Abstract
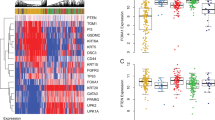
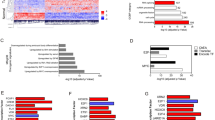
Urothelial bladder cancer (UBC) is heterogeneous at the clinical, pathological and genetic levels. Tumor invasiveness (T) and grade (G) are the main factors associated with outcome and determine patient management1. A discovery exome sequencing screen (n = 17), followed by a prevalence screen (n = 60), identified new genes mutated in this tumor coding for proteins involved in chromatin modification (MLL2, ASXL2 and BPTF), cell division (STAG2, SMC1A and SMC1B) and DNA repair (ATM, ERCC2 and FANCA). STAG2, a subunit of cohesin, was significantly and commonly mutated or lost in UBC, mainly in tumors of low stage or grade, and its loss was associated with improved outcome. Loss of expression was often observed in chromosomally stable tumors, and STAG2 knockdown in bladder cancer cells did not increase aneuploidy. STAG2 reintroduction in non-expressing cells led to reduced colony formation. Our findings indicate that STAG2 is a new UBC tumor suppressor acting through mechanisms that are different from its role in preventing aneuploidy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others

References
Sylvester, R.J. Natural history, recurrence, and progression in superficial bladder cancer. ScientificWorldJournal 6, 2617–2625 (2006).
Luis, N.M., Lopez-Knowles, E. & Real, F.X. Molecular biology of bladder cancer. Clin. Transl. Oncol. 9, 5–12 (2007).
Hernández, S. et al. FGFR3 mutations as a prognostic factor in non-muscle invasive urothelial bladder carcinomas: results of a prospective study. J. Clin. Oncol. 24, 3664–3671 (2006).
López-Knowles, E. et al. PIK3CA mutations are an early genetic alteration associated with FGFR3 mutations in superficial papillary bladder tumors. Cancer Res. 66, 7401–7404 (2006).
Real, F.X. p53: it has it all, but will it make to the clinic as a marker in bladder cancer? J. Clin. Oncol. 25, 5341–5344 (2007).
López-Knowles, E. et al. The p53 pathway and outcome among patients with T1G3 bladder tumors. Clin. Cancer Res. 12, 6029–6036 (2006).
Jebar, A.H. et al. FGFR3 and Ras gene mutations are mutually exclusive genetic events in urothelial cell carcinoma. Oncogene 24, 5218–5225 (2005).
Lindgren, D. et al. Combined gene expression and genomic profiling define two intrinsic molecular subtypes of urothelial carcinoma and gene signatures for molecular grading and outcome. Cancer Res. 70, 3463–3472 (2010).
Höglund, M. The bladder cancer genome; chromosomal changes as prognostic makers, opportunities, and obstacles. Urol. Oncol. 30, 533–540 (2012).
Blaveri, E. et al. Bladder cancer stage and outcome by array-based comparative genomic hybridization. Clin. Cancer Res. 11, 7012–7022 (2005).
Gui, Y. et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat. Genet. 43, 875–878 (2011).
Fujimoto, A. et al. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat. Genet. 44, 760–764 (2012).
Guichard, C. et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 44, 694–698 (2012).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Wilson, B.G. & Roberts, C.W. SWI/SNF nucleosome remodelers and cancer. Nat. Rev. Cancer 11, 481–492 (2011).
Solomon, D.A. et al. Mutational inactivation of STAG2 causes aneuploidy in human cancer. Science 333, 1039–1043 (2011).
Amaral, A.F.S. et al. Plasma 25-hydroxyvitamin D3 levels and bladder cancer risk according to tumor stage and FGFR3 status: a mechanism-based epidemiological study. J. Natl. Cancer Inst. 104, 1897–1904 (2012).
Welch, J.S. et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 150, 264–278 (2012).
Walter, M.J. et al. Acquired copy number alterations in adult acute myeloid leukemia genomes. Proc. Natl. Acad. Sci. USA 106, 12950–12955 (2009).
Walter, M.J. et al. Clonal architecture of secondary acute myeloid leukemia. N. Engl. J. Med. 366, 1090–1098 (2012).
Nasmyth, K. & Haering, C.H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009).
Remeseiro, S. & Losada, A. Cohesin, a chromatin engagement ring. Curr. Opin. Cell Biol. 25, 63–71 (2013).
Remeseiro, S. et al. Cohesin-SA1 deficiency drives aneuploidy and tumourigenesis in mice due to impaired replication of telomeres. EMBO J. 31, 2076–2089 (2012).
Remeseiro, S. et al. A unique role of cohesin-SA1 in gene regulation and development. EMBO J. 31, 2090–2102 (2012).
Shain, A.H. et al. Convergent structural alterations define SWItch/Sucrose NonFermentable (SWI/SNF) chromatin remodeler as a central tumor suppressive complex in pancreatic cancer. Proc. Natl. Acad. Sci. USA 109, E252–E259 (2012).
