Abstract
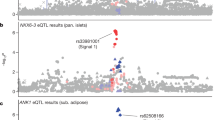
Genome-wide association studies (GWAS) have identified 36 loci associated with body mass index (BMI), predominantly in populations of European ancestry. We conducted a meta-analysis to examine the association of >3.2 million SNPs with BMI in 39,144 men and women of African ancestry and followed up the most significant associations in an additional 32,268 individuals of African ancestry. We identified one new locus at 5q33 (GALNT10, rs7708584, P = 3.4 × 10−11) and another at 7p15 when we included data from the GIANT consortium (MIR148A-NFE2L3, rs10261878, P = 1.2 × 10−10). We also found suggestive evidence of an association at a third locus at 6q16 in the African-ancestry sample (KLHL32, rs974417, P = 6.9 × 10−8). Thirty-two of the 36 previously established BMI variants showed directionally consistent effect estimates in our GWAS (binomial P = 9.7 × 10−7), five of which reached genome-wide significance. These findings provide strong support for shared BMI loci across populations, as well as for the utility of studying ancestrally diverse populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Flegal, K.M., Carroll, M.D., Kit, B.K. & Ogden, C.L. Prevalence of obesity and trends in the distribution of body mass index among US adults, 1999–2010. J. Am. Med. Assoc. 307, 491–497 (2012).
Bradfield, J.P. et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat. Genet. 44, 526–531 (2012).
Thorleifsson, G. et al. Genome-wide association yields new sequence variants at seven loci that associate with measures of obesity. Nat. Genet. 41, 18–24 (2009).
Frayling, T.M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Scuteri, A. et al. Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet. 3, e115 (2007).
Loos, R.J.F. et al. Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat. Genet. 40, 768–775 (2008).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Willer, C.J. et al. Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat. Genet. 41, 25–34 (2009).
Okada, Y. et al. Common variants at CDKAL1 and KLF9 are associated with body mass index in east Asian populations. Nat. Genet. 44, 302–306 (2012).
Wen, W. et al. Meta-analysis identifies common variants associated with body mass index in east Asians. Nat. Genet. 44, 307–311 (2012).
Chambers, J.C. et al. Common genetic variation near MC4R is associated with waist circumference and insulin resistance. Nat. Genet. 40, 716–718 (2008).
Meyre, D. et al. Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nat. Genet. 41, 157–159 (2009).
Scherag, A. et al. Two new loci for body-weight regulation identified in a joint analysis of genome-wide association studies for early-onset extreme obesity in French and german study groups. PLoS Genet. 6, e1000916 (2010).
Ng, M.C. et al. Genome-wide association of BMI in African Americans. Obesity (Silver Spring) 20, 622–627 (2012).
Kang, S.J. et al. Genome-wide association of anthropometric traits in African- and African-derived populations. Hum. Mol. Genet. 19, 2725–2738 (2010).
Shifman, S., Kuypers, J., Kokoris, M., Yakir, B. & Darvasi, A. Linkage disequilibrium patterns of the human genome across populations. Hum. Mol. Genet. 12, 771–776 (2003).
Campbell, M.C. & Tishkoff, S.A. African genetic diversity: implications for human demographic history, modern human origins, and complex disease mapping. Annu. Rev. Genomics Hum. Genet. 9, 403–433 (2008).
N'Diaye, A. et al. Identification, replication, and fine-mapping of loci associated with adult height in individuals of african ancestry. PLoS Genet. 7, e1002298 (2011).
Heid, I.M. et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat. Genet. 42, 949–960 (2010).
Teslovich, T.M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Saxena, R. et al. Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nat. Genet. 42, 142–148 (2010).
Voight, B.F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Ehret, G.B. et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 478, 103–109 (2011).
Nalls, M.A. et al. Imputation of sequence variants for identification of genetic risks for Parkinson's disease: a meta-analysis of genome-wide association studies. Lancet 377, 641–649 (2011).
Hernandez, D.G. et al. Distinct DNA methylation changes highly correlated with chronological age in the human brain. Hum. Mol. Genet. 20, 1164–1172 (2011).
Zhong, H., Yang, X., Kaplan, L.M., Molony, C. & Schadt, E.E. Integrating pathway analysis and genetics of gene expression for genome-wide association studies. Am. J. Hum. Genet. 86, 581–591 (2010).
Myers, A.J. et al. A survey of genetic human cortical gene expression. Nat. Genet. 39, 1494–1499 (2007).
Cheng, L. et al. Characterization of a novel human UDP-GalNAc transferase, pp-GalNAc-T10. FEBS Lett. 531, 115–121 (2002).
Xie, H., Lim, B. & Lodish, H.F. MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes 58, 1050–1057 (2009).
Ortega, F.J. et al. MiRNA expression profile of human subcutaneous adipose and during adipocyte differentiation. PLoS ONE 5, e9022 (2010).
Clerc, P. et al. Involvement of cholecystokinin 2 receptor in food intake regulation: hyperphagia and increased fat deposition in cholecystokinin 2 receptor-deficient mice. Endocrinology 148, 1039–1049 (2007).
