Abstract
Domestic yaks (Bos grunniens) provide meat and other necessities for Tibetans living at high altitude on the Qinghai-Tibetan Plateau and in adjacent regions. Comparison between yak and the closely related low-altitude cattle (Bos taurus) is informative in studying animal adaptation to high altitude. Here, we present the draft genome sequence of a female domestic yak generated using Illumina-based technology at 65-fold coverage. Genomic comparisons between yak and cattle identify an expansion in yak of gene families related to sensory perception and energy metabolism, as well as an enrichment of protein domains involved in sensing the extracellular environment and hypoxic stress. Positively selected and rapidly evolving genes in the yak lineage are also found to be significantly enriched in functional categories and pathways related to hypoxia and nutrition metabolism. These findings may have important implications for understanding adaptation to high altitude in other animal species and for hypoxia-related diseases in humans.
Similar content being viewed by others
Main
The yak (B. grunniens) is an iconic symbol of Tibet and of high altitude. More than 14 million domestic yaks provide the basic resources (such as meat, milk, transportation, dung for fuel and hides for tented accommodation) that are necessary for Tibetans and other nomadic pastoralists in high-altitude environments1. In contrast, closely related and cross-fertile taurine cattle (B. taurus) suffer from severe pulmonary hypertension when reared in the yak habitat2,3,4. Yaks have numerous anatomical and physiological traits that equip them for life at high altitude, including large lungs and hearts1, lack of hypoxic pulmonary vasoconstriction5, increased foraging ability6, strong environmental sense1 and high energy metabolism1,7. Thus, comparing yak and cattle contributes to the understanding of evolutionary adaptation to high altitude5,6,7.
Genomic comparisons between closely related species provide insights into the genetic basis of mammalian divergence and adaptation8,9. In this study, we sequenced the genome of a female domestic yak using a whole-genome shotgun strategy and the Illumina HiSeq 2000 platform. De novo assembly of 4.4 billion reads from paired-end libraries (Supplementary Table 1) yielded a draft assembly (65-fold coverage) with a total length of 2,657 Mb, close to the 2,649 Mb of sequence obtained for the cattle genome10 (UMD 3.1), and contig and scaffold N50 sizes of 20.4 kb and 1.4 Mb, respectively (Supplementary Table 2). Approximately 90% of the total sequence was covered by 2,083 scaffolds of >307 kb, with the largest scaffold spanning 8.8 Mb. The assembly metrics of the yak genome were comparable to those of other animal genome assemblies generated by next-generation sequencing technology (Supplementary Table 3). The sequencing depth of 98% of the assembly was more than 20-fold (Supplementary Fig. 1), ensuring high accuracy at the nucleotide level11. The genome assembly covered, without any obvious errors in assembly, 97% of 6 fosmids sequenced by Sanger sequencing and 98% of 81,020 Unigenes assembled from Illumina RNA sequencing (RNA-seq) data for 5 tissues (Supplementary Fig. 2 and Supplementary Tables 4 and 5), indicating coverage of most of the euchromatic regions. GC content distributions were similar to those of the cattle genome (Supplementary Fig. 3 and Supplementary Note). We predicted 22,282 protein-coding genes in yak on the basis of RNA-seq, homology and ab initio gene prediction (Supplementary Fig. 4 and Supplementary Table 6).
High coverage11,12 permitted the identification of 2.2 million heterozygous single-nucleotide variants (SNVs) within the sequenced individual (Supplementary Figs. 5 and 6 and Supplementary Tables 7 and 8). The heterozygosity rate (0.89 × 10−3) was approximately 1.5 times higher than that estimated for cattle (0.59 × 10−3) (refs. 10,13). This may be due to a longer and more systematic selection in cattle and/or to introgression from wild yaks still living on the Qinghai-Tibetan Plateau14,15. Yak and cattle both have 30 chromosomes and have similar karyotypes1, which allows chromosomal assignments on the basis of homology, despite a lack of physical maps for yak chromosomes. Using the human genome as an outgroup, we reconstructed 207 conserved ancestral homologous synteny blocks (aHSBs)16 covering 2.51 Gb (94%) of the yak genome. The existence of these blocks confirms the extensive synteny of the cattle and yak genomes and also allows the detection of breakpoint regions (Supplementary Table 9).
