Abstract
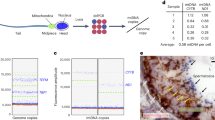
In mammals, mitochondrial DNA (mtDNA) sequence variants are observed to segregate rapidly between generations despite the high mtDNA copy number in the oocyte. This has led to the concept of a genetic bottleneck for the transmission of mtDNA1,2,3, but the mechanism remains contentious. Several studies have suggested that the bottleneck occurs during embryonic development, as a result of a marked reduction in germline mtDNA copy number4,5. Mitotic segregation of mtDNAs during preimplantation5, or during the expansion of primordial germ cells (PGCs) before they colonize the gonad4,5, is thought to account for the increase in genotypic variance observed among mature oocytes from heteroplasmic mothers. This view has, however, been challenged by studies suggesting that the bottleneck occurs without a reduction in germline mtDNA content6. To resolve this controversy, we measured mtDNA heteroplasmy and copy number in single germ cells isolated from heteroplasmic mice. By directly tracking the evolution of mtDNA genotypic variance during oogenesis, we show that the genetic bottleneck occurs during postnatal folliculogenesis and not during embryonic oogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hauswirth, W.W. & Laipis, P.J. Mitochondrial DNA polymorphism in a maternal lineage of Holstein cows. Proc. Natl. Acad. Sci. USA 79, 4686–4690 (1982).
Olivo, P.D., Van de Walle, M.J., Laipis, P.J. & Hauswirth, W.W. Nucleotide sequence evidence for rapid genotypic shifts in the bovine mitochondrial DNA D-loop. Nature 306, 400–402 (1983).
Laipis, P.J., Van de Walle, M.J. & Hauswirth, W.W. Unequal partitioning of bovine mitochondrial genotypes among siblings. Proc. Natl. Acad. Sci. USA 85, 8107–8110 (1988).
Jenuth, J.P., Peterson, A.C., Fu, K. & Shoubridge, E.A. Random genetic drift in the female germline explains the rapid segregation of mammalian mitochondrial DNA. Nat. Genet. 14, 146–151 (1996).
Cree, L.M. et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat. Genet. 40, 249–254 (2008).
Cao, L. et al. The mitochondrial bottleneck occurs without reduction of mtDNA content in female mouse germ cells. Nat. Genet. 39, 386–390 (2007).
Jansen, R.P. Germline passage of mitochondria: quantitative considerations and possible embryological sequelae. Hum. Reprod. 15 Suppl 2, 112–128 (2000).
Michaels, G.S., Hauswirth, W.W. & Laipis, P.J. Mitochondrial DNA copy number in bovine oocytes and somatic cells. Dev. Biol. 94, 246–251 (1982).
Yeom, Y.I. et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 122, 881–894 (1996).
Chiquoine, A.D. The identification, origin, and migration of the primordial germ cells in the mouse embryo. Anat. Rec. 118, 135–146 (1954).
Hayashi, K., de Sousa Lopes, S.M. & Surani, M.A. Germ cell specification in mice. Science 316, 394–396 (2007).
Snow, M.H. Gastrulation in the mouse: Growth and regionalization of the epiblast. J. Embryol. Exp. Morphol. 42, 293–303 (1977).
Kucej, M. & Butow, R.A. Evolutionary tinkering with mitochondrial nucleoids. Trends Cell Biol. 17, 586–592 (2007).
Kaufman, B.A. et al. The mitochondrial transcription factor TFAM coordinates the assembly of multiple DNA molecules into nucleoid-like structures. Mol. Biol. Cell 18, 3225–3236 (2007).
Legros, F., Malka, F., Frachon, P., Lombes, A. & Rojo, M. Organization and dynamics of human mitochondrial DNA. J. Cell Sci. 117, 2653–2662 (2004).
Kloc, M., Bilinski, S. & Etkin, L.D. The Balbiani body and germ cell determinants: 150 years later. Curr. Top. Dev. Biol. 59, 1–36 (2004).
Pepling, M.E., Wilhelm, J.E., O'Hara, A.L., Gephardt, G.W. & Spradling, A.C. Mouse oocytes within germ cell cysts and primordial follicles contain a Balbiani body. Proc. Natl. Acad. Sci. USA 104, 187–192 (2007).
