Abstract
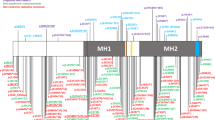
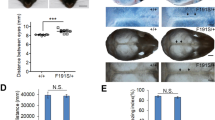
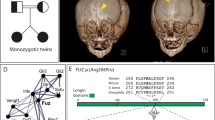
The extracellular signal–related kinases 1 and 2 (ERK1/2) are key proteins mediating mitogen-activated protein kinase signaling downstream of RAS: phosphorylation of ERK1/2 leads to nuclear uptake and modulation of multiple targets1. Here, we show that reduced dosage of ERF, which encodes an inhibitory ETS transcription factor directly bound by ERK1/2 (refs. 2,3,4,5,6,7), causes complex craniosynostosis (premature fusion of the cranial sutures) in humans and mice. Features of this newly recognized clinical disorder include multiple-suture synostosis, craniofacial dysmorphism, Chiari malformation and language delay. Mice with functional Erf levels reduced to ∼30% of normal exhibit postnatal multiple-suture synostosis; by contrast, embryonic calvarial development appears mildly delayed. Using chromatin immunoprecipitation in mouse embryonic fibroblasts and high-throughput sequencing, we find that ERF binds preferentially to elements away from promoters that contain RUNX or AP-1 motifs. This work identifies ERF as a novel regulator of osteogenic stimulation by RAS-ERK signaling, potentially by competing with activating ETS factors in multifactor transcriptional complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Plotnikov, A., Zehorai, E., Procaccia, S. & Seger, R. The MAPK cascades: signaling components, nuclear roles and mechanisms of nuclear translocation. Biochim. Biophys. Acta 1813, 1619–1633 (2011).
Mavrothalassitis, G.J. & Papas, T.S. Positive and negative factors regulate the transcription of the ETS2 gene via an oncogene-responsive–like unit within the ETS2 promoter region. Cell Growth Differ. 2, 215–224 (1991).
Sgouras, D.N. et al. ERF: an ETS domain protein with strong transcriptional repressor activity, can suppress ets-associated tumorigenesis and is regulated by phosphorylation during cell cycle and mitogenic stimulation. EMBO J. 14, 4781–4793 (1995).
Le Gallic, L., Sgouras, D., Beal, G. & Mavrothalassitis, G. Transcriptional repressor ERF is a Ras/mitogen-activated protein kinase target that regulates cellular proliferation. Mol. Cell Biol. 19, 4121–4133 (1999).
Le Gallic, L., Virgilio, L., Cohen, P., Biteau, B. & Mavrothalassitis, G. ERF nuclear shuttling, a continuous monitor of Erk activity that links it to cell cycle progression. Mol. Cell Biol. 24, 1206–1218 (2004).
Polychronopoulos, S. et al. The transcriptional ETS2 repressor factor associates with active and inactive Erks through distinct FXF motifs. J. Biol. Chem. 281, 25601–25611 (2006).
von Kriegsheim, A. et al. Cell fate decisions are specified by the dynamic ERK interactome. Nat. Cell Biol. 11, 1458–1464 (2009).
Ng, S.B. et al. Exome sequencing identifies the cause of a mendelian disorder. Nat. Genet. 42, 30–35 (2010).
Kim, H.-J. Erk pathway and activator protein 1 play crucial roles in FGF2-stimulated premature cranial suture closure. Dev. Dyn. 227, 335–346 (2003).
Shukla, V., Coumoul, X., Wang, R.-H., Kim, H.-S. & Deng, C.-X. RNA interference and inhibition of MEK-ERK signaling prevent abnormal skeletal phenotypes in a mouse model of craniosynostosis. Nat. Genet. 39, 1145–1150 (2007).
Hollenhorst, P.C., McIntosh, L.P. & Graves, B.J. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu. Rev. Biochem. 80, 437–471 (2011).
Wei, G.-H. Genome-wide analysis of ETS-family DNA-binding in vitro and in vivo. EMBO J. 29, 2147–2160 (2010).
Wilkie, A.O.M. et al. Prevalence and complications of single-gene and chromosomal disorders in craniosynostosis. Pediatrics 126, e391–e400 (2010).