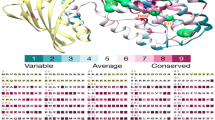
Frequent turnover of gene copy and/or protein domain number has been proposed as a major mechanism underlying the adaptive divergence of closely related species9,17,18,19. First, we used TreeFam20 to identify 13,810 homologous gene families shared by 4 species (yak, cattle, human and dog): 362 gene families were specific to yak and cattle, and 100 were found only in yak (Fig. 1). The sequence depth of these multiple-copy genes was in the same range as for single-copy genes (Supplementary Fig. 7). The yak-specific gene families contained 170 genes, 75 of which have known InterPro domains. Compared with the cattle-specific gene families, the yak-specific families were significantly over-represented in two major functional categories: olfactory sensation (14 genes; P < 0.01) and host defense and immunity (11 genes; P < 0.01) (Supplementary Table 10). Next, we identified 596 gene families that were substantially expanded in yak compared to other mammals (Fig. 2a). Functional categories that were enriched for significant gene family expansions mainly included sensory perception (gene ontology (GO) 0004984, olfactory receptor activity, P < 0.01; GO 0050909, sensory perception of taste, P < 0.01) and energy metabolism (GO 0004129, cytochrome-c oxidase activity, P < 0.01; GO 0015986, ATP synthesis coupled proton transport, P < 0.01) (Supplementary Table 11). Third, matching of ORFs to PFAM domain families at the protein level showed expansion in the numbers of specific domains (Supplementary Table 12). Thus, with regard to sensory perception receptors (olfactory and taste) and other G protein–coupled receptor (GPCR) rhodopsin-like receptors, known to be involved in sensing of the extracellular environment21, we found significantly more GPCR transmembrane domains in yak than in cattle (1,558 versus 1,358). Categories related to hypoxic stress seemed also to have enriched expansions of the corresponding domains in yak. For example, genes with the Hig_1_n (PF04588.6 in Pfam) domain were highly expressed under hypoxic stress22,23 and also had expanded copy numbers in yak (13 copies) relative to cattle (9 copies) and other mammals. Phylogenetic analysis of genes that encoded this domain showed an expansion in the numbers of the closely related Hig_1_n domains in both yak and cattle, as well as three additional copies in yak (Fig. 2b).
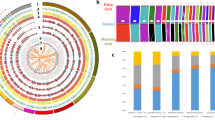
Adaptive divergence at the molecular level may also be expressed by an increased rate of nonsynonymous changes within genes involved in adaptation9,24. We identified 8,923 high-confidence 1:1:1 orthologous genes in the yak, cattle and human genomes, most of which also correspond to genes in the horse, dog, mouse and chimpanzee genomes (Supplementary Fig. 8). Overall, yak and cattle genes were highly similar, with 45% of encoded proteins identical and mean protein similarity approximating 99.5% (Supplementary Figs. 9 and 10). Average synonymous (Ks) and nonsynonymous (Ka) gene divergence values between yak and cattle were 0.0114 and 0.00207, respectively, close to the values between human and chimpanzee genes (Supplementary Fig. 11). Yak and cattle were estimated to have diverged approximately 4.9 million years ago, which is comparable to the time at which humans and chimpanzees diverged9 (Supplementary Fig. 12). Ka/Ks ratios of nonsynonymous-to-synonymous substitutions for different GO categories revealed an enrichment of elevated pairwise Ka/Ks values in the hypoxia response25,26,27 and energy metabolism28,29 categories, including in 'regulation of blood vessel size', 'regulation of angiogenesis', 'heme binding', 'glycerolipid biosynthetic process' and 'electron carrier activity' (Supplementary Table 13). Analysis of Ka/Ks ratios in the cattle and yak lineages verified that genes with elevated Ka/Ks values in yak were significantly enriched for these functions (Fig. 3a).
(a) Data points represent pairs of yak and cattle mean Ka/Ks ratios by GO category. GO categories with putatively accelerated (P < 0.05, binomial test) nonsynonymous divergence in the yak lineage (red) and in the cattle lineage (blue) are highlighted. A complete list of categories is provided in Supplementary Table 17. (b) Comparison of the proportions of genes showing evidence for positive selection in the yak and cattle lineages. The numbers of positively selected genes are given in parentheses. (c) Five genes involved in integrated nutrition pathways (according to KEGG pathway: map04971 and map00020) were found to show evidence of positive selection in the yak lineage. Solid lines indicate direct relationships between enzymes and metabolites. Dashed lines indicate that more than one step is involved in the process.
