Abstract
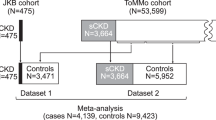
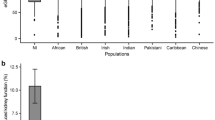
Chronic kidney disease (CKD), impairment of kidney function, is a serious public health problem, and the assessment of genetic factors influencing kidney function has substantial clinical relevance. Here, we report a meta-analysis of genome-wide association studies for kidney function–related traits, including 71,149 east Asian individuals from 18 studies in 11 population-, hospital- or family-based cohorts, conducted as part of the Asian Genetic Epidemiology Network (AGEN). Our meta-analysis identified 17 loci newly associated with kidney function–related traits, including the concentrations of blood urea nitrogen, uric acid and serum creatinine and estimated glomerular filtration rate based on serum creatinine levels (eGFRcrea) (P < 5.0 × 10−8). We further examined these loci with in silico replication in individuals of European ancestry from the KidneyGen, CKDGen and GUGC consortia, including a combined total of ∼110,347 individuals. We identify pleiotropic associations among these loci with kidney function–related traits and risk of CKD. These findings provide new insights into the genetics of kidney function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
National Kidney Foundation. K/DOQI clinical practice guidelines for chronic kidney disease: evaluation, classification, and stratification. Am. J. Kidney Dis. 39, S1–S266 (2002).
Horio, M., Imai, E., Yasuda, Y., Watanabe, T. & Matsuo, S. Modification of the CKD epidemiology collaboration (CKD-EPI) equation for Japanese: accuracy and use for population estimates. Am. J. Kidney Dis. 56, 32–38 (2010).
Fox, C.S. et al. Genome-wide linkage analysis to serum creatinine, GFR, and creatinine clearance in a community-based population: the Framingham Heart Study. J. Am. Soc. Nephrol. 15, 2457–2461 (2004).
Köttgen, A. et al. Multiple loci associated with indices of renal function and chronic kidney disease. Nat. Genet. 41, 712–717 (2009).
Kamatani, Y. et al. Genome-wide association study of hematological and biochemical traits in a Japanese population. Nat. Genet. 42, 210–215 (2010).
Chambers, J.C. et al. Genetic loci influencing kidney function and chronic kidney disease. Nat. Genet. 42, 373–375 (2010).
Köttgen, A. et al. New loci associated with kidney function and chronic kidney disease. Nat. Genet. 42, 376–384 (2010).
Gudbjartsson, D.F. et al. Association of variants at UMOD with chronic kidney disease and kidney stones—role of age and comorbid diseases. PLoS Genet. 6, e1001039 (2010).
Kim, Y.J. et al. Large-scale genome-wide association studies in east Asians identify new genetic loci influencing metabolic traits. Nat. Genet. 43, 990–995 (2011).
Döring, A. et al. SLC2A9 influences uric acid concentrations with pronounced sex-specific effects. Nat. Genet. 40, 430–436 (2008).
Vitart, V. et al. SLC2A9 is a newly identified urate transporter influencing serum urate concentration, urate excretion and gout. Nat. Genet. 40, 437–442 (2008).
Kolz, M. et al. Meta-analysis of 28,141 individuals identifies common variants within five new loci that influence uric acid concentrations. PLoS Genet. 5, e1000504 (2009).
Yang, Q. et al. Multiple genetic loci influence serum urate levels and their relationship with gout and cardiovascular disease risk factors. Circ. Cardiovasc. Genet. 3, 523–530 (2010).
Kato, N. et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat. Genet. 43, 531–538 (2011).
Okada, Y. et al. Common variants at CDKAL1 and KLF9 are associated with body mass index in east Asian populations. Nat. Genet. 44, 302–306 (2012).
Wen, W. et al. Meta-analysis identifies common variants associated with body mass index in east Asians. Nat. Genet. 44, 307–311 (2012).
