Abstract
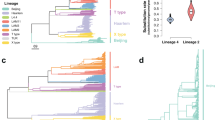
A key question in tuberculosis control is why some strains of M. tuberculosis are preferentially associated with resistance to multiple drugs. We demonstrate that M. tuberculosis strains from lineage 2 (East Asian lineage and Beijing sublineage) acquire drug resistances in vitro more rapidly than M. tuberculosis strains from lineage 4 (Euro-American lineage) and that this higher rate can be attributed to a higher mutation rate. Moreover, the in vitro mutation rate correlates well with the bacterial mutation rate in humans as determined by whole-genome sequencing of clinical isolates. Finally, using a stochastic mathematical model, we demonstrate that the observed differences in mutation rate predict a substantially higher probability that patients infected with a drug-susceptible lineage 2 strain will harbor multidrug-resistant bacteria at the time of diagnosis. These data suggest that interventions to prevent the emergence of drug-resistant tuberculosis should target bacterial as well as treatment-related risk factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Velayati, A.A. et al. Emergence of new forms of totally drug-resistant tuberculosis bacilli: super extensively drug-resistant tuberculosis or totally drug-resistant strains in Iran. Chest 136, 420–425 (2009).
Udwadia, Z.F., Amale, R.A., Ajbani, K.K. & Rodrigues, C. Totally drug-resistant tuberculosis in India. Clin. Infect. Dis. 54, 579–581 (2012).
Migliori, G.B., De Iaco, G., Besozzi, G., Centis, R. & Cirillo, D.M. First tuberculosis cases in Italy resistant to all tested drugs. Euro Surveill. 12, E070517.1 (2007).
Gandhi, N.R. et al. Multidrug-resistant and extensively drug-resistant tuberculosis: a threat to global control of tuberculosis. Lancet 375, 1830–1843 (2010).
David, H.L. Probability distribution of drug-resistant mutants in unselected populations of Mycobacterium tuberculosis. Appl. Microbiol. 20, 810–814 (1970).
Ford, C.B. et al. Use of whole genome sequencing to estimate the mutation rate of Mycobacterium tuberculosis during latent infection. Nat. Genet. 10.1038/ng.811 (2011).10.1038/ng.811
Bradford, W.Z. et al. The changing epidemiology of acquired drug-resistant tuberculosis in San Francisco, USA. Lancet 348, 928–931 (1996).
Goble, M. et al. Treatment of 171 patients with pulmonary tuberculosis resistant to isoniazid and rifampin. N. Engl. J. Med. 328, 527–532 (1993).
Pablos-Méndez, A., Knirsch, C.A., Barr, R.G., Lerner, B.H. & Frieden, T.R. Nonadherence in tuberculosis treatment: predictors and consequences in New York City. Am. J. Med. 102, 164–170 (1997).
Udwadia, Z.F., Pinto, L.M. & Uplekar, M.W. Tuberculosis management by private practitioners in Mumbai, India: has anything changed in two decades? PLoS ONE 5, e12023 (2010).
Johnson, R. et al. Drug-resistant tuberculosis epidemic in the Western Cape driven by a virulent Beijing genotype strain. Int. J. Tuberc. Lung Dis. 14, 119–121 (2010).
Streicher, E.M. et al. Genotypic and phenotypic characterization of drug-resistant Mycobacterium tuberculosis isolates from rural districts of the Western Cape Province of South Africa. J. Clin. Microbiol. 42, 891–894 (2004).
Casali, N. et al. Microevolution of extensively drug-resistant tuberculosis in Russia. Genome Res. 22, 735–745 (2012).
Ioerger, T.R. et al. The non-clonality of drug resistance in Beijing-genotype isolates of Mycobacterium tuberculosis from the Western Cape of South Africa. BMC Genomics 11, 670 (2010).
Sun, G. et al. Dynamic population changes in Mycobacterium tuberculosis during acquisition and fixation of drug resistance in patients. J. Infect. Dis. 10.1093/infdis/jis601 (2012).10.1093/infdis/jis601
European Concerted Action on New Generation Genetic Markers and Techniques for the Epidemiology and Control of Tuberculosis. Beijing/W genotype Mycobacterium tuberculosis and drug resistance. Emerg. Infect. Dis. 12, 736–743 (2006).