To test the hypothesis that these rapidly evolving genes in yak have been under positive selection, we used the branch-site likelihood ratio test to identify positively selected genes (PSGs) in both the yak and cattle lineages. We identified 85 PSGs (in yak) and 95 PSGs (in cattle) (Fig. 3b and Supplementary Table 14). The PSGs detected in yak were enriched for genes involved in the hypoxia response and energy metabolism (Supplementary Tables 15 and 16). Of 81 genes examined in the response to hypoxia functional category (GO 0001666), 3 (3.7%) showed evidence of positive selection in yak (compared to none in cattle), which is significantly higher than the background level of positive selection across the genome (P < 0.05) (Fig. 3b). The three yak PSGs comprise two important regulators (Adam17 and Arg2) and one target gene (Mmp3) of hypoxia-inducible factor–1α (Hif-1α). As a master regulator of the cellular response to hypoxia, Hif-1α triggers wide transcription of genes involved in angiogenesis, vasodilatation and energy metabolism30,31,32,33,34,35. Notably, alleles of human ADAM17 were previously shown to be present at significant different frequencies in Tibetans and low-altitude dwellers36, indicating a possible role for this gene in altitude adaptation. The Adam17 and Arg2 proteins affect Hif-1α stability and activity by regulating production of tumor necrosis factor α (TNF-α)31,32 and nitric oxide, respectively33,34, whereas Mmp3 has key roles in numerous physiological processes35. In their high-altitude environments, yaks must not only maintain normal energy production under hypoxic pressure7,37 but must also optimize nutritional assimilation, as a consequence of the limited herbal resources available1. Indeed, we found five key genes that show signs of positive selection in yak nutrition pathways (Fig. 3c): Camk2b regulates the secretion of gastric acid in the rumen, which contributes to the assimilation of volatile fatty acids produced by ruminal fermentation38,39,40,41,42, and Gcnt3, Hsd17b12, Whsc1 and Glul have important roles in polysaccharide, fatty acid and amino-acid metabolism, respectively43,44,45,46. In addition, the positively selected changes in Glul may be important for the high level of nitrogen utilization in yak7.
Our evolutionary analyses based on genomic data have provided important insights into adaptation to high altitude in yak. Further understanding may be gained by functional analysis of the identified genes with signs of adaptive evolution in comparative stress studies of yak and other animals living at high altitude. The identification of genes required for natural high-altitude adaptation may help to improve current understanding, treatment and prevention of altitude sickness and other hypoxia-related diseases in humans. In addition, this report of the yak genome sequence, together with the many SNVs identified, will facilitate genetic dissection of agronomically important traits in the species and will accelerate the genetic improvement of milk and meat production in this animal that is essential to the lifestyle and economy of the Tibetan people.
URLs.
SOAP, http://soap.genomics.org.cn/; Ensembl, http://www.ensembl.org/; TimeTree, http://www.timetree.org/.
Methods
Genome sequencing and assembly.
Genomic DNA was extracted from the liver of a female yak with an estimated inbreeding coefficient of 0.094 (ref. 1) that lived above 3,700 m in Huangyuan County of Qinghai Province, China. Sequencing libraries were constructed with multiple insert sizes (200 bp to 20 kb) according to the Illumina protocol. For short insert libraries (200 to 800 bp), 6 μg of DNA was fragmented to the desired insert size, end-repaired and ligated to Illumina paired-end adaptors. Ligated fragments were size selected at 200, 500 and 800 bp on agarose gels and were purified by PCR amplification to yield the corresponding libraries. For long insert sizes (2, 5, 10 and 20 kb) mate-pair library construction, 60 μg of genomic DNA was used; we circularized DNA, digested linear DNA, fragmented circularized DNA and purified biotinylated DNA and then performed adaptor ligation. All libraries were sequenced on an Illumina HiSeq 2000 platform.
Whole-genome shotgun assembly of the yak was performed using short oligonucleotide analysis package (SOAP)denovo47 (Supplementary Note). First, a series of strict filtering steps were performed before assembly to avoid artifactual duplication, adaptor contamination and low-quality reads. Second, reads from the short-insert (≤800-bp) libraries were assembled into distinct contigs on the basis of k-mer overlap information. Third, reads from the long-insert (≥800-bp) libraries were aligned to the contig sequence, and the paired-end relationships between reads were used to construct scaffolds. We used a hierarchical assembly strategy in which we added data step by step from short paired-end reads to long paired-end reads. Finally, in order to fill gaps between scaffolds, we used the paired-end information to retrieve read pairs that had one read well aligned on the contigs and another read located within the gap region. We then performed a local assembly of the collected reads.
Transcriptome and fosmid sequencing and assembly.
From the same yak animal, RNA was extracted from fresh heart, liver, brain, stomach and lung tissues for the generation of transcriptome data. The quality and integrity of the RNA samples were examined using the Agilent 2100 Bioanalyzer, their RNA integrity number (RIN) values ranged from 8.6–10.0, with no sign of degradation. Approximately 20 μg of total RNA (at a concentration of ≥400 ng/μl) from each tissue was used to construct cDNA libraries. Poly(A) mRNA was isolated using beads conjugated to oligo(dT), mRNA was fragmented, and cDNA was synthesized using random hexamer primers and reverse transcriptase (Invitrogen). After end repair, adaptor ligation and PCR amplification, the libraries were sequenced using the Illumina HiSeq 2000 platform. Libraries that gave reads that were unevenly distributed among the gene regions (for example, showing a strong bias toward 5′ or 3′ regions) were discarded and replaced. Transcripts were assembled using SOAPdenovo. Reads for the assembly were filtered as for the genome assembly, and duplicate reads were removed. For further study, we used only those transcripts that were longer than 150 bp and that were covered at least twice.