Balbás-Martínez, C. et al. ARID1A alterations are associated with FGFR3 wild type, poor-prognosis, urothelial bladder tumors. PLoS ONE 8, e62483 (2013).
Marco-Sola, S. et al. The GEM mapper: fast, accurate and versatile alignment by filtration. Nat. Methods 9, 1185–1188 (2012).
Reva, B. et al. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res. 39, e118 (2011).
Kumar, P. et al. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Gonzalez-Perez, A. et al. Functional impact bias reveals cancer drivers. Nucleic Acids Res. 40, e169 (2012).
Vazquez, M. et al. Chapter 14: cancer genome analysis. PLoS Comput. Biol. 8, e1002824 (2012).
Quesada, V. et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat. Genet. 44, 47–52 (2012).
Rodríguez-Santiago, B. et al. Mosaic uniparental disomies and aneuploidies as large structural variants of the human genome. Am. J. Hum. Genet. 87, 129–138 (2010).
Snijders, A.M. et al. Assembly of microarrays for genome-wide measurement of DNA copy number. Nat. Genet. 29, 263–264 (2001).
Acknowledgements
We thank F. Algaba, Y. Allory, A. Cuadrado, C. González, E. López, P. Lapunzina, T. Lobato, M. Malumbres, S. Remeseiro, V.J. Sánchez-Arévalo, F. Waldman and the CNIO core facilities for valuable contributions. We also thank TCGA investigators for providing unpublished information for analysis. This work was supported, in part, by grants from Ministerio de Economia y Competitividad, Madrid (grants Consolíder ONCOBIO, Consolider INESGEN, SAF-2010-21517 and SAF2011-15934-E), Instituto de Salud Carlos III (grants G03/174, 00/0745, PI051436, PI061614, G03/174, PI080440, PI120425 and Red Temática de Investigación Cooperativa en Cáncer (RTICC)), Asociación Española Contra el Cáncer, EU-FP7-201663 and US National Institutes of Health grant RO1 CA089715. C.B.-M. is the recipient of a La Caixa International PhD Fellowship. E.L. is supported by a grant from the Fundación Banco Santander Postdoctoral Programme.
Author information
Authors and Affiliations
Contributions
C.B.-M., E.L. and L.R. designed and performed in vitro functional studies. C.B.-M. performed immunohistochemical analysis of tumor samples. A.S. and C.B.-M. designed and performed mutation validation analyses. A.S. designed and prepared HaloPlex libraries for sequencing. E.C.-d.-S.-P., M.V., F.C.-G., S.B. and D.G.P. processed and analyzed exome sequencing and targeted resequencing data. J.E., E.L.-K., D.R. and S.C. performed gene copy number analyses of tumors. M.M. coordinated subject and sample data management. A.C., M.K., J.A.L. and A.T. contributed to subject recruitment and data collection. J.H. performed statistical analyses. X.L. provided technical support with subject samples. M.B. and M.G. coordinated library preparation and sequencing. O.D. contributed to library preparation and sequencing. J.C.C. and R.N.S. contributed to the in vitro analysis of the effects of STAG2 knockdown on aneuploidy. N.J. and J.L. performed pathological review of samples. I.G. coordinated exome sequencing and targeted resequencing. I.G., S.H. and A.V. supervised bioinformatics analyses. A.L. provided scientific insight and contributed with reagents. N.M. coordinated subject recruitment and the collection of clinical and pathological data and supervised clinical-pathological-molecular association and outcome analyses. F.X.R. and N.M. conceived the study. F.X.R. supervised the overall way in which the study was conducted. F.X.R. wrote the manuscript with N.M., C.B.-M., A.S., E.C.-d.-S.-P., A.V., A.L. and I.G. contributed to manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 1, 2, 5 and 7–20 (PDF 14705 kb)
Supplementary Table 3
Relevant mutations identified in the discovery screen (exome sequencing) (n = 17) (XLS 565 kb)
Supplementary Table 4
Pathway analyses of genes mutated in UBC (XLSX 38 kb)
Supplementary Table 6
Design of the prevalence screen and genes included therein, depth of coverage and relevant mutations detected (total and in each individual tumor) (XLSX 119 kb)
Rights and permissions
About this article
Cite this article
Balbás-Martínez, C., Sagrera, A., Carrillo-de-Santa-Pau, E. et al. Recurrent inactivation of STAG2 in bladder cancer is not associated with aneuploidy. Nat Genet 45, 1464–1469 (2013). https://doi.org/10.1038/ng.2799
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2799
This article is cited by
-
The synergism of SMC1A cohesin gene silencing and bevacizumab against colorectal cancer
Journal of Experimental & Clinical Cancer Research (2024)
-
Cohesin maintains replication timing to suppress DNA damage on cancer genes
Nature Genetics (2023)
-
Alterations of cohesin complex genes in acute myeloid leukemia: differential co-mutations, clinical presentation and impact on outcome
Blood Cancer Journal (2023)
-
The multifaceted roles of cohesin in cancer
Journal of Experimental & Clinical Cancer Research (2022)
-
Combinatorial gene targeting in primary human hematopoietic stem and progenitor cells
Scientific Reports (2022)