Abstract
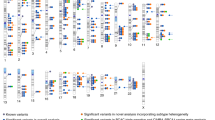
Estrogen receptor (ER)-negative tumors represent 20–30% of all breast cancers, with a higher proportion occurring in younger women and women of African ancestry1. The etiology2 and clinical behavior3 of ER-negative tumors are different from those of tumors expressing ER (ER positive), including differences in genetic predisposition4. To identify susceptibility loci specific to ER-negative disease, we combined in a meta-analysis 3 genome-wide association studies of 4,193 ER-negative breast cancer cases and 35,194 controls with a series of 40 follow-up studies (6,514 cases and 41,455 controls), genotyped using a custom Illumina array, iCOGS, developed by the Collaborative Oncological Gene-environment Study (COGS). SNPs at four loci, 1q32.1 (MDM4, P = 2.1 × 10−12 and LGR6, P = 1.4 × 10−8), 2p24.1 (P = 4.6 × 10−8) and 16q12.2 (FTO, P = 4.0 × 10−8), were associated with ER-negative but not ER-positive breast cancer (P > 0.05). These findings provide further evidence for distinct etiological pathways associated with invasive ER-positive and ER-negative breast cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Chu, K.C. & Anderson, W.F. Rates for breast cancer characteristics by estrogen and progesterone receptor status in the major racial/ethnic groups. Breast Cancer Res. Treat. 74, 199–211 (2002).
Yang, X.R. et al. Associations of breast cancer risk factors with tumor subtypes: a pooled analysis from the Breast Cancer Association Consortium studies. J. Natl. Cancer Inst. 103, 250–263 (2011).
Blows, F.M. et al. Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med. 7, e1000279 (2010).
Mavaddat, N., Antoniou, A.C., Easton, D.F. & Garcia-Closas, M. Genetic susceptibility to breast cancer. Mol. Oncol. 4, 174–191 (2010).
Haiman, C.A. et al. A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor–negative breast cancer. Nat. Genet. 43, 1210–1214 (2011).
Stevens, K.N. et al. 19p13.1 is a triple negative–specific breast cancer susceptibility locus. Cancer Res. 72, 1795–1803 (2012).
Siddiq, A. et al. A meta-analysis of genome-wide association studies of breast cancer identifies two novel susceptibility loci at 6q14 and 20q11. Hum. Mol. Genet. 21, 5373–5384 (2012).
Michailidou, K. et al. Large-scale genotyping identifies 41 new breast cancer susceptibility loci. Nat. Genet. published online; doi:10.1038/ng.2563 (27 March 2013).
Turashvili, G. et al. Novel markers for differentiation of lobular and ductal invasive breast carcinomas by laser microdissection and microarray analysis. BMC Cancer 7, 55 (2007).
Graham, K. et al. Gene expression in histologically normal epithelium from breast cancer patients and from cancer-free prophylactic mastectomy patients shares a similar profile. Br. J. Cancer 102, 1284–1293 (2010).
Wang, H. & Yan, C. A small-molecule p53 activator induces apoptosis through inhibiting MDMX expression in breast cancer cells. Neoplasia 13, 611–619 (2011).
Couch, F.J. et al. Genome-wide association study in BRCA1 mutation carriers identifies novel loci associated with breast and ovarian cancer risk. PLoS Genet. 9, e1003212 (2013).
Atwal, G.S. et al. Altered tumor formation and evolutionary selection of genetic variants in the human MDM4 oncogene. Proc. Natl. Acad. Sci. USA 106, 10236–10241 (2009).
Frayling, T.M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Bojesen, S.E. et al. Multiple independent TERT variants associated with telomere length and risks of breast and ovarian cancer. Nat. Genet. published online; doi:10.1038/ng.2566 (27 March 2013).
Shen, H. et al. Epigenetic analysis leads to identification of HNF1B as a subtype-specific susceptibility gene for ovarian cancer. Nat. Comm. published online; doi:10.1038/ncomms2629 (27 March 2013).
Hunter, D.J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat. Genet. 39, 870–874 (2007).
