type 2 Diabetes (GoT2D) project and the Type 2 Diabetes Genetic Exploration by Next-Generation Sequencing in Multi-Ethnic Samples (T2D-GENES) project—as well as four additional genotyping efforts. The overall study bringing together data from these efforts was coordinated by J.F. and D.A., with final analysis combining data from all variants collected by J.F. The manuscript was written by J.F., D.A. and K.S., and all authors reviewed, edited and approved the manuscript. Author contributions specific to the sequencing or genotyping studies are as follows.
Pfizer, Massachusetts General Hospital, Broad Institute and Lund University. Study design. B.F.V., F.B., S.P., N.P.B., D.R.C., T.R., L.G., S.K. and D.A. Clinical investigation and sample management. T. Forsén, B.I., T. Tuomi, L.G. and J. Kravic. Sequencing, genotyping and data processing. N.P.B., T. Fennell and S.G.; performed at the Broad Institute. Analysis. J.F. (p.Arg138*) and A.V.S. (p.Trp325Arg). Functional studies. N.L.B., S.B.R.J., Z.D. (cellular models), F.V.-S., H.S. and D.A.D. (screen for carrier autoantibodies). Leadership and management. N.P.B., A.-M.R., J. Brosnan, J.K.T., S.K., T.R., L.G., D.R.C. and D.A.
deCODE Genetics. Clinical investigation and sample management. A.B.H. and R.B. Sequencing, genotyping and data processing. G.T., V. Steinthorsdottir, P.S., G.M., D.F.G. and A.K.; performed at deCODE Genetics. Analysis. G.T., V. Steinthorsdottir, P.S., G.M., D.F.G. and A.K. Leadership and management. A.K., U.T. and K.S.
T2D-GENES and Go-T2D. Clinical investigation and sample management. G.A., J. Blangero, D.W.B., J.C., Y.S.C., R.D., B.G., C.H., J. Kooner, M.L., T.M., J.-Y.L., E.S.T., Y.Y.T. and J.G.W. Sequencing and data processing. N.P.B., Y.F., T. Fennell and S.G.; performed at the Broad Institute. Analysis. J.F. (SLC30A8), P.F., A.P.M. and T. Teslovich (exome wide). Leadership and management. M.I.M., M.B. and D.A. HUNT2 sample management, genotyping, analysis and leadership involved K.H., A. Molven, S.J. and P.R.N. Danish sample management, genotyping, analysis and leadership involved N.G., R.R.-M., M.E.J., C.C., I.B., A.L., T.J., T.H. and O.P. PIVUS/ULSAM sample management, genotyping, analysis and leadership involved A. Mahajan, C.M.L., L.L., E.I. and A.P.M. Finnish sample management, genotyping, analysis and leadership involved C.F., H.M.S., M.L., K.L.M., R.R., V. Salomaa, J.T. and M.B.
Corresponding authors
Ethics declarations
Competing interests
G.T., V. Steinthorsdottir, P.S., G.M., D.F.G., A.K., U.T. and K.S. are employed by deCODE Genetics/Amgen, Inc. S.P., A.-M.R., J. Brosnan, J.K.T., T.R. and D.R.C. are employees of Pfizer, Inc. F.B. is a former employee of Pfizer, Inc., and retains shares in the company. All other authors declare no competing financial interests.
Additional information
Full lists of members and affiliations appear in the Supplementary Note.
Full lists of members and affiliations appear in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 Technical quality control metrics of targeted sequencing.
We computed various metrics to evaluate the quality of the initial sequencing experiment. Shown from the left are the distributions (over all sequenced samples) of the number of variant sites with 10x sequence coverage and hence nonmissing genotypes (Call Rate), the number of minor alleles in genotypes called across all variant sites (Minor Alleles), the number of minor alleles at sites where no other samples have minor alleles (Singletons), the fraction of genotypes called heterozygous (Heterozygosity), the ratio of the number of heterozygous sites to the number of minor allele homozygous sites (Het to Hom ratio), the fraction of sequence reads with the minor allele (averaged over all heterozygous sites; Allele Balance) and the fraction of minor allele genotypes identical to those called from Exome Chip or Metabochip genotyping (Concordance). Absent differential technical artifacts or population structure, distributions are expected to be similar between cases and controls. Statistical comparison between case and control distributions was performed via a Kruskal-Wallis one-way analysis of variance test.
Supplementary Figure 2 Association with T2D of nonsynonymous variants from the initial sequencing experiment.
Based on data from the initial sequencing experiment, we performed two types of as- sociation tests for all low-frequency (below 1%) nonsynonymous variants. First, variants were tested individually using a linear mixed model. Second, variants were collapsed within each gene and tested for aggregate association using the same mixed model approach. Shown are QQ plots of association for each approach: the x-axis plots the expected distribution of (−log10) P-values under the null model whereby no association exists across the entire experiment (e.g. the uniform distribution); the y-axis plots the observed distribution of (−log10) p-values. The blue lines show estimated 95% confidence intervals for observed distributions under the null model. For each plot, only variants (or, analogously, genes) for which ten or more carriers are observed are plotted: for variants with small numbers of counts, the uniform distribution is not a good approximation to the P -value distribution expected under the null model.
