Abstract
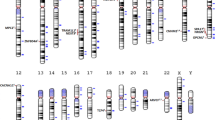
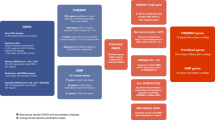
Schizophrenia is an idiopathic mental disorder with a heritable component and a substantial public health impact. We conducted a multi-stage genome-wide association study (GWAS) for schizophrenia beginning with a Swedish national sample (5,001 cases and 6,243 controls) followed by meta-analysis with previous schizophrenia GWAS (8,832 cases and 12,067 controls) and finally by replication of SNPs in 168 genomic regions in independent samples (7,413 cases, 19,762 controls and 581 parent-offspring trios). We identified 22 loci associated at genome-wide significance; 13 of these are new, and 1 was previously implicated in bipolar disorder. Examination of candidate genes at these loci suggests the involvement of neuronal calcium signaling. We estimate that 8,300 independent, mostly common SNPs (95% credible interval of 6,300–10,200 SNPs) contribute to risk for schizophrenia and that these collectively account for at least 32% of the variance in liability. Common genetic variation has an important role in the etiology of schizophrenia, and larger studies will allow more detailed understanding of this disorder.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Saha, S., Chant, D. & McGrath, J. A systematic review of mortality in schizophrenia: is the differential mortality gap worsening over time? Arch. Gen. Psychiatry 64, 1123–1131 (2007).
World Health Organization. The Global Burden of Disease: 2004 Update (World Health Organization Press, Geneva, 2008).
Knapp, M., Mangalore, R. & Simon, J. The global costs of schizophrenia. Schizophr. Bull. 30, 279–293 (2004).
Lichtenstein, P. et al. Recurrence risks for schizophrenia in a Swedish national cohort. Psychol. Med. 36, 1417–1425 (2006).
Lichtenstein, P. et al. Common genetic influences for schizophrenia and bipolar disorder: a population-based study of 2 million nuclear families. Lancet 373, 234–239 (2009).
Sullivan, P.F., Kendler, K.S. & Neale, M.C. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch. Gen. Psychiatry 60, 1187–1192 (2003).
Lee, S.H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Sullivan, P.F., Daly, M.J. & O'Donovan, M. Genetic architectures of psychiatric disorders: the emerging picture and its implications. Nat. Rev. Genet. 13, 537–551 (2012).
Ng, M.Y. et al. Meta-analysis of 32 genome-wide linkage studies of schizophrenia. Mol. Psychiatry 14, 774–785 (2009).
Girard, S.L. et al. Increased exonic de novo mutation rate in individuals with schizophrenia. Nat. Genet. 43, 860–863 (2011).
Xu, B. et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat. Genet. 44, 1365–1369 (2012).
Need, A.C. et al. Exome sequencing followed by large-scale genotyping suggests a limited role for moderately rare risk factors of strong effect in schizophrenia. Am. J. Hum. Genet. 91, 303–312 (2012).
Kim, Y., Zerwas, S., Trace, S.E. & Sullivan, P.F. Schizophrenia genetics: where next? Schizophr. Bull. 37, 456–463 (2011).
International Schizophrenia Consortium. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Shi, J. et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Wray, N.R. & Visscher, P.M. Narrowing the boundaries of the genetic architecture of schizophrenia. Schizophr. Bull. 36, 14–23 (2010).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Major Depressive Disorder Working Group of the PGC. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Hindorff, L.A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 106, 9362–9367 (2009).
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat. Genet. 43, 977–983 (2011).
Ferreira, M.A. et al. Collaborative genome-wide association analysis of 10,596 individuals supports a role for Ankyrin-G (ANK3) and the α-1C subunit of the L-type voltage-gated calcium channel (CACNA1C) in bipolar disorder. Nat. Genet. 40, 1056–1058 (2008).
Hamshere, M.L. et al. Genome-wide significant associations in schizophrenia to ITIH3/4, CACNA1C and SDCCAG8, and extensive replication of associations reported by the Schizophrenia PGC. Mol. Psychiatry 18, 708–712 (2013).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Cichon, S. et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am. J. Hum. Genet. 88, 372–381 (2011).
Bidaud, I., Mezghrani, A., Swayne, L.A., Monteil, A. & Lory, P. Voltage-gated calcium channels in genetic diseases. Biochim. Biophys. Acta 1763, 1169–1174 (2006).