Aulchenko, Y.S., Struchalin, M.V. & van Duijn, C.M. ProbABEL package for genome-wide association analysis of imputed data. BMC Bioinformatics 11, 134 (2010).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Coetzee, S.G., Rhie, S.K., Berman, B.P., Coetzee, G.A. & Noushmehr, H. FunciSNP: an R/bioconductor tool integrating functional non-coding data sets with genetic association studies to identify candidate regulatory SNPs. Nucleic Acids Res. 40, e139 (2012).
Acknowledgements
The authors wish to thank all the individuals who took part in these studies and all the researchers, clinicians and administrative staff who have enabled this work to be carried out. We are very grateful to Illumina, in particular J. Stone, S. McBean, J. Hadlington, A. Mustafa and K. Cook, for their help with designing the array. BCAC is funded by Cancer Research UK (C1287/A10118 and C1287/A12014) and by the European Community's Seventh Framework Programme under grant agreement 223175 (HEALTH-F2-2009-223175) (COGS). Meetings of BCAC have been funded by the European Union European Cooperation in Science and Technology (COST) programme (BM0606). BPC3 is funded by US National Cancer Institute cooperative agreements U01-CA98233, U01-CA98710, U01-CA98216 and U01-CA98758 and the Intramural Research Program of the US National Institutes of Health (NIH)/National Cancer Institute, Division of Cancer Epidemiology and Genetics. TNBCC is supported by Mayo Clinic Breast Cancer Study (MCBCS) (US NIH grants CA122340 and a Specialized Program of Research Excellence (SPORE) in Breast Cancer (CA116201)), grants from the Komen Foundation for the Cure and the Breast Cancer Research Foundation. Genotyping on the iCOGS array was funded by the European Union (HEALTH-F2-2009-223175), Cancer Research UK (C1287/A10710), US NIH grant CA122340, the Komen Foundation for the Cure, the Breast Cancer Research Foundation, the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer program (J. Simiard and D.E.) and Ministry of Economic Development, Innovation and Export Trade of Quebec grant PSR-SIIRI-701 (J. Simiard, D.E. and P.H.). J. Simiard holds the Canada Research Chair in Oncogenetics. Combination of the GWAS data was supported in part by US NIH Cancer Post-Cancer GWAS initiative grant U19 CA 148065-01 (DRIVE, part of the GAME-ON initiative) and Breakthrough Breast Cancer Research.
Author information
Authors and Affiliations
Consortia
Contributions
M.G.-C., F.J.C., S.L., K. Michailidou, M.K.S., P.D.P.P., C.V., D.F.E., C.A.H. and P. Kraft formed the writing group and drafted the manuscript. M.G.-C. coordinated the writing group. F.J.C., S.L., K. Michaildou, D.F.E., C.A.H. and P. Kraft performed statistical analyses of GWAS data. M.G.-C. and M.N.B. performed statistical analyses of BCAC follow-up studies and meta-analyses. P. Kraft coordinated the BPC3 GWAS, and M.G.-C., E.R., H.S.F., L.L.M., J.E.B., W.C.W., D.J.H. and S.J.C. led individual studies in the BPC3 scan. F.J.C. and C.V. coordinated the TNBCC GWAS, and D.E., P. Miron, P.A.F., J.C.-C., J.C., A.A., H.N., H. Brauch and G.G.G. led individual studies in the TNBCC scan. D.E. coordinated the C-BCAC GWAS, and H.N., J.L.H., J.C.