Abstract
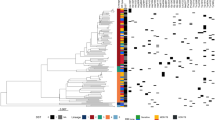
The worldwide emergence of multidrug-resistant (MDR) and extensively drug-resistant (XDR) tuberculosis threatens to make this disease incurable1,2. Drug resistance mechanisms are only partially understood3,4,5, and whether the current understanding of the genetic basis of drug resistance in M. tuberculosis is sufficiently comprehensive remains unclear. Here we sequenced and analyzed 161 isolates with a range of drug resistance profiles, discovering 72 new genes, 28 intergenic regions (IGRs), 11 nonsynonymous SNPs and 10 IGR SNPs with strong, consistent associations with drug resistance. On the basis of our examination of the dN/dS ratios of nonsynonymous to synonymous SNPs among the isolates6,7,8, we suggest that the drug resistance–associated genes identified here likely contain essentially all the nonsynonymous SNPs that have arisen as a result of drug pressure in these isolates and should thus represent a near-complete set of drug resistance–associated genes for these isolates and antibiotics. Our work indicates that the genetic basis of drug resistance is more complex than previously anticipated and provides a strong foundation for elucidating unknown drug resistance mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gandhi, N.R. et al. Multidrug-resistant and extensively drug-resistant tuberculosis: a threat to global control of tuberculosis. Lancet 375, 1830–1843 (2010).
Zumla, A. et al. Drug-resistant tuberculosis—current dilemmas, unanswered questions, challenges, and priority needs. J. Infect. Dis. 205 (suppl. 2), S228–S240 (2012).
Goldberg, D.E., Siliciano, R.F. & Jacobs, W.R. Jr. Outwitting evolution: fighting drug-resistant TB, malaria, and HIV. Cell 148, 1271–1283 (2012).
Laurenzo, D. & Mousa, S.A. Mechanisms of drug resistance in Mycobacterium tuberculosis and current status of rapid molecular diagnostic testing. Acta Trop. 119, 5–10 (2011).
Zhang, Y. & Yew, W.W. Mechanisms of drug resistance in Mycobacterium tuberculosis. Int. J. Tuberc. Lung Dis. 13, 1320–1330 (2009).
Elena, S.F. & Lenski, R.E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nat. Rev. Genet. 4, 457–469 (2003).
Barrick, J.E. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461, 1243–1247 (2009).
Woods, R., Schneider, D., Winkworth, C.L., Riley, M.A. & Lenski, R.E. Tests of parallel molecular evolution in a long-term experiment with Escherichia coli. Proc. Natl. Acad. Sci. USA 103, 9107–9112 (2006).
World Health Organization. Global Tuberculosis Control 2011 (World Health Organization, Geneva, 2011).
Zhao, Y. et al. National survey of drug-resistant tuberculosis in China. N. Engl. J. Med. 366, 2161–2170 (2012).
Ford, C.B. et al. Mycobacterium tuberculosis mutation rate estimates from different lineages predict substantial differences in the emergence of drug-resistant tuberculosis. Nat. Genet. 45, 784–790 (2013).
Casali, N. et al. Microevolution of extensively drug-resistant tuberculosis in Russia. Genome Res. 22, 735–745 (2012).
Walker, T.M. et al. Whole-genome sequencing to delineate Mycobacterium tuberculosis outbreaks: a retrospective observational study. Lancet Infect. Dis. 13, 137–146 (2013).
Ford, C.B. et al. Use of whole genome sequencing to estimate the mutation rate of Mycobacterium tuberculosis during latent infection. Nat. Genet. 43, 482–486 (2011).
Ioerger, T.R. et al. Genome analysis of multi- and extensively-drug-resistant tuberculosis from KwaZulu-Natal, South Africa. PLoS ONE 4, e7778 (2009).
Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Hershberg, R. et al. High functional diversity in Mycobacterium tuberculosis driven by genetic drift and human demography. PLoS Biol. 6, e311 (2008).
Gagneux, S. & Small, P.M. Global phylogeography of Mycobacterium tuberculosis and implications for tuberculosis product development. Lancet Infect. Dis. 7, 328–337 (2007).
Müller, B., Borrell, S., Rose, G. & Gagneux, S. The heterogeneous evolution of multidrug-resistant Mycobacterium tuberculosis. Trends Genet. 29, 160–169 (2013).