Bichet, D. et al. The I-II loop of the Ca2+ channel α1 subunit contains an endoplasmic reticulum retention signal antagonized by the β subunit. Neuron 25, 177–190 (2000).
Müller, C.S. et al. Quantitative proteomics of the Cav2 channel nano-environments in the mammalian brain. Proc. Natl. Acad. Sci. USA 107, 14950–14957 (2010).
Woodside, B.L., Borroni, A.M., Hammonds, M.D. & Teyler, T.J. NMDA receptors and voltage-dependent calcium channels mediate different aspects of acquisition and retention of a spatial memory task. Neurobiol. Learn. Mem. 81, 105–114 (2004).
Moosmang, S. et al. Role of hippocampal Cav1.2 Ca2+ channels in NMDA receptor–independent synaptic plasticity and spatial memory. J. Neurosci. 25, 9883–9892 (2005).
White, J.A. et al. Conditional forebrain deletion of the L-type calcium channel Cav1.2 disrupts remote spatial memories in mice. Learn. Mem. 15, 1–5 (2008).
Mangoni, M.E. et al. Voltage-dependent calcium channels and cardiac pacemaker activity: from ionic currents to genes. Prog. Biophys. Mol. Biol. 90, 38–63 (2006).
Perrin, M.J. & Gollob, M.H. Genetics of cardiac electrical disease. Can. J. Cardiol. 29, 89–99 (2013).
Splawski, I. et al. Cav1.2 calcium channel dysfunction causes a multisystem disorder including arrhythmia and autism. Cell 119, 19–31 (2004).
Koponen, H. et al. Schizophrenia and sudden cardiac death: a review. Nord. J. Psychiatry 62, 342–345 (2008).
Stöllberger, C., Huber, J.O. & Finsterer, J. Antipsychotic drugs and QT prolongation. Int. Clin. Psychopharmacol. 20, 243–251 (2005).
McKusick, V.A. Mendelian Inheritance in Man and its online version, OMIM. Am. J. Hum. Genet. 80, 588–604 (2007).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Szulwach, K.E. et al. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell Biol. 189, 127–141 (2010).
Smrt, R.D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 28, 1060–1070 (2010).
Willemsen, M.H. et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J. Med. Genet. 48, 810–818 (2011).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Lee, P.H., O'Dushlaine, C., Thomas, B. & Purcell, S. InRich: interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Majewski, J. & Pastinen, T. The study of eQTL variations by RNA-seq: from SNPs to phenotypes. Trends Genet. 27, 72–79 (2011).
Cookson, W., Liang, L., Abecasis, G., Moffatt, M. & Lathrop, M. Mapping complex disease traits with global gene expression. Nat. Rev. Genet. 10, 184–194 (2009).
Nicolae, D.L. et al. Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet. 6, e1000888 (2010).
Stranger, B.E. et al. Patterns of cis regulatory variation in diverse human populations. PLoS Genet. 8, e1002639 (2012).
Grundberg, E. et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat. Genet. 44, 1084–1089 (2012).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
de Jong, S. et al. Expression QTL analysis of top loci from GWAS meta-analysis highlights additional schizophrenia candidate genes. Eur. J. Hum. Genet. 20, 1004–1008 (2012).
Fransen, K. et al. Analysis of SNPs with an effect on gene expression identifies UBE2L3 and BCL3 as potential new risk genes for Crohn's disease. Hum. Mol. Genet. 19, 3482–3488 (2010).
Luo, R. et al. Genome-wide transcriptome profiling reveals the functional impact of rare de novo and recurrent CNVs in autism spectrum disorders. Am. J. Hum. Genet. 91, 38–55 (2012).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Zeller, T. et al. Genetics and beyond—the transcriptome of human monocytes and disease susceptibility. PLoS ONE 5, e10693 (2010).
Gamazon, E.R., Huang, R.S., Cox, N.J. & Dolan, M.E. Chemotherapeutic drug susceptibility associated SNPs are enriched in expression quantitative trait loci. Proc. Natl. Acad. Sci. USA 107, 9287–9292 (2010).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Degner, J.F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Saha, S., Chant, D., Welham, J. & McGrath, J. A systematic review of the prevalence of schizophrenia. PLoS Med. 2, e141 (2005).
Stahl, E.A. et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nat. Genet. 44, 483–489 (2012).