Kan, S.-H. et al. Genomic screening of fibroblast growth-factor receptor 2 reveals a wide spectrum of mutations in patients with syndromic craniosynostosis. Am. J. Hum. Genet. 70, 472–486 (2002).
Lajeunie, E. et al. Mutation screening in patients with syndromic craniosynostoses indicates that a limited number of recurrent FGFR2 mutations accounts for severe forms of Pfeiffer syndrome. Eur. J. Hum. Genet. 14, 289–298 (2006).
Hammond, P. & Suttie, M.J. Large-scale phenotyping of 3D facial morphology. Hum. Mutat. 33, 817–825 (2012).
Papadaki, C. et al. Transcriptional repressor Erf determines extraembryonic ectoderm differentiation. Mol. Cell Biol. 27, 5201–5213 (2007).
Mundlos, S. et al. Mutations involving the transcription factor CBFA1 cause cleidocranial dysplasia. Cell 89, 773–779 (1997).
Park, M.-H. et al. Differential expression patterns of Runx2 isoforms in cranial suture morphogenesis. J. Bone Miner. Res. 16, 885–892 (2001).
Komori, T. Signaling networks in RUNX2-dependent bone development. J. Cell Biochem. 112, 750–755 (2011).
Mefford, H.C. et al. Copy number variation analysis in single-suture craniosynostosis: multiple rare variants including RUNX2 duplication in two cousins with metopic craniosynostosis. Am. J. Med. Genet. 152A, 2203–2210 (2010).
Varvagiannis, K. et al. Pure de novo partial trisomy 6p in a girl with craniosynostosis. Am. J. Med. Genet. A. published online; 10.1002/ajmg.a.35727 (10 January 2013).
Iseki, S., Wilkie, A.O.M. & Morriss-Kay, G.M. Fgfr1 and Fgfr2 have distinct differentation- and proliferation-related roles in the developing mouse skull vault. Development 126, 5611–5620 (1999).
Hecht, J. et al. Detection of novel skeletogenesis target genes by comprehensive analysis of a Runx2−/− mouse model. Gene Expr. Patterns 7, 102–112 (2007).
Morriss-Kay, G.M. & Wilkie, A.O.M. Growth of the normal skull vault and its alteration in craniosynostosis: insights from human genetics and experimental studies. J. Anat. 207, 637–653 (2005).
Verykokakis, M., Papadaki, C., Vorgia, E., Le Gallic, L. & Mavrothalassitis, G. The RAS-dependent ERF control of cell proliferation and differentiation is mediated by c-Myc repression. J. Biol. Chem. 282, 30285–30294 (2007).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Dejosez, M. et al. Ronin/Hcf-1 binds to a hyperconserved enhancer element and regulates genes involved in the growth of embryonic stem cells. Genes Dev. 24, 1479–1484 (2010).
Hollenhorst, P.C. et al. DNA specificity determinants associate with distinct transcription factor functions. PLoS Genet. 5, e1000778 (2009).
Hollenhorst, P.C. et al. Oncogenic ETS proteins mimic activated RAS/MAPK signaling in prostate cells. Genes Dev. 25, 2147–2157 (2011).
Goetz, T.L., Gu, T.L., Speck, N.A. & Graves, B.J. Auto-inhibition of Ets-1 is counteracted by DNA binding cooperativity with core-binding factor α2. Mol. Cell Biol. 20, 81–90 (2000).
Hess, J., Porte, D., Munz, C. & Angel, P. AP-1 and Cbfa/runt physically interact and regulate parathyroid hormone–dependent MMP13 expression in osteoblasts through a new osteoblast-specific element 2/AP-1 composite element. J. Biol. Chem. 276, 20029–20038 (2001).
D′Alonzo, R.C., Selvamurugan, N., Karsenty, G. & Partridge, N.C. Physical interaction of the activator protein-1 factors c-Fos and c-Jun with Cbfa1 for collagenase-3 promoter activation. J. Biol. Chem. 277, 816–822 (2002).