Cox, R.T. & Spradling, A.C. A Balbiani body and the fusome mediate mitochondrial inheritance during Drosophila oogenesis. Development 130, 1579–1590 (2003).
Fan, W. et al. A mouse model of mitochondrial disease reveals germline selection against severe mtDNA mutations. Science 319, 958–962 (2008).
Stewart, J.B. et al. Strong purifying selection in transmission of mitochondrial DNA. PLoS Biol. 6, e10 (2008).
Shoubridge, E.A. & Wai, T. Sidestepping mutational meltdown. Science 319, 914–915 (2008).
Zhang, Y.H. et al. Mouth cell collection device for newborn mice. Mol. Genet. Metab. 89, 164–167 (2006).
El Shourbagy, S.H., Spikings, E.C., Freitas, M. & St John, J.C. Mitochondria directly influence fertilisation outcome in the pig. Reproduction 131, 233–245 (2006).
Ramakers, C., Ruijter, J.M., Deprez, R.H. & Moorman, A.F. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 339, 62–66 (2003).
Battersby, B.J., Redpath, M.E. & Shoubridge, E.A. Mitochondrial DNA segregation in hematopoietic lineages does not depend on MHC presentation of mitochondrially encoded peptides. Hum. Mol. Genet. 14, 2587–2594 (2005).
McKee, M.D. & Nanci, A. Postembedding colloidal-gold immunocytochemistry of noncollagenous extracellular matrix proteins in mineralized tissues. Microsc. Res. Tech. 31, 44–62 (1995).
Brown, M.B. & Forsythe, A.B. Robust tests for the equality of variances. J. Am. Stat. Assoc. 69, 364–367 (1974).
Pan, G. Confidence intervals for comparing two scale parameters based on Levene's statistics. J. Nonparam. Stat. 14, 459–476 (2007).
Lim, T.-S. & Loh, W.-Y. Acomparison of tests of equality of variances. Comput. Stat. Data Anal. 22, 287–301 (1995).
Battersby, B.J., Loredo-Osti, J.C. & Shoubridge, E.A. Nuclear genetic control of mitochondrial DNA segregation. Nat. Genet. 33, 183–186 (2003).
Acknowledgements
We thank L. Villeneuve for help with confocal microscopy; J. Mui for assistance with electron microscopy; T. Johns, F. Jones and D. Sabour for technical assistance; and J. Correa for statistical design. We are grateful for antibodies directed against TFAM (B. Kaufman, MNI), mt-SSB (M. Zeviani, Instituto Carlo Besta) and POLG (W. Copeland, US National Institutes of Health). The OCT4Δ PE-EGFP mice were obtained from H.R. Scholer (University of Pennsylvania). This research was supported by the Canadian Institutes of Health Research and the US National Institutes of Health. E.A.S. is an International Scholar of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.W. and E.A.S. designed the study and wrote the manuscript. D.T. performed genotyping, and T.W. did all other experiments.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1 and 2 (PDF 1950 kb)
Supplementary Video 1
3D view of the merged maximum projection close-up for primary oocytes in Fig. 2 (MOV 234 kb)
Supplementary Video 2
3D view of the merged maximum projection close-up for neonatal heart in Fig. 2 (MOV 463 kb)
Rights and permissions
About this article
Cite this article
Wai, T., Teoli, D. & Shoubridge, E. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat Genet 40, 1484–1488 (2008). https://doi.org/10.1038/ng.258
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.258
This article is cited by
-
Effects of high-altitude hypoxia on embryonic developmental potential in women undergoing IVF/ICSI procedures
Archives of Gynecology and Obstetrics (2023)
-
LONP-1 and ATFS-1 sustain deleterious heteroplasmy by promoting mtDNA replication in dysfunctional mitochondria
Nature Cell Biology (2022)
-
Comparison of mitochondrial DNA sequences from whole blood and lymphoblastoid cell lines
Scientific Reports (2022)
-
Oocytes maintain ROS-free mitochondrial metabolism by suppressing complex I
Nature (2022)
-
Reduction of mtDNA heteroplasmy in mitochondrial replacement therapy by inducing forced mitophagy
Nature Biomedical Engineering (2022)