Abstract
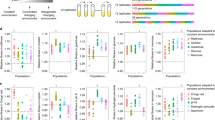
Evolution depends on genetic variation generated by mutation or recombination from standing genetic variation. In sexual organisms, little is known about the molecular population genetics of adaptation and reverse evolution1,2,3,4,5,6,7,8,9,10,11. We carry out 50 generations of experimental reverse evolution in populations of Drosophila melanogaster, previously differentiated by forward evolution, and follow changes in the frequency of SNPs in both arms of the third chromosome. We characterize the effects of sampling finite population sizes and natural selection at the genotype level. We demonstrate that selection has occurred at several loci and further that there is no general loss or gain of allele diversity. We also observe that despite the complete convergence to ancestral levels of adaptation, allele frequencies only show partial return.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Cohan, F.M. Can uniform selection retard random genetic divergence between isolated conspecific populations. Evolution Int. J. Org. Evolution 38, 495–504 (1984).
Travisano, M., Mongold, J.A., Bennett, A.F. & Lenski, R.E. Experimental tests of the roles of adaptation, chance, and history in evolution. Science 267, 87–90 (1995).
Crill, W.D., Wichman, H.A. & Bull, J.J. Evolutionary reversals during viral adaptation to alternating hosts. Genetics 154, 27–37 (2000).
Teotónio, H. & Rose, M.R. Variation in the reversibility of evolution. Nature 408, 463–466 (2000).
Bull, J.J. & Charnov, E.L. On irreversible evolution. Evolution Int. J. Org. Evolution 39, 1149–1155 (1985).
Teotónio, H. & Rose, M.R. Perspective: reverse evolution. Evolution Int. J. Org. Evolution 55, 653–660 (2001).
Teotónio, H., Matos, M. & Rose, M.R. Reverse evolution of fitness in Drosophila melanogaster. J. Evol. Biol. 15, 608–617 (2002).
Grant, P.R. & Grant, B.R. Unpredictable evolution in a 30-year study of Darwin's finches. Science 296, 707–711 (2002).
Whiting, M.F., Bradler, S. & Maxwell, T. Loss and recovery of wings in stick insects. Nature 421, 264–267 (2003).
Whitlock, M.C., Phillips, P.C. & Fowler, K. Persistence of changes in the genetic covariance matrix after a bottleneck. Evolution Int. J. Org. Evolution 56, 1968–1975 (2002).
Porter, M.L. & Crandall, K.A. Lost along the way: the significance of evolution in reverse. Trends Ecol. Evol. 18, 541–547 (2003).
Colosimo, P.F. et al. Widespread parallel evolution in sticklebacks by repeated fixation of Ectodysplasin alleles. Science 307, 1928–1933 (2005).
Teotónio, H., Matos, M. & Rose, M.R. Quantitative genetics of functional characters in Drosophila melanogaster populations subjected to laboratory selection. J. Genet. 83, 265–277 (2004).
Hudson, R.R., Sáez, A.G. & Ayala, F.J. DNA variation at the Sod locus of Drosophila meanogaster: an unfolding story of natural selection. Proc. Natl. Acad. Sci. USA 94, 7725–7729 (1997).
Deckert-Cruz, D.J., Tyler, R.H., Landmesser, J.E. & Rose, M.R. Allozymic differentiation in response to laboratory demographic selection of Drosophila melanogaster. Evolution Int. J. Org. Evolution 51, 865–872 (1997).
Verrelli, B.C. & Eanes, W.F. Clinal variation for amino acid polymorphisms at the Pgm locus in Drosophila melanogaster. Genetics 157, 1649–1663 (2001).
Balakirev, E.S., Anisimova, M. & Ayala, F.J. Positive and negative selection in the β-Esterase gene cluster of the Drosophila melanogaster subgroup. J. Mol. Evol. 62, 496–510 (2006).
Smith, J.M. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 23, 23–35 (1974).
Hartl, D.L. & Clark, A.G. Principles of Population Genetics (Sinauer, Sunderland, Massachusetts, 1989).
Macdonald, S.J., Pasten, T. & Long, A.D. The effect of polymorphisms in the Enhancer of split gene complex on bristle number variation in a large wild-caught cohort of Drosophila melanogaster. Genetics 171, 1741–1756 (2005).
Scheet, P. & Stephens, M. Fast and flexible model for LD. Am. J. Hum. Genet. 78, 629–644 (2006).
McVean, G., Awadalla, P. & Fearnhead, P. A coalescent-based method for detecting and estimating recombination rates from gene sequences. Genetics 160, 1231–1241 (2002).
Nei, M. & Tajima, F. Genetic drift and the estimation of effective population size. Genetics 98, 625–640 (1981).
Goldringer, I. & Bataillon, T. On the distribution of temporal variations in allele frequency: consequences for the estimation of effective population size and the detection of loci undergoing selection. Genetics 168, 563–568 (2004).
Nei, M. Definition and estimation of fixation indices. Evolution Int. J. Org. Evolution 40, 643–645 (1986).
Churchill, G.A. & Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics 138, 963–971 (1994).
Hermisson, J. & Pennings, P.S. Soft sweeps: molecular population genetics of adaptation from standing genetic variation. Genetics 169, 2335–2352 (2005).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2006).
Haag-Liautard, C. et al. Direct estimation of per nucleotide and genomic deleterious mutation rates in Drosophila. Nature 445, 82–85 (2007).
Acknowledgements
We thank T. Aires, D. Brites, J. Costa and I. Marques for support with SNP discovery and genotyping, and G. McVean for support with LDhat. We thank R. Azevedo, S. Carvalho, A. Coutinho, S. Estes, L. Mueller, P. Phillips, S. Proulx, M. Che Soares and É. Sucena for comments on the project and manuscript. Financial support was provided by The National Science Foundation to A.D.L. (DEB-0614429), and Fundação para a Ciência e a Tecnologia (FCT/FEDER POCTI/BSE/48228/2002) and Fundação Calouste Gulbenkian to H.T.
Author information
Authors and Affiliations
Contributions
H.T., I.M.C. and M.B. performed the experiments. H.T., I.M.C. and A.D.L. analysed the data. H.T., M.R.R. and A.D.L. conceived the project and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–6 (PDF 513 kb)
Rights and permissions
About this article
Cite this article
Teotónio, H., Chelo, I., Bradić, M. et al. Experimental evolution reveals natural selection on standing genetic variation. Nat Genet 41, 251–257 (2009). https://doi.org/10.1038/ng.289
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.289