Wray, N.R. & Gottesman, I.I. Using summary data from the Danish national registers to estimate heritabilities for schizophrenia, bipolar disorder, and major depressive disorder. Front. Genet. 3, 118 (2012).
Lander, E.S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Sullivan, P. Don't give up on GWAS. Mol. Psychiatry 17, 2–3 (2012).
Park, J.H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat. Genet. 42, 570–575 (2010).
Stahl, E.A. et al. Genome-wide association study meta-analysis identifies seven new rheumatoid arthritis risk loci. Nat. Genet. 42, 508–514 (2010).
Betancur, C. Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res. 1380, 42–77 (2011).
Chiurazzi, P., Schwartz, C.E., Gecz, J. & Neri, G. XLMR genes: update 2007. Eur. J. Hum. Genet. 16, 422–434 (2008).
Inlow, J.K. & Restifo, L.L. Molecular and comparative genetics of mental retardation. Genetics 166, 835–881 (2004).
Bergen, S.E. et al. Genome-wide association study in a Swedish population yields support for greater CNV and MHC involvement in schizophrenia compared to bipolar disorder. Mol. Psychiatry 17, 880–886 (2012).
Kristjansson, E., Allebeck, P. & Wistedt, B. Validity of the diagnosis of schizophrenia in a psychiatric inpatient register. Nord. Psykiatr. Tidsskr. 41, 229–234 (1987).
Dalman, C., Broms, J., Cullberg, J. & Allebeck, P. Young cases of schizophrenia identified in a national inpatient register—are the diagnoses valid? Soc. Psychiatry Psychiatr. Epidemiol. 37, 527–531 (2002).
World Health Organization. International Classification of Diseases (World Health Organization, Geneva, 1967).
World Health Organization. International Classification of Diseases (World Health Organization, Geneva, 1978).
World Health Organization. International Classification of Diseases (World Health Organization, Geneva, 1992).
Hultman, C.M., Sparen, P., Takei, N., Murray, R.M. & Cnattingius, S. Prenatal and perinatal risk factors for schizophrenia, affective psychosis, and reactive psychosis of early onset: case-control study. Br. Med. J. 318, 421–426 (1999).
Zammit, S. et al. Investigating the association between cigarette smoking and schizophrenia in a cohort study. Am. J. Psychiatry 160, 2216–2221 (2003).
Andersson, R.E., Olaison, G., Tysk, C. & Ekbom, A. Appendectomy and protection against ulcerative colitis. N. Engl. J. Med. 344, 808–814 (2001).
Hansson, L.E. et al. The risk of stomach cancer in patients with gastric or duodenal ulcer disease. N. Engl. J. Med. 335, 242–249 (1996).
Schwartz, S. & Susser, E. Genome-wide association studies: does only size matter? Am. J. Psychiatry 167, 741–744 (2010).
Craddock, N. & Owen, M.J. The Kraepelinian dichotomy—going, going but still not gone. Br. J. Psychiatry 196, 92–95 (2010).
Hartge, P. Participation in population studies. Epidemiology 17, 252–254 (2006).
Morton, L.M., Cahill, J. & Hartge, P. Reporting participation in epidemiologic studies: a survey of practice. Am. J. Epidemiol. 163, 197–203 (2006).
Bulik, C.M. et al. Patterns of remission, continuation and incidence of broadly defined eating disorders during early pregnancy in the Norwegian Mother and Child Cohort Study (MoBa). Psychol. Med. 37, 1109–1118 (2007).
Purcell, S. et al. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am. J. Hum. Genet. 81, 559–575 (2007).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 1, 457–470 (2011).
Delaneau, O., Marchini, J. & Zagury, J.F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2012).
McGrath, J.J. Variations in the incidence of schizophrenia: data versus dogma. Schizophr. Bull. 32, 195–197 (2006).
Lips, E.S. et al. Functional gene group analysis identifies synaptic gene groups as risk factor for schizophrenia. Mol. Psychiatry 17, 996–1006 (2012).
Irish Schizophrenia Genomics Consortium & Wellcome Trust Case Control Consortium. Genome-wide association study implicates HLA-C*01:02 as a risk factor at the MHC locus in schizophrenia. Biol. Psychiatry 72, 620–628 (2012).