Supplementary Figure 3 Association with T2D of 71 variants from small-scale genotyping.
We genotyped select variants from the initial sequencing experiment in up to 13,884 additional Finnish and Swedish individuals. Shown is a QQ plot of associations (with the same format as in Supplementary Figure 2), tested via a logistic regression.
Supplementary Figure 4 Frequency of the p.Arg138* variant in Europe.
The p.Arg138* variant was genotyped in individuals from Finland, Sweden, Denmark, the UK, and Poland (through the Illumina Human Exome Array or custom Sequenom genotyping), as well as Iceland (through whole genome sequencing). Shown are the observed frequencies of the variant in each country; frequency estimates from Finland are stratified by the Botnia region (individuals from the Botnia cohort) and other regions of Finland. N indicates the number of individuals genotyped.
Supplementary Figure 5 Frequency of the p.Arg138* variant in Botnia.
Frequency estimates for Botnia (as computed in Supplementary Figure 4) were further stratified by town in Botnia. Information on the center of sample selection was available for all samples from the Botnia region of Finland. Individuals were grouped by study center, and frequency estimates were computed for each center separately. N indicates the number of individuals genotyped.
Supplementary Figure 6 Position of the p.Lys34Serfs*50 frameshift mutation.
Shown is the genomic position of the frameshift mutation p.Lys34Serfs*50, as visualized in the UCSC genome browser (16).
Supplementary Figure 7 Partial sequence chromatogram for the p.Lys34Serfs*50 frameshift variant.
Sanger sequencing was used to confirm carriers of the frameshift mutation. Shown is a chromatogram for one individual carrier (from Norway) heterozygous for the variant.
Supplementary Figure 8 Transcript levels of SLC30A8 variants.
nCounter analysis of SLC30A8 transcript levels in HeLa cells, following transient over-expression of C-terminal, V5-tagged ZnT8 variants and control ORFs. Data shown are mean, normalized mRNA counts ± s.e.m. of three independent plasmid transfections from two experiments. Non-specific binding is indicated by the red line. Gene expression detected by probes directed against the SLC30A8 sequence encoding the N terminus, the sequence encoding the V5 tag, and TUBB are shown.
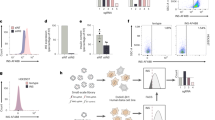
Supplementary Figure 9 Coexpression of ZnT8 variants.
HeLa cells overexpressed either (a) singly transfected C-terminal, V5-tagged ZnT8 variants or (b) cotransfections that also included C-terminally HA-tagged Trp325-ZnT8 for 24h, after which ZnT8 protein levels were observed. Cells were immunostained for ZnT8 expression using anti-HA and anti-V5 antibodies and costained with Hoechst 33342 to mark nuclei. Scale bars, 100μm (c) Protein blot analysis of HeLa cell lysates following cotransfection of Trp325-HA ZnT8 and V5-tagged ZnT8 variants. ZnT8 expression was detected using anti-V5 and anti-HA antibodies.
Supplementary Figure 10 Inhibition of protein degradation.
HeLa cells transiently overexpressing V5-tagged ZnT8 variants were treated (+) with chloroquine (100 μM) or MG132 (10 μM) or left untreated in medium absent these chemicals, over a 4-h incubation period, to inhibit lysosomal and proteasomal degradation, respectively. Protein blot analysis was performed using anti-V5 and anti-tubulin antibodies.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–12, Supplementary Figures 1–10 and Supplementary Note (PDF 9205 kb)
Supplementary Data Set 1
Sequence read data for variants called from additional sequencing. (ZIP 205 kb)
Rights and permissions
About this article
Cite this article
Flannick, J., Thorleifsson, G., Beer, N. et al. Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nat Genet 46, 357–363 (2014). https://doi.org/10.1038/ng.2915
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2915
This article is cited by
-
Urinary metal profiles in mother-offspring pairs and their association with early dysglycemia in the International Hyperglycemia and Adverse Pregnancy Outcome Follow Up Study (HAPO-FUS)
Journal of Exposure Science & Environmental Epidemiology (2023)
-
Stomach-derived human insulin-secreting organoids restore glucose homeostasis
Nature Cell Biology (2023)
-
Genetics implicates overactive osteogenesis in the development of diffuse idiopathic skeletal hyperostosis
Nature Communications (2023)
-
Prioritization of genes associated with type 2 diabetes mellitus for functional studies
Nature Reviews Endocrinology (2023)
-
Protective effect of the rare variant rs13266634C/T of the SLC30A8 gene in the genetic background of the development of type 2 diabetes in Turkish population
International Journal of Diabetes in Developing Countries (2023)