Ramaswamy, S. & Musser, J.M. Molecular genetic basis of antimicrobial agent resistance in Mycobacterium tuberculosis: 1998 update. Tuber. Lung Dis. 79, 3–29 (1998).
Sandgren, A. et al. Tuberculosis drug resistance mutation database. PLoS Med. 6, e2 (2009).
Sekiguchi, J. et al. Detection of multidrug resistance in Mycobacterium tuberculosis. J. Clin. Microbiol. 45, 179–192 (2007).
Zaunbrecher, M.A., Sikes, R.D. Jr., Metchock, B., Shinnick, T.M. & Posey, J.E. Overexpression of the chromosomally encoded aminoglycoside acetyltransferase eis confers kanamycin resistance in Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 106, 20004–20009 (2009).
World Health Organization. Guidelines for Surveillance of Drug Resistance in Tuberculosis (World Health Organization, Geneva, 2009).
World Health Organization. Treatment of Tuberculosis: Guidelines for National Programmes 4th edn. (World Health Organization, Geneva, 2009).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Mohanty, D., Sankaranarayanan, R. & Gokhale, R.S. Fatty acyl-AMP ligases and polyketide synthases are unique enzymes of lipid biosynthetic machinery in Mycobacterium tuberculosis. Tuberculosis (Edinb.) 91, 448–455 (2011).
Schroeder, E.K., de Souza, N., Santos, D.S., Blanchard, J.S. & Basso, L.A. Drugs that inhibit mycolic acid biosynthesis in Mycobacterium tuberculosis. Curr. Pharm. Biotechnol. 3, 197–225 (2002).
Heath, R.J., White, S.W. & Rock, C.O. Lipid biosynthesis as a target for antibacterial agents. Prog. Lipid Res. 40, 467–497 (2001).
Birch, H.L. et al. Biosynthesis of mycobacterial arabinogalactan: identification of a novel α(1→3) arabinofuranosyltransferase. Mol. Microbiol. 69, 1191–1206 (2008).
Gavalda, S. et al. The Pks13/FadD32 crosstalk for the biosynthesis of mycolic acids in Mycobacterium tuberculosis. J. Biol. Chem. 284, 19255–19264 (2009).
Domenech, P., Reed, M.B. & Barry, C.E. III. Contribution of the Mycobacterium tuberculosis MmpL protein family to virulence and drug resistance. Infect. Immun. 73, 3492–3501 (2005).
La Rosa, V. et al. MmpL3 is the cellular target of the antitubercular pyrrole derivative BM212. Antimicrob. Agents Chemother. 56, 324–331 (2012).
Tahlan, K. et al. SQ109 targets MmpL3, a membrane transporter of trehalose monomycolate involved in mycolic acid donation to the cell wall core of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 56, 1797–1809 (2012).
Deidda, D. et al. Bactericidal activities of the pyrrole derivative BM212 against multidrug-resistant and intramacrophagic Mycobacterium tuberculosis strains. Antimicrob. Agents Chemother. 42, 3035–3037 (1998).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nat. Genet. 44, 106–110 (2012).
Ramaswamy, S.V. et al. Single nucleotide polymorphisms in genes associated with isoniazid resistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 47, 1241–1250 (2003).
Rindi, L. et al. Mutations responsible for Mycobacterium tuberculosis isoniazid resistance in Italy. Int. J. Tuberc. Lung Dis. 9, 94–97 (2005).
Gagneux, S. et al. Impact of bacterial genetics on the transmission of isoniazid-resistant Mycobacterium tuberculosis. PLoS Pathog. 2, e61 (2006).
Fivian-Hughes, A.S., Houghton, J. & Davis, E.O. Mycobacterium tuberculosis thymidylate synthase gene thyX is essential and potentially bifunctional, while thyA deletion confers resistance to p-aminosalicylic acid. Microbiology 158, 308–318 (2012).
Reese, M.G. Application of a time-delay neural network to promoter annotation in the Drosophila melanogaster genome. Comput. Chem. 26, 51–56 (2001).
Nellen, W. & Hammann, C. Small RNAs: Analysis and Regulatory Functions (Springer, Heidelberg, Germany, 2005).
Song, T. & Wai, S.N. A novel sRNA that modulates virulence and environmental fitness of Vibrio cholerae. RNA Biol. 6, 254–258 (2009).