Lyman, J.L. Blood urea nitrogen and creatinine. Emerg. Med. Clin. North Am. 4, 223–233 (1986).
Okada, Y. et al. Genome-wide association study for C-reactive protein levels identified pleiotropic associations in the IL6 locus. Hum. Mol. Genet. 20, 1224–1231 (2011).
LaMarca, M.E. et al. A novel alteration in metaxin 1, F202L, is associated with N370S in Gaucher disease. J. Hum. Genet. 49, 220–222 (2004).
Tong, G.X. et al. Expression of PAX8 in normal and neoplastic renal tissues: an immunohistochemical study. Mod. Pathol. 22, 1218–1227 (2009).
Kataoka, K. et al. Evi1 is essential for hematopoietic stem cell self-renewal, and its expression marks hematopoietic cells with long-term multilineage repopulating activity. J. Exp. Med. 208, 2403–2416 (2011).
MHC Sequencing Consortium. Complete sequence and gene map of a human major histocompatibility complex. Nature 401, 921–923 (1999).
de Bakker, P.I. et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 38, 1166–1172 (2006).
Mansouri, A. et al. Paired-related murine homeobox gene expressed in the developing sclerotome, kidney, and nervous system. Dev. Dyn. 210, 53–65 (1997).
Reiner, O. et al. The evolving doublecortin (DCX) superfamily. BMC Genomics 7, 188 (2006).
Meyer, T.E. et al. Genome-wide association studies of serum magnesium, potassium, and sodium concentrations identify six loci influencing serum magnesium levels. PLoS Genet. 6, e1001045 (2010).
Sahali, D. et al. A novel approach to investigation of the pathogenesis of active minimal-change nephrotic syndrome using subtracted cDNA library screening. J. Am. Soc. Nephrol. 13, 1238–1247 (2002).
Vanita, V., Singh, D., Robinson, P.N., Sperling, K. & Singh, J.R. A novel mutation in the DNA-binding domain of MAF at 16q23.1 associated with autosomal dominant “cerulean cataract” in an Indian family. Am. J. Med. Genet. A 140, 558–566 (2006).
Ehret, G.B. et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 478, 103–109 (2011).
Okada, Y. et al. Identification of nine novel loci associated with white blood cell subtypes in a Japanese population. PLoS Genet. 7, e1002067 (2011).
Okada, Y. et al. Meta-analysis identifies nine new loci associated with rheumatoid arthritis in the Japanese population. Nat. Genet. 44, 511–516 (2012).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Nakamura, Y. The BioBank Japan Project. Clin. Adv. Hematol. Oncol. 5, 696–697 (2007).
Acknowledgements
The authors acknowledge the essential roles of AGEN in developing the study. BBJ was supported by the Ministry of Education, Culture, Sports, Science and Technology, Japan (MEXT). SP2 was funded by grants from the Biomedical Research Council of Singapore (BMRC 05/1/36/19/413 and 03/1/27/18/216) and the National Medical Research Council of Singapore (NMRC/1174/2008). SiMES was funded by the National Medical Research Council of Singapore (NMRC 0796/2003, IRG07nov013 and NMRC/STaR/0003/2008) and the Biomedical Research Council of Singapore (BMRC 09/1/35/19/616). SINDI and SCES were funded by grants from the Biomedical Research Council of Singapore (BMRC 09/1/35/19/616 and BMRC 08/1/35/19/550) and the National Medical Research Council of Singapore (NMRC/STaR/0003/2008). Y.-Y.T. acknowledges support from the Singapore National Research Foundation (NRF-RF-2010-05). E.