Abstract
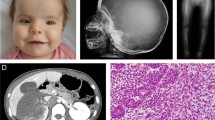
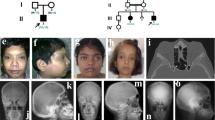
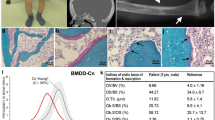
Abnormalities in WNT signaling are implicated in a broad range of developmental anomalies and also in tumorigenesis1. Here we demonstrate that germline mutations in WTX (FAM123B), a gene that encodes a repressor of canonical WNT signaling2, cause an X-linked sclerosing bone dysplasia, osteopathia striata congenita with cranial sclerosis (OSCS; MIM300373)3. This condition is typically characterized by increased bone density and craniofacial malformations in females and lethality in males. The mouse homolog of WTX is expressed in the fetal skeleton, and alternative splicing implicates plasma membrane localization of WTX as a factor associated with survival in males with OSCS. WTX has also been shown to be somatically inactivated in 11–29% of cases of Wilms tumor4,5,6. Despite being germline for such mutations, individuals with OSCS are not predisposed to tumor development. The observed phenotypic discordance dependent upon whether a mutation is germline or occurs somatically suggests the existence of temporal or spatial constraints on the action of WTX during tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Clevers, H. Wnt/β-catenin signaling in development and disease. Cell 127, 469–480 (2006).
Major, M.B. et al. Wilms tumor suppressor WTX negatively regulates WNT/β-catenin signaling. Science 316, 1043–1046 (2007).
Hurt, R.L. Osteopathia striata-Voorhoeve's disease; report of a case presenting the features of osteopathia striata and osteopetrosis. J. Bone Joint Surg. Br. 35-B, 89–96 (1953).
Rivera, M.N. et al. An X chromosome gene, WTX, is commonly inactivated in Wilms tumor. Science 315, 642–645 (2007).
Ruteshouser, E.C., Robinson, S.M. & Huff, V. Wilms tumor genetics: mutations in WT1, WTX, and CTNNB1 account for only about one-third of tumors. Genes Chromosom. Cancer 47, 461–470 (2008).
Perotti, D. et al. Functional inactivation of the WTX gene is not a frequent event in Wilms' tumors. Oncogene 27, 4625–4632 (2008).
Behninger, C. & Rott, H.D. Osteopathia striata with cranial sclerosis: literature reappraisal argues for X-linked inheritance. Genet. Couns. 11, 157–167 (2000).
Nannya, Y. et al. A robust algorithm for copy number detection using high-density oligonucleotide single nucleotide polymorphism genotyping arrays. Cancer Res. 65, 6071–6079 (2005).
Bennett, C.N. et al. Wnt10b increases postnatal bone formation by enhancing osteoblast differentiation. J. Bone Miner. Res. 22, 1924–1932 (2007).
Morvan, F. et al. Deletion of a single allele of the Dkk1 gene leads to an increase in bone formation and bone mass. J. Bone Miner. Res. 21, 934–945 (2006).
Li, X. et al. Sclerostin binds to LRP5/6 and antagonizes canonical Wnt signaling. J. Biol. Chem. 280, 19883–19887 (2005).
Little, R.D. et al. A mutation in the LDL receptor-related protein 5 gene results in the autosomal dominant high-bone-mass trait. Am. J. Hum. Genet. 70, 11–19 (2002).
Boyden, L.M. et al. High bone density due to a mutation in LDL-receptor-related protein 5. N. Engl. J. Med. 346, 1513–1521 (2002).
Clement-Lacroix, P. et al. Lrp5-independent activation of Wnt signaling by lithium chloride increases bone formation and bone mass in mice. Proc. Natl. Acad. Sci. USA 102, 17406–17411 (2005).
Grzeschik, K.H. et al. Deficiency of PORCN, a regulator of Wnt signaling, is associated with focal dermal hypoplasia. Nat. Genet. 39, 833–835 (2007).
Wang, X. et al. Mutations in X-linked PORCN, a putative regulator of Wnt signaling, cause focal dermal hypoplasia. Nat. Genet. 39, 836–838 (2007).
Grohmann, A., Tanneberger, K., Alzner, A., Schneikert, J. & Behrens, J. AMER1 regulates the distribution of the tumor suppressor APC between microtubules and the plasma membrane. J. Cell Sci. 120, 3738–3747 (2007).
