Abstract
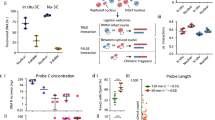
Gene expression during development and differentiation is regulated in a cell- and stage-specific manner by complex networks of intergenic and intragenic cis-regulatory elements whose numbers and representation in the genome far exceed those of structural genes. Using chromosome conformation capture, it is now possible to analyze in detail the interaction between enhancers, silencers, boundary elements and promoters at individual loci, but these techniques are not readily scalable. Here we present a high-throughput approach (Capture-C) to analyze cis interactions, interrogating hundreds of specific interactions at high resolution in a single experiment. We show how this approach will facilitate detailed, genome-wide analysis to elucidate the general principles by which cis-acting sequences control gene expression. In addition, we show how Capture-C will expedite identification of the target genes and functional effects of SNPs that are associated with complex diseases, which most frequently lie in intergenic cis-acting regulatory elements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kowalczyk, M.S. et al. Intragenic enhancers act as alternative promoters. Mol. Cell 45, 447–458 (2012).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
de Laat, W. & Dekker, J. 3C-based technologies to study the shape of the genome. Methods 58, 189–191 (2012).
Sajan, S.A. & Hawkins, R.D. Methods for identifying higher-order chromatin structure. Annu. Rev. Genomics Hum. Genet. 13, 59–82 (2012).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
van de Werken, H.J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
Schaub, M.A., Boyle, A.P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012).
Bainbridge, M.N. et al. Whole exome capture in solution with 3 Gbp of data. Genome Biol. 11, R62 (2010).
Göndör, A., Rougier, C. & Ohlsson, R. High-resolution circular chromosome conformation capture assay. Nat. Protoc. 3, 303–313 (2008).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Hughes, J.R. et al. Annotation of cis-regulatory elements by identification, subclassification, and functional assessment of multispecies conserved sequences. Proc. Natl. Acad. Sci. USA 102, 9830–9835 (2005).
De Gobbi, M. et al. Tissue-specific histone modification and transcription factor binding in α globin gene expression. Blood 110, 4503–4510 (2007).
Vavouri, T., McEwen, G.K., Woolfe, A., Gilks, W.R. & Elgar, G. Defining a genomic radius for long-range enhancer action: duplicated conserved non-coding elements hold the key. Trends Genet. 22, 5–10 (2006).
Wallace, H.A. et al. Manipulating the mouse genome to engineer precise functional syntenic replacements with human sequence. Cell 128, 197–209 (2007).
Lower, K.M. et al. Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc. Natl. Acad. Sci. USA 106, 21771–21776 (2009).
Vernimmen, D. et al. Chromosome looping at the human α-globin locus is mediated via the major upstream regulatory element (HS-40). Blood 114, 4253–4260 (2009).
Baù, D. et al. The three-dimensional folding of the α-globin gene domain reveals formation of chromatin globules. Nat. Struct. Mol. Biol. 18, 107–114 (2011).
Hughes, J.R. et al. High-resolution analysis of cis-acting regulatory networks at the α-globin locus. Phil. Trans. R. Soc. Lond. B 368, 20120361 (2013).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J.A., Wood, W.G. & Higgs, D.R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Ferreira, R. et al. Impaired in vitro erythropoiesis following deletion of the Scl (Tal1) +40 enhancer is largely compensated for in vivo despite a significant reduction in expression. Mol. Cell. Biol. 33, 1254–1266 (2013).
Ogilvy, S. et al. The SCL +40 enhancer targets the midbrain together with primitive and definitive hematopoiesis and is regulated by SCL and GATA proteins. Mol. Cell. Biol. 27, 7206–7219 (2007).
Göttgens, B. et al. The scl +18/19 stem cell enhancer is not required for hematopoiesis: identification of a 5′ bifunctional hematopoietic-endothelial enhancer bound by Fli-1 and Elf-1. Mol. Cell. Biol. 24, 1870–1883 (2004).
Flygare, J., Rayon Estrada, V., Shin, C., Gupta, S. & Lodish, H.F. HIF1α synergizes with glucocorticoids to promote BFU-E progenitor self-renewal. Blood 117, 3435–3444 (2011).
Shaw, G.C. et al. Mitoferrin is essential for erythroid iron assimilation. Nature 440, 96–100 (2006).
Amigo, J.D. et al. Identification of distal cis-regulatory elements at mouse mitoferrin loci using zebrafish transgenesis. Mol. Cell. Biol. 31, 1344–1356 (2011).
Sanyal, A., Lajoie, B.R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Hosseini, M. et al. Causes and consequences of chromatin variation between inbred mice. PLoS Genet. 9, e1003570 (2013).