Borrell, S. & Gagneux, S. Infectiousness, reproductive fitness and evolution of drug-resistant Mycobacterium tuberculosis. Int. J. Tuberc. Lung Dis. 13, 1456–1466 (2009).
Hershberg, R. et al. High functional diversity in Mycobacterium tuberculosis driven by genetic drift and human demography. PLoS Biol. 6, e311 (2008).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Glynn, J.R., Whiteley, J., Bifani, P.J., Kremer, K. & Van Soolingen, D. Worldwide occurrence of Beijing/W strains of Mycobacterium tuberculosis: a systematic review. Emerg. Infect. Dis. 8, 843–849 (2002).
Kato-Maeda, M. et al. Beijing sublineages of Mycobacterium tuberculosis differ in pathogenicity in the guinea pig. Clin. Vaccine Immunol. 19, 1227–1237 (2012).
Drobniewski, F. et al. Drug-resistant tuberculosis, clinical virulence, and the dominance of the Beijing strain family in Russia. J. Am. Med. Assoc. 293, 2726–2731 (2005).
Coscolla, M. & Gagneux, S. Does M. tuberculosis genomic diversity explain disease diversity? Drug Discov. Today Dis. Mech. 7, e43–e59 (2010).
de Jong, B.C. et al. Progression to active tuberculosis, but not transmission, varies by Mycobacterium tuberculosis lineage in The Gambia. J. Infect. Dis. 198, 1037–1043 (2008).
Pang, Y. et al. Spoligotyping and drug resistance analysis of Mycobacterium tuberculosis strains from national survey in China. PLoS ONE 7, e32976 (2012).
Vadwai, V., Shetty, A., Supply, P. & Rodrigues, C. Evaluation of 24-locus MIRU-VNTR in extrapulmonary specimens: study from a tertiary centre in Mumbai. Tuberculosis (Edinb.) 92, 264–272 (2012).
Huang, H.Y. et al. Mixed infection with Beijing and non-Beijing strains and drug resistance pattern of Mycobacterium tuberculosis. J. Clin. Microbiol. 48, 4474–4480 (2010).
Anh, D.D. et al. Mycobacterium tuberculosis Beijing genotype emerging in Vietnam. Emerg. Infect. Dis. 6, 302–305 (2000).
Taype, C.A. et al. Genetic diversity, population structure and drug resistance of Mycobacterium tuberculosis in Peru. Infect. Genet. Evol. 12, 577–585 (2012).
Mestre, O. et al. Phylogeny of Mycobacterium tuberculosis Beijing strains constructed from polymorphisms in genes involved in DNA replication, recombination and repair. PLoS ONE 6, e16020 (2011).
de Steenwinkel, J.E.M. et al. Drug susceptibility of Mycobacterium tuberculosis Beijing genotype and association with MDR TB. Emerg. Infect. Dis. 18, 660–663 (2012).
Werngren, J. Drug-susceptible Mycobacterium tuberculosis Beijing genotype does not develop mutation-conferred resistance to rifampin at an elevated rate. J. Clin. Microbiol. 41, 1520–1524 (2003).
Rosche, W.A. & Foster, P.L. Determining mutation rates in bacterial populations. Methods 20, 4–17 (2000).
Luria, S.E. & Delbrück, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Lang, G.I. & Murray, A.W. Estimating the per-base-pair mutation rate in the yeast Saccharomyces cerevisiae. Genetics 178, 67–82 (2008).
Stewart, F.M. Fluctuation tests: how reliable are the estimates of mutation rates? Genetics 137, 1139–1146 (1994).
Stewart, F.M., Gordon, D.M. & Levin, B.R. Fluctuation analysis: the probability distribution of the number of mutants under different conditions. Genetics 124, 175–185 (1990).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 19, 716–723 (1974).
Burnham, K.P. Multimodel inference: understanding AIC and BIC in model selection. Sociol. Methods Res. 33, 261–304 (2004).