Psychosis Endophenotypes International Consortium & Wellcome Trust Case-Control Consortium 2. A genome-wide association analysis of a broad psychosis phenotype identifies three loci for further investigation. Biol. Psychiatry doi:10.1016/j.biopsych.2013.03.033 (17 July 2013).10.1016/j.biopsych.2013.03.033
Levinson, D.F. et al. Genome-wide association study of multiplex schizophrenia pedigrees. Am. J. Psychiatry 169, 963–973 (2012).
Acknowledgements
We are deeply grateful for the participation of all subjects contributing to this research and to the collection team that worked to recruit them: E. Flordal-Thelander, A.-B. Holmgren, M. Hallin, M. Lundin, A.-K. Sundberg, C. Pettersson, R. Satgunanthan-Dawoud, S. Hassellund, M. Rådstrom, B. Ohlander, L. Nyrén and I. Kizling. Funding support was provided by the NIMH (R01 MH077139 to P.F.S. and R01 MH095034 to P.S.), the Stanley Center for Psychiatric Research, the Sylvan Herman Foundation, the Friedman Brain Institute at the Mount Sinai School of Medicine, the Karolinska Institutet, Karolinska University Hospital, the Swedish Research Council, the Swedish County Council, the Söderström Königska Foundation and the Netherlands Scientific Organization (NWO 645-000-003). SGENE was supported by European Union grant HEALTH-F2-2009-223423 (project PsychCNVs). The study of the Aarhus sample was supported by grants from the Danish Strategic Research Council, H. Lundbeck A/S, the Faculty of Health Sciences at Aarhus University, the Lundbeck Foundation and the Stanley Research Foundation. The Wellcome Trust Case Control Consortium 2 project collection was funded by the Wellcome Trust (085475/B/08/Z and 085475/Z/08/Z). The funders had no role in study design, execution or analysis or in manuscript preparation.
Author information
Authors and Affiliations
Consortia
Contributions
S.R., C.O., E.A.S., M.F., N.R.W., N.S., S.E.B., S.H.L., A.B.S., A.L.R., B.K.B.-S., B.M.N., C.d.L., D.P., D. Ruderfer, F.B., J.P., K.L., M.L.H., M.V., P.H., S.S., S.A.M., S.P. and P.F.S. conducted statistical analyses. A.D.B., D.M.H., D. Rujescu, E. Sigurdsson, J.S., M.P.M., N.D., O.M., P.B.M., S.T., T.S. and V.G. ascertained subjects. A.L.C., J.J.C., S.W., Y.K., K.X. and P.F.S. performed bioinformatics analyses. K.C., J.L.M. and S.A. managed the project. B.P.R., D.W.M., F.A.O., H.S., J.T.W., K.S.K., M.G., M.J.O., N.C., P.C., the Multicenter Genetic Studies of Schizophrenia, the Psychosis Endophenotypes International Consortium, the Wellcome Trust Case Control Consortium 2, A.P.C., E.B., K.S. and M.C.O. provided replication samples and genotypes. A.K.K. interfaced with Swedish national registers. The manuscript was written by P.K.E.M., S.A.M., S.P., P.S., C.M.H. and P.F.S. The study was designed by S.P., P.S., C.M.H. and P.F.S. Funding was obtained by E. Scolnick, P.S., C.M.H. and P.F.S.
Corresponding author
Ethics declarations
Competing interests
P.F.S. was on the scientific advisory board of Expression Analysis (Durham, North Carolina, USA). P.S. is on the Board of Directors of Catalytic, Inc. The other authors report no conflicts.
Supplementary information
Supplementary Figures and Text
Supplementary Figures 1–13 and Supplementary Table 1–9 and Supplementary Note (PDF 6339 kb)
Rights and permissions
About this article
Cite this article
Ripke, S., O'Dushlaine, C., Chambert, K. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 45, 1150–1159 (2013). https://doi.org/10.1038/ng.2742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2742
This article is cited by
-
Polygenic effects on brain functional endophenotype for deficit and non-deficit schizophrenia
Schizophrenia (2024)
-
Genomic analysis of firework fear and noise reactivity in standard poodles
Canine Medicine and Genetics (2023)
-
Methylation in MAD1L1 is associated with the severity of suicide attempt and phenotypes of depression
Clinical Epigenetics (2023)
-
Evaluating 17 methods incorporating biological function with GWAS summary statistics to accelerate discovery demonstrates a tradeoff between high sensitivity and high positive predictive value
Communications Biology (2023)
-
A higher dysregulation burden of brain DNA methylation in female patients implicated in the sex bias of Schizophrenia
Molecular Psychiatry (2023)