Allegra, M. et al. Semaphorin-7a reverses the ERF-induced inhibition of EMT in Ras-dependent mouse mammary epithelial cells. Mol. Biol. Cell 23, 3873–3881 (2012).
Lou, Y. et al. A Runx2 threshold for the cleidocranial dysplasia phenotype. Hum. Mol. Genet. 18, 556–568 (2009).
Sumarsono, S.H. et al. Down′s syndrome–like skeletal abnormalities in Ets2 transgenic mice. Nature 379, 534–537 (1996).
Vary, C.P.H. et al. Involvement of Ets transcription factors and targets in osteoblast differentiation and matrix mineralization. Exp. Cell Res. 257, 213–222 (2000).
Little, G.H. et al. Genome-wide Runx2 occupancy in prostate cancer cells suggests a role in regulating secretion. Nucleic Acids Res. 40, 3538–3547 (2012).
Wai, P.Y. et al. Ets-1 and runx2 regulate transcription of a metastatic gene, osteopontin, in murine colorectal cancer cells. J. Biol. Chem. 281, 18973–18982 (2006).
Ge, C. et al. Identification and functional characterization of ERK/MAPK phosphorylation sites in the Runx2 transcription factor. J. Biol. Chem. 284, 32533–32543 (2009).
Park, O.-J., Kim, H.-J., Woo, K.-M., Baek, J.-H. & Ryoo, H.-M. FGF2-activated ERK mitogen-activated protein kinase enhances Runx2 acetylation and stabilization. J. Biol. Chem. 285, 3568–3574 (2010).
Matsushita, T. et al. Extracellular signal–regulated kinase 1 (ERK1) and ERK2 play essential roles in osteoblast differentiation and in supporting osteoclastogenesis. Mol. Cell Biol. 29, 5843–5857 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Hammond, P. The use of 3D face shape modelling in dysmorphology. Arch. Dis. Child. 92, 1120–1126 (2007).
Watson, D.K. et al. The ERGB/Fli-1 gene: isolation and characterization of a new member of the family of human ETS transcription factors. Cell Growth Differ. 3, 705–713 (1992).
Spyropoulos, D.D. et al. Hemorrhage, impaired hematopoiesis, and lethality in mouse embryos carrying a targeted disruption of the Fli1 transcription factor. Mol. Cell Biol. 20, 5643–5652 (2000).
Hogan, B. Manipulating the Mouse Embryo: a Laboratory Manual (Cold Spring Harbor Laboratory Press, Plainview, New York, 1994).
Tallquist, M.D. & Soriano, P. Epiblast-restricted Cre expression in MORE mice: a tool to distinguish embryonic vs. extra-embryonic gene function. Genesis 26, 113–115 (2000).
Wilkinson, D.G. In-situ Hybridization: a Practical Approach (Oxford University Press, Oxford, 1992).
Kowalczyk, M.S. et al. Intragenic enhancers act as alternative promoters. Mol. Cell 45, 447–458 (2012).
Acknowledgements
We thank R. Boehm, H. Care, C. Langman, J. Phipps and E. Sweeney for clinical assistance, L. Gregory, P. Piazza and staff at the High-Throughput Genomics facility at the Wellcome Trust Centre for Human Genetics for exome sequencing, C. Babbs, S. Butler, J. Frankland, C. Rode and T. Rostron for technical help, K. Kourouniotis for blastocyst injections and expert animal support, E. Giannoulatou for bioinformatics assistance, and B. Graves and P. Hollenhorst for discussions. We thank J. Heath (University of Birmingham), G. Schwabe (Charité University Hospital) and D. Rice (University of Helsinki) for the gifts of the Spp1, Runx2 and Bglap2 probes, respectively. This work was funded by the Greek Ministry of Education grants PYTHAGORAS II KA2092, PENED 03ED626, HERAKLEITOS II KA 3396 and SYNERGASIA 09SYN-11-902 to G.M., the NIHR Biomedical Research Centre, with funding from the Department of Health's NIHR Biomedical Research Centres funding scheme (S.J.L.K. and A.O.M.W.), the Oxford Craniofacial Unit Charitable Fund (V.P.S.), the Department of Health, UK, Quality, Improvement, Development and Initiative Scheme (QIDIS) (V.P.S.) and the Wellcome Trust (090532, S.J.L.K.; 093329, S.R.F.T. and A.O.M.W.). The views expressed in this publication are those of the authors and not necessarily those of the Department of Health, UK.