-C. and P.H. led individual studies in the C-BCAC scan. D.F.E. conceived and coordinated the synthesis of the iCOGS array and led BCAC. P.H. coordinated COGS, and J.B. led the BCAC genotyping working group. A.G.-N., G.P., M.R.A., D.V., F.B., D.C.T. and F.J.C. coordinated genotyping of the iCOGS array. M.G.-C., P.D.P.P. and M.K.S. led the pathology working group in BCAC. M.E.S. was the lead pathologist in BCAC. W.J.H. performed automated scoring of tissue microarrays. A.M.D. and G.C.-T. led the quality control working group. J.D. and N.O. provided bioinformatics support. S.K.R. and G.A.C. performed FunciSNP bioinformatics analyses. M.K.B. and Q. Wang provided data management support for BCAC. G.G., A.A., A. Broeks, A.B.E., A.C., U.H., A.-S.D., A.G.U., A.H., A.H.W., A.I., the ABCTB Investigators, A.J.-V., A.J., A.K.G., R.W., A. Lindblom, A. Lophatananon, A.M.D., A.M.M., A.M.W.v.d.O., A.R., A. Swerdlow, A. Schneeweiss, B.B., B.E.H., B.G.N., B.M.-M., B.P., C.B., C.B.A., C.-Y.C., C.C., C.D.B., C.-N.H., C.H.M.v.D., C.H.Y., C.J., C.M., C.M.S., C.O., C.R., C.-Y.S., C. Sohn, C. Stegmaier, C.-C.T., C.T., C.W.C., D.C., D.C.T., D.F.-J., D.G., D.I.C., D.J.P., D.J.S., D.K., D.L., D.O.S., D.S., D.T., D.V.D.B., E.D., C.V., E.J.R., E.J.S., E.M., E.M.J., E.V.B., E.W., F.A., FBCS, F.C.-C., F.C., F.H., F.L., F.M., F.R., F.S., G.A.C., G.C.-T., G.K.C., G.S., G.W.M., H.A.-C., H.C., H.F., H. Ito, H. Iwata, H. Müller, H. Miao, H.M.-H., H.P., H.T., H.W., I.d.S.S., I.K., I.L.A., I.T., J.A.K., J.D.F., J.E.O., J.I.A.P., J.J.H., J. Long, J. Lubinski, J. Liu, J. Lissowska, J.L.R.-G., J.M.H., J.P., J. Stone, J. Simard, J.W., J.-C.Y., K. Aittomäki, K. Aaltonen, K.C., K.D., K.J., K.-T.K., K.L., K. Muir, K. Matsuo, K.P., K.S., K.S.C., L. Bernard, L. Baglietto, L. Bernstein, L. Beckmann, L.D., L.G., L.J.V.V., L.N.K., L.S., M.B., M.C.S., M.D., M.F.P., M.G.S., M. Jones, M. Johansson, M.J.H., M.J.K., M.K., M.K.B., M.L., M.M.G., M.P.L., M. Shrubsole, M. Shah, M.W.B., M.W.R.R., N.A.M.T., N.D., N.G.M., N.J., N.M., N.N.A., N.R., N.S., N.V.B., O.F., P.G., P.H., P.H.P., P. Kerbrat, P.L.-P., P.L., P. Menénde, P.N., P.P., P.R., P. Siriwanarangsan, P. Sharma, P.-E.W., Q.C., Q. Wang, Q. Waisfisz, R.B., R.G.Z., R.H., R.K., R.K.S., R.L.M., R.M.M., R.N.H., R.P., R.A.E.M.T., R. Tumino, R. Travis, S.A.I., S.E.B., S.E.H., S.F.N., S.G., S.H.T., S.K., S.K.R., S.L.D.-H., S.M., S.M.J., S. Nickels, S. Nyante, S.P.B., S. Sangrajrang, S.S.-B., S. Slager, S.S.C., T.A.M., T.B., T.D., T.H., T.T., V.A., V. Kristensen, V. Kataja, V.-M.K., W.B., W.L., W.R.D., W.T., X.-O.S., X.W., Y.F., Y.-T.G., Y.-D.K. and Y.Y. contributed to GWAS and/or BCAC follow-up studies. M.G.-C., F.J.C., S.L., K. Michailidou, M.K.S., P.D.P.P., C.V., D.F.E., C.A.H., P. Kraft, M.N.B., E.R., H.S.F., L.L.M., J.E.B., W.C.W., D.J.H., S.J.C., D.E., P.A.F., J.C.-C., J.C., A. Broeks, H.N., H. Brauch, H. Brenner, G.P., G.G.G., J.L.H., P. Miron, J.B., A.G.-N., M.R.A., D.V., F.B., M.E.S., W.J.H., G.G., A.A., A. Beck, A.B.E., A.C., U.H., A.-S.D., A.G.U., A.H., A.H.W., A.I., the ABCTB Investigators, A.J.