Abstract
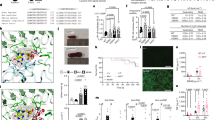
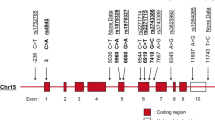
The c-Src tyrosine kinase, Csk, physically interacts with the intracellular phosphatase Lyp (encoded by PTPN22) and can modify the activation state of downstream Src kinases, such as Lyn, in lymphocytes. We identified an association of CSK with systemic lupus erythematosus (SLE) and refined its location to the intronic polymorphism rs34933034 (odds ratio (OR) = 1.32; P = 1.04 × 10−9). The risk allele at this SNP is associated with increased CSK expression and augments inhibitory phosphorylation of Lyn. In carriers of the risk allele, there is increased B-cell receptor (BCR)-mediated activation of mature B cells, as well as higher concentrations of plasma immunoglobulin M (IgM), relative to individuals with the non-risk haplotype. Moreover, the fraction of transitional B cells is doubled in the cord blood of carriers of the risk allele, due to an expansion of late transitional cells in a stage targeted by selection mechanisms. This suggests that the Lyp-Csk complex increases susceptibility to lupus at multiple maturation and activation points in B cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cho, J.H. & Gregersen, P.K. Genomics and the multifactorial nature of human autoimmune disease. N. Engl. J. Med. 365, 1612–1623 (2011).
Liston, A., Lesage, S., Gray, D.H.D., Boyd, R.L. & Goodnow, C.C. Genetic lesions in T-cell tolerance and thresholds for autoimmunity. Immunol. Rev. 204, 87–101 (2005).
Hom, G. et al. Association of systemic lupus erythematosus with C8orf13-BLK and ITGAM-ITGAX. N. Engl. J. Med. 358, 900–909 (2008).
Nath, S.K. et al. A nonsynonymous functional variant in integrin-αM (encoded by ITGAM) is associated with systemic lupus erythematosus. Nat. Genet. 40, 152–154 (2008).
Levinson, N.M., Seeliger, M.A., Cole, P.A. & Kuriyan, J. Structural basis for the recognition of c-Src by its inactivator Csk. Cell 134, 124–134 (2008).
Gregersen, P.K., Lee, H.-S., Batliwalla, F. & Begovich, A.B. PTPN22: setting thresholds for autoimmunity. Semin. Immunol. 18, 214–223 (2006).
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat. Genet. 43, 1193–1201 (2011).
Martin, J.-E. et al. Identification of CSK as a systemic sclerosis genetic risk factor through Genome Wide Association Study follow-up. Hum. Mol. Genet. 21, 2825–2835 (2012).
Su, A.I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
Zikherman, J. et al. CD45-Csk phosphatase-kinase titration uncouples basal and inducible T cell receptor signaling during thymic development. Immunity 32, 342–354 (2010).
Hasegawa, M. et al. A CD19-dependent signaling pathway regulates autoimmunity in Lyn-deficient mice. J. Immunol. 167, 2469–2478 (2001).
Arechiga, A.F. et al. Cutting edge: the PTPN22 allelic variant associated with autoimmunity impairs B cell signaling. J. Immunol. 182, 3343–3347 (2009).
Zikherman, J. et al. PTPN22 deficiency cooperates with the CD45 E613R allele to break tolerance on a non-autoimmune background. J. Immunol. 182, 4093–4106 (2009).
Zhang, J. et al. The autoimmune disease–associated PTPN22 variant promotes calpain-mediated Lyp/Pep degradation associated with lymphocyte and dendritic cell hyperresponsiveness. Nat. Genet. 43, 902–907 (2011).
Menard, L. et al. The PTPN22 allele encoding an R620W variant interferes with the removal of developing autoreactive B cells in humans. J. Clin. Invest. 121, 3635–3644 (2011).
Lu, R. et al. Genetic associations of LYN with systemic lupus erythematosus. Genes Immun. 10, 397–403 (2009).
Simpfendorfer, K.R. et al. The autoimmunity-associated BLK haplotype exhibits cis-regulatory effects on mRNA and protein expression that are prominently observed in B cells early in development. Hum. Mol. Genet. 21, 3918–3925 (2012).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Thorburn, C.M. et al. Association of PDCD1 genetic variation with risk and clinical manifestations of systemic lupus erythematosus in a multiethnic cohort. Genes Immun. 8, 279–287 (2007).
Criswell, L.A. et al. Analysis of families in the multiple autoimmune disease genetics consortium (MADGC) collection: the PTPN22 620W allele associates with multiple autoimmune phenotypes. Am. J. Hum. Genet. 76, 561–571 (2005).