Arnvig, K.B. et al. Sequence-based analysis uncovers an abundance of non-coding RNA in the total transcriptome of Mycobacterium tuberculosis. PLoS Pathog. 7, e1002342 (2011).
Miotto, P. et al. Genome-wide discovery of small RNAs in Mycobacterium tuberculosis. PLoS ONE 7, e51950 (2012).
Griffin, J.E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7, e1002251 (2011).
Telenti, A. et al. Detection of rifampicin-resistance mutations in Mycobacterium tuberculosis. Lancet 341, 647–650 (1993).
Koenig, R. Few mutations divide some drug-resistant TB strains. Science 318, 901–902 (2007).
Fleischmann, R.D. et al. Whole-genome comparison of Mycobacterium tuberculosis clinical and laboratory strains. J. Bacteriol. 184, 5479–5490 (2002).
World Health Organization. Policy Guidance on TB Drug Susceptibility Testing (DST) of Second-Line Drugs (World Health Organization, Geneva, 2008).
Kamerbeek, J. et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J. Clin. Microbiol. 35, 907–914 (1997).
Demay, C. et al. SITVITWEB—a publicly available international multimarker database for studying Mycobacterium tuberculosis genetic diversity and molecular epidemiology. Infect. Genet. Evol. 12, 755–766 (2012).
van Soolingen, D., Hermans, P.W., de Haas, P.E., Soll, D.R. & van Embden, J.D. Occurrence and stability of insertion sequences in Mycobacterium tuberculosis complex strains: evaluation of an insertion sequence–dependent DNA polymorphism as a tool in the epidemiology of tuberculosis. J. Clin. Microbiol. 29, 2578–2586 (1991).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics Chapter 4, Unit 4.10 (2009).
Zhang, Z. et al. KaKs_Calculator: calculating Ka and Ks through model selection and model averaging. Genomics Proteomics Bioinformatics 4, 259–263 (2006).
Acknowledgements
We would like to thank S.M. Wang for helpful discussions and input and the anonymous reviewers for their constructive comments. We thank D. Chatterji (Indian Institute of Science) for the pSD5B vector. L.B. was supported by the National Natural Science Foundation of China (grant 31170132), the National Basic Research Program of China (grants 2009CB825402 and 2012CB518703), the Chinese Academy of Sciences (grant KSZD-EW-Z-006) and the Key Project Specialized for Infectious Diseases of the Chinese Ministry of Health (grants 2012ZX10003002 and 2013ZX10003006).
Author information
Authors and Affiliations
Contributions
H.Z. conducted the experiments. D.L., J.F., H.Z., N.L., Y.H., Z.L., Y. Zhu, S.W., M.W., Y. Zhou, J.D., J.W., R.Y., G.Z., N.G., H.H., L. Zhao, J. Zhou and J. Zhang contributed to data analysis. K.W., L. Zhao, C.L., Q.Z., L. Zhou, T.C. and F.L. collected the isolates. Y.G., H.Z., L. Zhao, N.L. and T.C. cultured the isolates and prepared DNA samples. Tong Wang performed genome sequencing and assembly. H.Z. and Ting Wang performed the IGR promoter experiments. J.F. and H.Z. wrote and revised the manuscript. X.-E.Z. directed part of the work, and L.B. conceived, designed and provided overall direction for the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–19 (PDF 21834 kb)
Rights and permissions
About this article
Cite this article
Zhang, H., Li, D., Zhao, L. et al. Genome sequencing of 161 Mycobacterium tuberculosis isolates from China identifies genes and intergenic regions associated with drug resistance. Nat Genet 45, 1255–1260 (2013). https://doi.org/10.1038/ng.2735
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2735
This article is cited by
-
Whole genome sequencing reveals candidate genes involving in PAS resistance in M. Tuberculosis isolated from patients in Thailand
World Journal of Microbiology and Biotechnology (2024)
-
Association between fatty acid metabolism gene mutations and Mycobacterium tuberculosis transmission revealed by whole genome sequencing
BMC Microbiology (2023)
-
Evolution and epidemic success of Mycobacterium tuberculosis in eastern China: evidence from a prospective study
BMC Genomics (2023)
-
Structural insights into RNase J that plays an essential role in Mycobacterium tuberculosis RNA metabolism
Nature Communications (2023)
-
Omics analysis of Mycobacterium tuberculosis isolates uncovers Rv3094c, an ethionamide metabolism-associated gene
Communications Biology (2023)