-V., A.J., A.K.G., R.W., A. Lindblom, A. Lophatananon, A.M.D., A.M.M., A.M.W.v.d.O., A.R., A. Swerdlow, A. Schneeweiss, B.B., B.E.H., B.G.N., B.M.-M., B.P., C.B., C.B.A., C.-Y.C., C.C., C.D.B., C.-N.H., C.H.M.v.D., C.H.Y., C.J., C.M., C.M.S., C.O., C.R., C.-Y.S., C. Sohn, C. Stegmaier, C.-C.T., C.T., C.W.C., D.C., D.C.T., D.F.-J., D.G., D.I.C., D.J.P., D.J.S., D.K., D.L., D.O.S., D.S., D.T., D.V.D.B., E.D., C.V., E.J.R., E.J.S., E.M., E.M.J., E.V.B., E.W., F.A., FBCS, F.C.-C., F.C., F.H., F.L., F.M., F.R., F.S., G.A.C., G.C.-T., G.K.C., G.S., G.W.M., H.A.-C., H.C., H.F., H. Ito, H. Iwata, H. Müller, H. Miao, H.M.-H., H.P., H.T., H.W., I.d.S.S., I.K., I.L.A., I.T., J.A.K., J.D., J.D.F., J.E.O., J.I.A.P., J.J.H., J. Long, J. Lubinski, J. Liu, J. Lissowska, J.L.R.-G., J.M.H., J.P., J. Stone, J. Simard, J.W., J.-C.Y., K. Aittomäki, K. Aaltonen, K.C., K.D., K.J., K.-T.K., K.L., K. Muir, K. Matsuo, K.P., K.S., K.S.C., L. Bernard, L. Baglietto, L. Bernstein, L. Beckmann, L.D., L.G., L.J.V.V., L.N.K., L.S., M.B., M.C.S., M.D., M.F.P., M.G.S., M.H., M. Jones, M. Johansson, M.J.H., M.J.K., M.K., M.K.B., M.L., M.M.G., M.P.L., M. Shrubsole, M. Shah, M.W.B., M.W.R.R., N.A.M.T., N.D., N.G.M., N.J., N.M., N.N.A., N.O., N.R., N.S., N.V.B., O.F., P.G., P.H., P.H.P., P. Kerbrat, P.L.-P., P.L., P. Menénde, P.N., P.P., P.R., P. Siriwanarangsan, P. Sharma, P.-E.W., Q.C., Q. Wang, Q. Waisfisz, R.B., R.G.Z., R.H., R.K., R.K.S., R.L.M., R.M.M., R.N.H., R.P., R.A.E.M.T., R. Tumino, R. Travis, S.A.I., S.E.B., S.E.H., S.F.N., S.G., S.H.T., S.K., S.K.R., S.L.D.-H., S.M., S.M.J., S. Nickels, S. Nyante, S.P.B., S. Sangrajrang, S.S.-B., S. Slager, S.S.C., T.A.M., T.B., T.D., T.H., T.T., V.A., V. Kristensen, V. Kataja, V.-M.K., W.B., W.L., W.R.D., W.T., X.-O.S., X.W., Y.F., Y.-T.G., Y.-D.K. A. Mannermaa, A. Meindl, W.Z., P.D., M.S.G. and Y.Y. provided critical review of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
A list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–9 and Supplementary Note (PDF 6460 kb)
Rights and permissions
About this article
Cite this article
Garcia-Closas, M., Couch, F., Lindstrom, S. et al. Genome-wide association studies identify four ER negative–specific breast cancer risk loci. Nat Genet 45, 392–398 (2013). https://doi.org/10.1038/ng.2561
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2561
This article is cited by
-
Understanding the contribution of lifestyle in breast cancer risk prediction: a systematic review of models applicable to Europe
BMC Cancer (2023)
-
MDM4: What do we know about the association between its polymorphisms and cancer?
Medical Oncology (2022)
-
MiR-556-5p modulates migration, invasion, and epithelial-mesenchymal transition in breast cancer cells via targeting PTHrP
Journal of Molecular Histology (2022)
-
Association of KLK3, VAMP8 and MDM4 Genetic Variants within microRNA Binding Sites with Prostate Cancer: Evidence from Serbian Population
Pathology & Oncology Research (2020)
-
Genome-wide association study identifies 32 novel breast cancer susceptibility loci from overall and subtype-specific analyses
Nature Genetics (2020)