Graham, R.R. et al. A common haplotype of interferon regulatory factor 5 (IRF5) regulates splicing and expression and is associated with increased risk of systemic lupus erythematosus. Nat. Genet. 38, 550–555 (2006).
Cunninghame Graham, D.S. et al. Association of NCF2, IKZF1, IRF8, IFIH1, and TYK2 with systemic lupus erythematosus. PLoS Genet. 7, e1002341 (2011).
Plenge, R.M. et al. TRAF1-C5 as a risk locus for rheumatoid arthritis—a genomewide study. N. Engl. J. Med. 357, 1199–1209 (2007).
Gregersen, P.K. et al. REL, encoding a member of the NF-κB family of transcription factors, is a newly defined risk locus for rheumatoid arthritis. Nat. Genet. 41, 820–823 (2009).
Wetzels, J.F.M., Kiemeney, L.A.L.M., Swinkels, D.W., Willems, H.L. & Heijer, M.d. Age- and gender-specific reference values of estimated GFR in Caucasians: The Nijmegen Biomedical Study. Kidney Int. 72, 632–637 (2007).
Tian, C. et al. European population genetic substructure: further definition of ancestry informative markers for distinguishing among diverse European ethnic groups. Mol. Med. 15, 371–383 (2009).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Suryani, S. et al. Differential expression of CD21 identifies developmentally and functionally distinct subsets of human transitional B cells. Blood 115, 519–529 (2010).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2 − Δ Δ C T method. Methods 25, 402–408 (2001).
Acknowledgements
The authors thank the volunteers who participated in this study; the Genotype and Phenotype (GaP) registry; M. Keogh, M. DeFranco, C. Mason, C. Metz and the Biorepository at the Feinstein Institute for Medical Research (FIMR) for recruiting subjects and collecting samples; H. Borrero for technical assistance; and the Biostatistics Unit of the FIMR and M. Akerman for assistance. This work was supported by US National Institutes of Health (NIH) grant RC2AR059092; The Alliance for Lupus Research, a Kirkland Scholar Award and NIH/National Center for Research Resources (NCRR) grant 5 M01 RR-00079 (L.A.C.); P01 AR49084 and UL1 TR000165 (R.P.K.); and NIH/National Center for Advancing Translational Sciences grant KL2TR000143 and an American College of Rheumatology Physician Scientist Development Award (S.A.C.). The principal investigators of the Nijmegen Biomedical Study are L.A.L.M. Kiemeney, M. den Heijer, A.L.M. Verbeek, D.W. Swinkels and B. Franke. We thank the Lupus Family Registry and Repository (LFRR) investigators, J.B. Harley, K.L. Moser Sivils, M.H. Weisman and D.J. Wallace, funded by N01AR62277 (K.L.M.S. and J.B.H.).
Author information
Authors and Affiliations
Contributions
N.M.-O., B.D. and P.K.G. designed the study. N.M.-O., S.A.C., D.S.C.G., J.F., D.H.B., T.J.V., L.A.C. and A.T.L. performed genetic analysis. N.M.-O., E.M., J.F.K., M.S.K., K.R.S. and T.J.H. performed experiments. E.N. gave the initial insight into Csk. M.J.H.C., B.F., D.W.S., R.R.G., R.P.K., T.J.V., T.W.B., P.M.G. and L.A.C. provided samples. N.M.-O., B.D. and P.K.G. analyzed and interpreted the data and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
T.W.B. and R.R.G. are full-time employees of Genentech.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4 and Supplementary Figures 1 and 2 (PDF 1701 kb)
Rights and permissions
About this article
Cite this article
Manjarrez-Orduño, N., Marasco, E., Chung, S. et al. CSK regulatory polymorphism is associated with systemic lupus erythematosus and influences B-cell signaling and activation. Nat Genet 44, 1227–1230 (2012). https://doi.org/10.1038/ng.2439
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2439
This article is cited by
-
Proteomic analysis identifies subgroups of patients with active systemic lupus erythematosus
Clinical Proteomics (2023)
-
Regulation of activated T cell survival in rheumatic autoimmune diseases
Nature Reviews Rheumatology (2022)
-
SH3-domain mutations selectively disrupt Csk homodimerization or PTPN22 binding
Scientific Reports (2022)
-
Src Family Protein Kinase Controls the Fate of B Cells in Autoimmune Diseases
Inflammation (2021)
-
Gut microbiota promote the inflammatory response in the pathogenesis of systemic lupus erythematosus
Molecular Medicine (2019)