-S.T. receives support from the National Medical Research Council of Singapore through a Clinician Scientist Award. We thank the Singapore BioBank and the Genome Institute of Singapore, Agency for Science, Technology and Research, Singapore, for providing services for tissue archiving and genotyping, respectively. KARE was supported by grants from the Korea Centers for Disease Control and Prevention (4845-301, 4851-302 and 4851-307) and an intramural grant from the Korea National Institute of Health (2010-N73002-00). Y.S.C. acknowledges support from a National Research Foundation of Korea (NRF) grant funded by the Korean government (MEST) (2012R1A2A1A03006155). TWSC and TWT2D were supported by the Academia Sinica Genomic Medicine Multicenter Study (40-05-GMM). We acknowledge the National Center for Genome Medicine (NSC100-2319-B-001-001), the National Core Facility Program for Biotechnology of the National Science Council, Taiwan, for technical help in sample management and genotyping. GenSalt was supported by grants (U01HL072507, R01HL087263 and R01HL090682) from the National Heart, Lung, and Blood Institute, the US National Institutes of Health. CAGE was supported by grants for Core Research for Evolutional Science and Technology (CREST) from the Japan Science Technology Agency; the Program for Promotion of Fundamental Studies in Health Sciences, the National Institute of Biomedical Innovation Organization (NIBIO) and the grant of National Center for Global Health and Medicine (NCGM). We thank all the people who supported the Hospital-based Cohort Study at NCGM and the Amagasaki Study. We thank A. Taniguchi, H. Rakugi, K. Sugimoto, K. Kamide and C. Makibayashi for supporting the study.
Author information
Authors and Affiliations
Consortia
Contributions
Y.O. and T. Tanaka designed the overall study. Y.O., X.S., M.J.G., C.-H.C., D.G., F.T. and P.C. analyzed GWAS data. Y.O. performed meta-analysis and other statistical analysis. Y.O., A.T., S.M., T. Tsunoda, K.Y., M.K., Y.N., N. Kamatani and T. Tanaka managed GWAS data of BBJ. X.S., P.C., S.-C.L., T.-Y.W., J.L., T.L.Y., T.A., M.S., Y.-Y.T. and E.-S.T. managed the GWAS data from SP2, SiMES, SINDI and SCES. M.J.G., Y.J.K., J.-Y.L., B.-G.H., D.K. and Y.S.C. managed the GWAS data from KARE and HEXA. C.-H.C., F.-J.T., L.-C.C., S.-J.C.F., Y.-T.C. and J.-Y.W. managed the GWAS data from TWSC and TWT2D. D.G., H.M., D.C.R., J.E.H., S.C. and J.H. managed the GWAS data from GenSalt. F.T., T.K., M.I., T.O. and N. Kato managed the GWAS data from CAGE. J.C.C., W.Z. and J.S.K. managed the data from the KidneyGen Consortium. E.A. managed the data from the GUGC consortium. Y.O., T. Tanaka, E.-S.T., Y.S.C., J.-Y.W., J.H. and N. Kato directed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of contributing members and affiliations is provided in the Supplementary Note.
A full list of contributing members and affiliations is provided in the Supplementary Note.
A full list of contributing members and affiliations is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–8, Supplementary Figures 1 and 2 and Supplementary Note (PDF 1512 kb)
Rights and permissions
About this article
Cite this article
Okada, Y., Sim, X., Go, M. et al. Meta-analysis identifies multiple loci associated with kidney function–related traits in east Asian populations. Nat Genet 44, 904–909 (2012). https://doi.org/10.1038/ng.2352
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2352
This article is cited by
-
Mother–child correlation of kidney function: data from the Yamanashi Adjunct Study of Japan Environment and Children’s Study (JECS)
Pediatric Nephrology (2024)
-
Identification of genetic loci associated with renal dysfunction after lung transplantation using an ethnic-specific single-nucleotide polymorphism array
Scientific Reports (2023)
-
Association of MPPED2 gene variant rs10767873 with kidney function and risk of cardiovascular disease in patients with hypertension
Journal of Human Genetics (2023)
-
Imputation-powered whole-exome analysis identifies genes associated with kidney function and disease in the UK Biobank
Nature Communications (2023)
-
Polygenic risk score trend and new variants on chromosome 1 are associated with male gout in genome-wide association study
Arthritis Research & Therapy (2022)