Bergval, I.L., Schuitema, A.R.J., Klatser, P.R. & Anthony, R.M. Resistant mutants of Mycobacterium tuberculosis selected in vitro do not reflect the in vivo mechanism of isoniazid resistance. J. Antimicrob. Chemother. 64, 515–523 (2009).
Sandgren, A. et al. Tuberculosis drug resistance mutation database. PLoS Med. 6, e2 (2009).
Gardy, J.L. et al. Whole-genome sequencing and social-network analysis of a tuberculosis outbreak. N. Engl. J. Med. 364, 730–739 (2011).
Drummond, A.J., Suchard, M.A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Drummond, A.J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Kimura, M. & Ota, T. On the rate of molecular evolution. J. Mol. Evol. 1, 1–17 (1971).
Walker, T.M. et al. Whole-genome sequencing to delineate Mycobacterium tuberculosis outbreaks: a retrospective observational study. Lancet Infect. Dis. 13, 137–146 (2013).
Colijn, C., Cohen, T., Ganesh, A. & Murray, M.B. Spontaneous emergence of multiple drug resistance in tuberculosis before and during therapy. PLoS ONE 6, e18327 (2011).
Portevin, D., Gagneux, S., Comas, I. & Young, D.B. Human macrophage responses to clinical isolates from the Mycobacterium tuberculosis complex discriminate between ancient and modern lineages. PLoS Pathog. 7, e1001307 (2011).
Bromham, L. & Penny, D. The modern molecular clock. Nat. Rev. Genet. 4, 216–224 (2003).
Rocha, E.P.C. et al. Comparisons of dN/dS are time dependent for closely related bacterial genomes. J. Theor. Biol. 239, 226–235 (2006).
Hall, L.M.C. & Henderson-Begg, S.K. Hypermutable bacteria isolated from humans—a critical analysis. Microbiology 152, 2505–2514 (2006).
Blázquez, J. Hypermutation as a factor contributing to the acquisition of antimicrobial resistance. Clin. Infect. Dis. 37, 1201–1209 (2003).
Oliver, A., Cantón, R., Campo, P., Baquero, F. & Blázquez, J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science 288, 1251–1254 (2000).
Mizrahi, V. & Andersen, S.J. DNA repair in Mycobacterium tuberculosis. What have we learnt from the genome sequence? Mol. Microbiol. 29, 1331–1339 (1998).
Springer, B. et al. Lack of mismatch correction facilitates genome evolution in mycobacteria. Mol. Microbiol. 53, 1601–1609 (2004).
Fallow, A., Domenech, P. & Reed, M.B. Strains of the East Asian (W/Beijing) lineage of Mycobacterium tuberculosis are DosS/DosT-DosR two-component regulatory system natural mutants. J. Bacteriol. 192, 2228–2238 (2010).
Huet, G. et al. A lipid profile typifies the Beijing strains of Mycobacterium tuberculosis: identification of a mutation responsible for a modification of the structures of phthiocerol dimycocerosates and phenolic glycolipids. J. Biol. Chem. 284, 27101–27113 (2009).
Schmalstieg, A.M. et al. The antibiotic resistance arrow of time: efflux pump induction is a general first step in the evolution of mycobacterial drug resistance. Antimicrob. Agents Chemother. 56, 4806–4815 (2012).
Louw, G.E. et al. Rifampicin reduces susceptibility to ofloxacin in rifampicin resistant Mycobacterium tuberculosis through efflux. Am. J. Respir. Crit. Care Med. 184, 269–276 (2011).
Adams, K.N. et al. Drug tolerance in replicating mycobacteria mediated by a macrophage-induced efflux mechanism. Cell 145, 39–53 (2011).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nat. Genet. 44, 106–110 (2012).
Williams, K. et al. Sterilizing activities of novel combinations lacking first- and second-line drugs in a murine model of tuberculosis. Antimicrob. Agents Chemother. 56, 3114–3120 (2012).
Lienhardt, C. et al. New drugs for the treatment of tuberculosis: needs, challenges, promise, and prospects for the future. J. Infect. Dis. 205, S241–S249 (2012).