Author information
Authors and Affiliations
Contributions
S.R.F.T. designed and performed experiments and wrote the manuscript. M.A., I.P., A.L.F., E.V., A.Z., E.S.A., S.J.L.K., H.L. and T.L. performed experiments. S.J.M., J.H. and S.T. performed bioinformatic analyses. V.P.S. performed experiments and assessed patients. L.I., A.K.L., S.N.M., F.J.S., A.V., L.C.W., D.J. and S.A.W. identified and assessed patients. C.H. and P.T.S. performed and analyzed μCT scans. P.H. performed and analyzed three-dimensional facial imaging. G.M. conceived the project, designed experiments and wrote the manuscript. A.O.M.W. conceived the project, assessed patients, designed experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Foundation for Research and Technology–Hellas has filed a patent application with the UK Intellectual Property Office for the use of animals with decreased Erf activity in drug development for ossification defects.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3, 9 and 10 and Supplementary Figures 1–10 (PDF 7508 kb)
Supplementary Table 4
ChIP-Seq targets identified with -FCS/+FCS >3 (non-TSS). (XLSX 339 kb)
Supplementary Table 5
ChIP-Seq targets identified with -FCS/+FCS >3 (TSS). (XLSX 250 kb)
Supplementary Table 6
Sequences near differentially bound ChIP-Seq targets (-FCS/+FCS >3; non-TSS) enriched for combinations of AP1, RUNX and ETS consensus binding sites – AP1. (XLSX 1153 kb)
Supplementary Table 7
Sequences near differentially bound ChIP-Seq targets (-FCS/+FCS >3; non-TSS) enriched for combinations of AP1, RUNX and ETS consensus binding sites – RUNX2. (XLSX 2863 kb)
Supplementary Table 8
Orthologous genes within 40 kb of ChIP-Seq peaks shared between datasets for Erf (-FCS/+FCS >3; non-TSS) in mouse embryonic fibroblasts (this work) and RUNX2 in human prostate carcinoma cells.38 (XLSX 23 kb)
Supplementary Movie 1
Animated morphs of faces of 4 ERF mutation-positive individuals from Family 1. (GIF 1693 kb)
Supplementary Movie 2
Animated morphs of faces of 2 ERF mutation-positive individuals from Family 4. (GIF 805 kb)
Supplementary Movie 3
Animated morphs of faces of 3 ERF mutation-positive individuals from Family 6. (GIF 1312 kb)
Supplementary Movie 4
Animated morphs of faces of 3 ERF mutation-positive individuals from Family 8. (GIF 1233 kb)
Supplementary Movie 5
Animated morphs of face of mutation-positive individual II-1 from Family 10. (GIF 338 kb)
Supplementary Movie 6
Animated morphs of faces of 3 ERF mutation-positive individuals from Family 11. (GIF 1171 kb)
Rights and permissions
About this article
Cite this article
Twigg, S., Vorgia, E., McGowan, S. et al. Reduced dosage of ERF causes complex craniosynostosis in humans and mice and links ERK1/2 signaling to regulation of osteogenesis. Nat Genet 45, 308–313 (2013). https://doi.org/10.1038/ng.2539
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2539
This article is cited by
-
ERF-related craniosynostosis and surgical management in the paediatric cohort
Child's Nervous System (2023)
-
Retroperitoneal liposarcoma and craniosynostosis: possible genomic relationship, case report, and literature review
Functional & Integrative Genomics (2023)
-
Genetic aetiologies for childhood speech disorder: novel pathways co-expressed during brain development
Molecular Psychiatry (2023)
-
Detection of untreated sewage discharges to watercourses using machine learning
npj Clean Water (2021)
-
Chiari 1 malformation and exome sequencing in 51 trios: the emerging role of rare missense variants in chromatin-remodeling genes
Human Genetics (2021)