Chevillard, G. & Blank, V. NFE2L3 (NRF3): the Cinderella of the Cap'n'Collar transcription factors. Cell Mol. Life Sci. 68, 3337–3348 (2011).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Price, A.L. et al. Sensitive detection of chromosomal segments of distinct ancestry in admixed populations. PLoS Genet. 5, e1000519 (2009).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Coetzee, S.G., Rhie, S.K., Berman, B.P., Coetzee, G.A. & Noushmehr, H. FunciSNP: an R/bioconductor tool integrating functional non-coding data sets with genetic association studies to identify candidate regulatory SNPs. Nucleic Acids Res. 40, e139 (2012).
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Bhatia, G. et al. Genome-wide comparison of African-ancestry populations from CARe and other cohorts reveals signals of natural selection. Am. J. Hum. Genet. 89, 368–381 (2011).
Murray, T. et al. African and non-African admixture components in African Americans and an African Caribbean population. Genet. Epidemiol. 34, 561–568 (2010).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Pickrell, J.K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Sabeti, P.C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Falush, D., Stephens, M. & Pritchard, J.K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Acknowledgements
A full listing of acknowledgments is detailed in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Design and/or management of the individual studies: A.A., A.A.A., C.B.A., C.I.A., D.K.A., L.A., M.C.A., M.A.A., S.A., C.H.B., D.M.B., D.W.B., E.P.B., E.V.B., G.B., I.B.B., J.P.B., L.B., S.I.B., W.J.B., C.S.C., G. Casey, G.K.C., J.C., L.C., M.C., N.E.C., Q.C., R.S.C., S.J.C., J.D., P.D., R.W.D., S.L.D.-H., M.K.E., T.L.E., C.F., J.K.F., E.M.G., P.J.G., S.F.A.G., A.H., A.J.M.H., B.E.H., B.V.H., C.A.H., C.C.H., D.H., H.H., K.J.H., J.J.H., J.N.H., V.J.H., S.A.I., E.M.J., J.M.J., C.K., D.L.K., E.A.K., E.K.K., L.N.K., L.H.K., R.A.K., S.J.K., S. Kolb, L.L.M., A.M.L., R.J.F.L., S.L., Yongmei Liu, A.B.M., B.M., K.R.M., R.C.M., T.H.M., J.H.M., J.C.M., I.M.-B., K.E.N., M.C.Y.N., S.N., C.N.-D., U.N., B.N., K.L.N., T.O.O., O.O., O.I.O., B.P., U.P., B.M.P., C.A.P., G.J.P., J.R.P., M.F.P., P.A.P., S.R.P., E.A.R.-N., B.A.R., C.N.R., S.R., J.L.R.-G., A.B.S., A.G.S., J.L.S., L.B.S., P.J.S., S.B.S., S.W.-S., M.R.S., M.M.S., I.J.S., S.S.S., H.T., M.J.T., M.A.T., M.V., J.S.W., X.W., J.K.W., S.M.W., L.K.W., M.W., J.J.Y., N.A.Z., R.G.Z., W. Zheng, A.B.Z., K.A.Z., Y.Z. and X.Z.
Genotyping: A.B., U.B., S.J.C., Y.-D.I.C., D.D., T.L.E., S.F.A.G., X.G., D.G.H., J.N.H., T.D.H., T. Haritunians, K.C.J., Yongmei Liu, Y. Lu, W.M., R.N., J.R.P., N.D.P., S.B.S. and D.J.V.D.B.
Phenotyping: A.A.A., D.K.A., M.A.A., E.P.B., R.S.C., E.D., B.I.F., O.G., S.F.A.G., J.N.H., T. Haritunians, K.C.J., A.K., C.K., E.K.K., S.L., J.E.M., M.N., R.N., A.O., H.O.-B., B.M.P., J.R.P., S.R.P., C.N.R., E.R., S.R., B.S., D.S., L.S., B.O.T. and T.R.Y.
Statistical methods and data analysis: A.A.A., D.K.A., L.F.B., C.W.K.C., G.K.C., G. Casey, N.E.C., W.-M.C., G.A.C., Y.-D.I.C., J.D., P.D., T.L.E., C.F., M.F.F., J.P.B., E.M.G., M.G., O.G., X.G., C.A.H., M.R.I., A.K., B.J.K., C.K., E.K.K., S.J.K., C.D.L., G. Lettre, G. Li, H.L., K.L., L.A.L., R.J.F.L., V.L., Yongmei Liu, Youfang Liu, Y. Lu, B.M., K.L.M., Y.A.M., A.N., K.E.N., M.A.N., M.C.Y.N., C.P., J.R.P., E.A.R.-N., S.K.R., B.P., A.P.R., L.J.R.-T., D.A.S., E.K.S., E.E.S., Y.V.S., B.O.T., K.C.T., D.R.V.E., M.K.W., Z.W., L.K.W., T.W.W., L.R.Y., J.Z., J.H.Z., N.A.Z., J.M.Z., W. Zhao, the NABEC Consortium, the UKBEC Consortium, the BioBank Japan Project and the AGEN Consortium.