Menzies, D. et al. Standardized treatment of active tuberculosis in patients with previous treatment and/or with mono-resistance to isoniazid: a systematic review and meta-analysis. PLoS Med. 6, e1000150 (2009).
Gelmanova, I.Y. et al. Barriers to successful tuberculosis treatment in Tomsk, Russian Federation: non-adherence, default and the acquisition of multidrug resistance. Bull. World Health Organ. 85, 703–711 (2007).
Seung, K.J. et al. The effect of initial drug resistance on treatment response and acquired drug resistance during standardized short-course chemotherapy for tuberculosis. Clin. Infect. Dis. 39, 1321–1328 (2004).
Helb, D. et al. Rapid detection of Mycobacterium tuberculosis and rifampin resistance by use of on-demand, near-patient technology. J. Clin. Microbiol. 48, 229–237 (2010).
Weyer, K. Laboratory Services in Tuberculosis Control. Part II: Microscopy (World Health Organization, Geneva, 1998).
Dowdy, D.W., Basu, S. & Andrews, J.R. Is passive diagnosis enough? The impact of subclinical disease on diagnostic strategies for tuberculosis. Am. J. Respir. Crit. Care Med. 187, 543–551 (2013).
Shaw, J.B. & Wynn-Williams, N. Infectivity of pulmonary tuberculosis in relation to sputum status. Am. Rev. Tuberc. 69, 724–732 (1954).
Tsolaki, A.G. et al. Functional and evolutionary genomics of Mycobacterium tuberculosis: insights from genomic deletions in 100 strains. Proc. Natl. Acad. Sci. USA 101, 4865–4870 (2004).
Gagneux, S. The competitive cost of antibiotic resistance in Mycobacterium tuberculosis. Science 312, 1944–1946 (2006).
Acknowledgements
This work was supported by a New Innovator's Award (DP2 0D001378) from the Director's Office of the US National Institutes of Health (S.M.F.), by a subcontract from the National Institute of Allergy and Infectious Diseases (U19 AI076217 to S.M.F. and M.B.M.), by the US National Institutes of Health Models of Infectious Disease Agent Study program, through cooperative agreement 1 U54 GM088558 (M.L.), by a Howard Hughes Medical Institute Physician Scientist Early Career Award (S.M.F.), by a Merit Fellowship from the Harvard University Graduate School of Arts and Sciences (C.B.F.) and by a Doris Duke Charitable Foundation Clinical Scientist Development Award (S.M.F.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences, National Institute of Allergy and Infectious Diseases or the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
C.B.F. designed the study, performed experimental and molecular studies, developed the mathematical model and conducted analyses, prepared the figures and drafted the manuscript. R.R.S. performed experimental studies. M.K.M. and S.G. isolated clinical strains. M.B.M. and T.C. advised development of the mathematical model. J.C.J. and J.G. contributed to analyses. M.L. advised design of the study and data analysis, including development of the mathematical model. S.M.F. designed the study, supervised experimental and molecular studies, and drafted the manuscript. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Tables 1–6 (PDF 489 kb)
Rights and permissions
About this article
Cite this article
Ford, C., Shah, R., Maeda, M. et al. Mycobacterium tuberculosis mutation rate estimates from different lineages predict substantial differences in the emergence of drug-resistant tuberculosis. Nat Genet 45, 784–790 (2013). https://doi.org/10.1038/ng.2656
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2656
This article is cited by
-
Molecular epidemiology and transmission dynamics of multi-drug resistant tuberculosis strains using whole genome sequencing in the Amhara region, Ethiopia
BMC Genomics (2023)
-
Mixed infections in genotypic drug-resistant Mycobacterium tuberculosis
Scientific Reports (2023)
-
The evolution of antibiotic resistance is associated with collateral drug phenotypes in Mycobacterium tuberculosis
Nature Communications (2023)
-
The relative transmission fitness of multidrug-resistant Mycobacterium tuberculosis in a drug resistance hotspot
Nature Communications (2023)
-
CT and 18F-FDG PET abnormalities in contacts with recent tuberculosis infections but negative chest X-ray
Insights into Imaging (2022)