Abstract
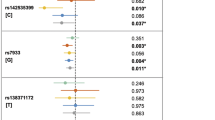
We have conducted the first meta-analyses for nonsyndromic cleft lip with or without cleft palate (NSCL/P) using data from the two largest genome-wide association studies published to date. We confirmed associations with all previously identified loci and identified six additional susceptibility regions (1p36, 2p21, 3p11.1, 8q21.3, 13q31.1 and 15q22). Analysis of phenotypic variability identified the first specific genetic risk factor for NSCLP (nonsyndromic cleft lip plus palate) (rs8001641; PNSCLP = 6.51 × 10−11; homozygote relative risk = 2.41, 95% confidence interval (CI) 1.84–3.16).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Accession codes
Primary accessions
NCBI Reference Sequence
References
Mangold, E., Ludwig, K.U. & Nöthen, M.M. Trends Mol. Med. 17, 725–733 (2011).
Rahimov, F. et al. Nat. Genet. 40, 1341–1347 (2008).
Beaty, T.H. et al. Nat. Genet. 42, 525–529 (2010).
Mangold, E. et al. Nat. Genet. 42, 24–26 (2010).
Mansouri, A., Stoykova, A., Torres, M. & Gruss, P. Development 122, 831–838 (1996).
Sull, J.W. et al. Eur. J. Hum. Genet. 17, 831–839 (2009).
Himanen, J.P., Saha, N. & Nikolov, D.B. Curr. Opin. Cell Biol. 19, 534–542 (2007).
Maunakea, A.K. et al. Nature 466, 253–257 (2010).
Drieschner, N. et al. Gene 403, 110–117 (2007).
Goodnough, L.H., Brugmann, S.A., Hu, D. & Helms, J.A. Dev. Dyn. 236, 1918–1928 (2007).
Welsh, I.C., Hagge-Greenberg, A. & O'Brien, T.P. Mech. Dev. 124, 746–761 (2007).
Jugessur, A., Farlie, P.G. & Kilpatrick, N. Oral Dis. 15, 437–453 (2009).
Grosen, D. et al. J. Med. Genet. 47, 162–168 (2010).
Matsumura, K. et al. Biochem. Biophys. Res. Commun. 404, 1076–1082 (2011).
Vieira, A.R. et al. PLoS Genet. 1, e64 (2005).
Acknowledgements
We thank all affected individuals and their families for their participation in this study, as well as the German support group for people with cleft lip and/or palate (Deutsche Selbsthilfevereinigung für Lippen-Gaumen-Fehlbildungen e.V.). The study was supported by the Deutsche Forschungsgemeinschaft (FOR 423 and individual grants MA 2546/3-1, KR 1912/7-1, NO 246/6-1 and WI 1555/5-1) and the Austrian Cleft Palate Craniofacial Association (ACPCA). T.A. and E.N. are supported by grants from the Ministry of Higher Education, Syrian Arab Republic. The data sets used for the analyses described in this manuscript were obtained from dbGaP under accession phs000094.v1.p1. Additional acknowledgments are provided in the Supplementary Note.
Author information
Authors and Affiliations
Contributions
E.M., F.-J.K., T.F.W., P.P. and M.M.N. initiated the study. E.M., S.N., K.U.L., P.H., M.K., M.R., P.A.M. and M.M.N. contributed to the study design. M.M.N., E.M., S.C., P.H. and K.U.L. coordinated the work. K.U.L., E.M., M.K. and M.M.N. prepared the manuscript, with feedback from the other authors. S.N., H.R., A.P., C. Lauster, B. Braumann, R.H.R., A.H., S.P., B. Blaumeiser, N.D., T.K., R.P.S.-T., F.-J.K., M.R. and P.A.M. clinically characterized the families with cleft lip and collected the blood samples. K.U.L., R.H., E.N., T.A., S.B., A.C.B., N.K., M.A.A. and J.B. prepared the DNA and performed the molecular genetic analyses in the Bonn-II study. J.C.M., M.L.M., I.R., A.F.S. and T.H.B. contributed data from the Baltimore study. M.K., S.H., C. Lange and M.M. conducted the statistical analyses. M.M.N., E.M., M.K., K.U.L., M.R. and P.P. analyzed and interpreted the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ludwig, K., Mangold, E., Herms, S. et al. Genome-wide meta-analyses of nonsyndromic cleft lip with or without cleft palate identify six new risk loci. Nat Genet 44, 968–971 (2012). https://doi.org/10.1038/ng.2360
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2360
This article is cited by
-
Genetic association and functional validation of ZFP36L2 in non-syndromic orofacial cleft subtypes
Journal of Human Genetics (2024)
-
Identification of putative regulatory single-nucleotide variants in NTN1 gene associated with NSCL/P
Journal of Human Genetics (2023)
-
Global perspectives in orofacial cleft management and research
British Dental Journal (2023)
-
Genetic markers for non-syndromic orofacial clefts in populations of European ancestry: a meta-analysis
Scientific Reports (2022)
-
Allele-specific transcription factor binding in a cellular model of orofacial clefting
Scientific Reports (2022)