Writing group: G.K.C., T.L.E., M.G., B.E.H., J.N.H., R.J.F.L., C.A.H., L.A.L., K.L.M., K.E.N., M.C.Y.N., C.P., G.J.P., A.P.R. and K.C.T.
Critical review of the manuscript: A.A.A., A.A., C.B.A., C.I.A., D.K.A., L.A., M.A.A., M.C.A., S.A., A.B., C.H.B., D.M.B., D.W.B., E.P.B., E.V.B., G.B., I.B.B., J.P.B., L.F.B., L.B., S.I.B., U.B., W.J.B., C.W.K.C., C.S.C., F.C., G.A.C., G.K.C., G. Casey, G. Chen, J.C., L.C., M.C., N.E.C., Q.C., R.S.C., S.J.C., W.-M.C., Y.-D.I.C., D.D., E.D., J.D., P.D., R.W.D., S.L.D.-H., M.K.E., T.L.E., C.F., J.K.F., M.F.F., B.I.F., Y.F., E.M.G., G.S.G., M.G., O.G., P.J.G., S.F.A.G., W.T.G., X.G., A.J.M.H., A.H., B.E.H., B.V.H., C.A.H., C.C.H., D.G.H., D.H., H.H., J.J.H., J.N.H., K.J.H., T.D.H., T. Haritunians, T. Harris, V.J.H., J.H.M., M.R.I., S.A.I., E.M.J., J.M.J., K.C.J., A.K., B.J.K., C.K., D.L.K., E.A.K., E.K.K., L.N.K., L.H.K., R.A.K., S.J.K., S.L.R.K., S. Kolb, S. Ketkar, A.M.L., C.D.L., G. Lettre, G. Li, H.L., K.L., L.A.L., L.L., M.C.L., R.J.F.L., S.L., V.L., Yongmei Liu, Youfang Liu, Y. Lu, A.B.M., B.M., J.C.M., J.E.M., L.H.M., K.L.M., K.R.M., R.C.M., T.H.M., W.M., Y.A.M., I.M.-B., A.N., B.N., K.E.N., K.L.N., M.A.N., M.C.Y.N., M.N., R.N., S.N., U.N., C.N.-D., O.I.O., O.O., T.O.O., A.O., H.O.-B., B.M.P., B.P., C.A.P., C.P., G.J.P., J.R.P., M.F.P., N.D.P., P.A.P., S.R.P., U.P., A.P.R., B.A.R., C.N.R., E.R., S.K.R., S.R., J.L.R.-G., E.A.R.-N., L.J.R.-T., A.B.S., A.G.S., B.S., D.A.S., D.S., E.K.S., E.E.S., I.J.S., J.L.S., L.B.S., L.S., M.M.S., M.R.S., P.J.S., S.B.S., S.S.S., Y.V.S., B.O.T., K.C.T., M.A.T., H.T., M.J.T., M.V., D.J.V.D.B., D.R.V.E., J.K.W., M.W., S.M.W., S.-Y.W., S.W.-S., T.W.W., X.W., Z.W., L.K.W., J.S.W., M.K.W., J.J.Y., L.R.Y., T.R.Y., A.B.Z., J.Z., J.M.Z., J.H.Z., K.A.Z., N.A.Z., R.G.Z., W. Zhao, W Zheng, X.Z. and Y.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of contributing members appears in the Supplementary Note.
A list of contributing members appears in the Supplementary Note.
A list of contributing members appears in the Supplementary Note.
A list of contributing members appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–18, Supplementary Figures 1–9 and Supplementary Note (PDF 7576 kb)
Supplementary Table 19
Comprehensive results of the biofeature analysis for the three novel loci (XLS 190 kb)
Rights and permissions
About this article
Cite this article
Monda, K., Chen, G., Taylor, K. et al. A meta-analysis identifies new loci associated with body mass index in individuals of African ancestry. Nat Genet 45, 690–696 (2013). https://doi.org/10.1038/ng.2608
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2608
This article is cited by
-
The rs1421085 variant within FTO promotes brown fat thermogenesis
Nature Metabolism (2023)
-
Identification of novel genes whose expression in adipose tissue affects body fat mass and distribution: an RNA-Seq and Mendelian Randomization study
European Journal of Human Genetics (2022)
-
Admixture mapping of anthropometric traits in the Black Women’s Health Study: evidence of a shared African ancestry component with birth weight and type 2 diabetes
Journal of Human Genetics (2022)
-
Association of fat mass and obesity-associated (FTO) gene polymorphisms with non-communicable diseases (NCDs) in the Iranian population: A systematic review of observational studies
Journal of Diabetes & Metabolic Disorders (2022)
-
Identification of stromal microenvironment characteristics and key molecular mining in pancreatic cancer
Discover Oncology (2022)