Gaur, T. et al. Canonical WNT signaling promotes osteogenesis by directly stimulating Runx2 gene expression. J. Biol. Chem. 280, 33132–33140 (2005).
Babij, P. et al. High bone mass in mice expressing a mutant LRP5 gene. J. Bone Miner. Res. 18, 960–974 (2003).
Bodine, P.V. et al. The Wnt antagonist secreted frizzled-related protein-1 is a negative regulator of trabecular bone formation in adult mice. Mol. Endocrinol. 18, 1222–1237 (2004).
Holmen, S.L. et al. Essential role of β-catenin in postnatal bone acquisition. J. Biol. Chem. 280, 21162–21168 (2005).
Glass, D.A. II et al. Canonical Wnt signaling in differentiated osteoblasts controls osteoclast differentiation. Dev. Cell 8, 751–764 (2005).
Hill, T.P., Taketo, M.M., Birchmeier, W. & Hartmann, C. Multiple roles of mesenchymal β-catenin during murine limb patterning. Development 133, 1219–1229 (2006).
Zhou, H., Mak, W., Zheng, Y., Dunstan, C.R. & Seibel, M.J. Osteoblasts directly control lineage commitment of mesenchymal progenitor cells through Wnt signaling. J. Biol. Chem. 283, 1936–1945 (2008).
Day, T.F., Guo, X., Garrett-Beal, L. & Yang, Y. Wnt/β-catenin signaling in mesenchymal progenitors controls osteoblast and chondrocyte differentiation during vertebrate skeletogenesis. Dev. Cell 8, 739–750 (2005).
Rodda, S.J. & McMahon, A.P. Distinct roles for Hedgehog and canonical Wnt signaling in specification, differentiation and maintenance of osteoblast progenitors. Development 133, 3231–3244 (2006).
Weinberg, R.A. Tumor suppressor genes. Science 254, 1138–1146 (1991).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-ΔΔC(T)) Method. Methods 25, 402–408 (2001).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 8, R19 (2007).
Acknowledgements
The investigators are grateful for the generosity of the individuals and families with OSCS for their participation in this work. Thanks to A. Reeve, J. Matheson and P. Daniel for comments on the manuscript, A. Wilkie and C. Beck for discussions and R. McPhee for assistance with manuscript preparation. S.P.R. is supported by the Child Health Research Foundation of New Zealand and Lotteries Health New Zealand.
Author information
Authors and Affiliations
Contributions
Z.A.J. conceived, designed and performed the experiments addressing WTX functions and co-wrote the paper; M.v.K. analysed the microarray data, conceived and performed experiments validating genomic deletions and co-wrote the manuscript; T.M. performed and validated mutation detection; A.J. and R.F. assisted in qPCR and MLPA design and execution; E.P. and C.T. designed and performed the in situ hybridization analysis; A.V.H., M.E.P., S.G.-M., A.B., D.L., F.S., T.F., L.B., S.B., L.C.A., M.T., A.D., L.C.W., R.C.M.H., D.D., S.M. and V.C.-D. ascertained affected individuals, and evaluated and characterized their phenotypes; and S.P.R. led the project, conceived and designed experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The corresponding author has filed a provisional patent for the application of insights gained from this study. The patent refers to exploiting the properties of WTX with the aim of improving bone quality and structure.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 956 kb)
Rights and permissions
About this article
Cite this article
Jenkins, Z., van Kogelenberg, M., Morgan, T. et al. Germline mutations in WTX cause a sclerosing skeletal dysplasia but do not predispose to tumorigenesis. Nat Genet 41, 95–100 (2009). https://doi.org/10.1038/ng.270
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.270
This article is cited by
-
Hallmark discoveries in the biology of Wilms tumour
Nature Reviews Urology (2024)
-
Intestinal malrotation in a female newborn affected by Osteopathia Striata with Cranial Sclerosis due to a de novo heterozygous nonsense mutation of the AMER1 gene
Italian Journal of Pediatrics (2022)
-
A primer on skeletal dysplasias
Japanese Journal of Radiology (2022)
-
Wilms tumor in patients with osteopathia striata with cranial sclerosis
European Journal of Human Genetics (2021)
-
Human Genetics of Sclerosing Bone Disorders
Current Osteoporosis Reports (2018)