Kang, J.H. et al. Genomic organization, tissue distribution and deletion mutation of human pyridoxine 5′-phosphate oxidase. Eur. J. Biochem. 271, 2452–2461 (2004).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Donnelly, P. Progress and challenges in genome-wide association studies in humans. Nature 456, 728–731 (2008).
Bush, W.S. & Moore, J.H. Chapter 11: genome-wide association studies. PLoS Comput. Biol. 8, e1002822 (2012).
Pennacchio, L.A., Bickmore, W., Dean, A., Nobrega, M.A. & Bejerano, G. Enhancers: five essential questions. Nat. Rev. Genet. 14, 288–295 (2013).
Kolovos, P., Knoch, T.A., Grosveld, F.G., Cook, P.R. & Papantonis, A. Enhancers and silencers: an integrated and simple model for their function. Epigenetics Chromatin 5, 1 (2012).
Spivak, J.L., Toretti, D. & Dickerman, H.W. Effect of phenylhydrazine-induced hemolytic anemia on nuclear RNA polymerase activity of the mouse spleen. Blood 42, 257–266 (1973).
Nichols, J., Evans, E.P. & Smith, A.G. Establishment of germ-line–competent embryonic stem (ES) cells using differentiation inhibiting activity. Development 110, 1341–1348 (1990).
Hagège, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nat. Protoc. 2, 1722–1733 (2007).
Stadhouders, R. et al. Multiplexed chromosome conformation capture sequencing for rapid genome-scale high-resolution detection of long-range chromatin interactions. Nat. Protoc. 8, 509–524 (2013).
Kassouf, M.T. et al. Genome-wide identification of TAL1's functional targets: insights into its mechanisms of action in primary erythroid cells. Genome Res. 20, 1064–1083 (2010).
Tallack, M.R. et al. A global role for KLF1 in erythropoiesis revealed by ChIP-seq in primary erythroid cells. Genome Res. 20, 1052–1063 (2010).
Papadopoulos, G.L. et al. GATA-1 genome-wide occupancy associates with distinct epigenetic profiles in mouse fetal liver erythropoiesis. Nucleic Acids Res. 41, 4938–4948 (2013).
Zhou, Z. et al. USF and NF-E2 cooperate to regulate the recruitment and activity of RNA polymerase II in the β-globin gene locus. J. Biol. Chem. 285, 15894–15905 (2010).
Horakova, A.H., Moseley, S.C., McLaughlin, C.R., Tremblay, D.C. & Chadwick, B.P. The macrosatellite DXZ4 mediates CTCF-dependent long-range intrachromosomal interactions on the human inactive X chromosome. Hum. Mol. Genet. 21, 4367–4377 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics Chapter 4, Unit 4.10 (2009).
McGowan, S.J., Hughes, J.R., Han, Z.P. & Taylor, S. MIG: Multi-Image Genome Viewer. Bioinformatics 29, 2477–2478 (2013).
Stein, L.D. Using GBrowse 2.0 to visualize and share next-generation sequence data. Brief. Bioinform. 14, 162–171 (2013).
Acknowledgements
We thank J. Davies and P. Piazza for technical suggestions. We thank M. Suciu, B. Graham and T. Milne for suggestions and critically reading the manuscript. We thank Z.-P. Han and J. Telenius for computational support. We thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics (funded by Wellcome Trust grant reference 090532/Z/09/Z and MRC Hub grant G0900747 91070) for the generation of the sequencing data. This work was supported by the MRC (UK) and by the Blood theme within Oxford Biomedical Research Centre (which is part of the National Institute for Health Research Biomedical Research Centres scheme). We also thank EpiGeneSys for support.
Author information
Authors and Affiliations
Contributions
J.R.H., D.R.H. and R.G. designed experiments. J.R.H., N.R., D.H., M.L. and M.D.G. performed experiments. J.R.H. performed bioinformatic analysis. E.G. performed statistical analysis. S.M. and S.T. provided bioinformatic support. J.R.H. and D.R.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Note and Supplementary Tables 1 and 2 (PDF 11216 kb)
Rights and permissions
About this article
Cite this article
Hughes, J., Roberts, N., McGowan, S. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat Genet 46, 205–212 (2014). https://doi.org/10.1038/ng.2871
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2871
This article is cited by
-
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Nature Communications (2024)
-
Increased enhancer–promoter interactions during developmental enhancer activation in mammals
Nature Genetics (2024)
-
3D Enhancer–promoter networks provide predictive features for gene expression and coregulation in early embryonic lineages
Nature Structural & Molecular Biology (2024)
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
3D genomics and its applications in precision medicine
Cellular & Molecular Biology Letters (2023)