We also constructed a fosmid (inset size of ∼40 kb) library from the same DNA resource and randomly selected six clones for Sanger sequencing. These fosmids were assembled using the Celera Assembler48. We then evaluated the completeness and accuracy of the genome assembly by comparing the assembled scaffolds with 6 complete fosmid clones and 81,020 Unigene sequences using BLAST searches.
Heterozygous SNV detection.
To evaluate the heterozygosity rate and its distribution, high-quality reads (average quality score of >30) from short-insert libraries were realigned to the assembly with SOAP (see URLs). The probabilities of each possible genotype at every position on the reference genome were calculated, and a statistical model based on Bayesian theory and the Illumina quality system was used to call SNVs. The allelic sequence with the highest probability was used as the reference sequence, and heterozygous SNVs were called if other alleles also had high probability11. To estimate the accuracy of the identified heterozygous SNVs, we randomly selected 150 heterozygous SNVs and validated them by PCR amplification and Sanger sequencing. In 146 of the 150 sequences, SNVs were validated by double sequence peaks (Supplementary Fig. 6 and Supplementary Table 8).
Annotation.
Transposable elements in the yak genome assembly were first identified using a combination of homology-based and de novo approaches at both the DNA and protein levels47. We then used homology and ab initio prediction, as well as RNA-seq to identify protein-coding genes, building a consensus gene set by merging all predicted genes. For homology-based gene prediction, we aligned human and cattle protein sequences to the repeat-masked yak genome using TBLASTN and Genewise49 for fast alignment and accurate spliced alignment, respectively. Next, we used the ab initio gene prediction methods Genscan50 and Augustus51 to predict protein-coding genes, using parameters trained from a set of high-quality homolog prediction proteins. RNA-seq–based gene prediction was performed by aligning all RNA-seq data against the assembled genome using TopHat52, and Cufflinks53 was used to predict cDNAs from the resultant data. The final gene set was generated by merging these three gene prediction resources by GLEAN54. Gene functions were assigned according to the best match of the alignment to the SwissProt and Translated EMBL Nucleotide Sequence Data Library (TEMBL) databases, using BLASTP. For yak reference genes, motifs and domains were determined by searches in InterProScan55 of the sequences against publicly available databases, including Pfam, PRINTS, PROSITE, ProDom and SMART. Gene Ontology56 IDs for each gene were obtained from the corresponding InterPro entry. We also mapped yak reference genes to KEGG57 pathway databases and identified the best match for each gene.
Gene families.
The protein-coding genes from 6 mammalian species (Canis familiaris, Homo sapiens, Mus musculus, B. taurus, Equus caballus and Monodelphis domestica) downloaded from Ensembl (release 56; see URLs) were used in addition to yak genes to define gene families that descended from a single gene in the last common ancestor20. The longest translation form was chosen to represent each gene, and stretches of genes encoding fewer than 30 amino acids were filtered out. The 9,393 single-copy families obtained from this analysis were used to reconstruct phylogenies and estimate the times since divergence. Data from fourfold-degenerate sites were extracted from each family and attributed to one 'super gene' for each species. Modeltest58 was used to select the best substitution model (GTR + gamma + I), and Mrbayes59 was used to reconstruct the phylogenetic tree. The MCMCtree program implemented in the Phylogenetic Analysis by Maximum Likelihood (PAML)60 package was used to estimate the time since cattle-yak divergence. Calibration time was obtained from the TimeTree database (see URLs). To identify gene families that had undergone expansion or contraction, we applied the Café program, which is based on a probabilistic graphical model61, to infer the rate and direction of change in gene family size over a given phylogeny.
Evolutionary analyses.
We used conserved genome synteny methodology62 to establish a high-confidence orthologous gene set that included yak, cattle (UMD 3.1), horse (EquCab2.0), dog (CanFam2.0), mouse (mm9), chimpanzee (panTro2) and human (hg19) genes. Briefly, whole-genome multiple alignments were constructed for the relevant genomes using the MULTIZ63 alignment pipeline, with the human genome serving as the reference genome. To minimize the effect of annotation errors, variations in sequence quality and changes in gene structure on subsequent evolutionary rate analyses, we mapped all the human protein-coding genes from RefSeq64, KnownGene65 and VEGA66 to each of the other species via their syntenic alignments, then passed the resulting blocks through a series of rigorous filters that selected for large-scale synteny, high alignment quality and conservation of exon-intron structure. All orthologs were aligned using the codon option in the Probabilistic Alignment Kit (PRANK)67 program, and alignments shorter than 150 bp were discarded. The values of Ka and Ks and the Ka/Ks ratio were estimated for each gene using the Codeml program with the free-ratio model in the PAML package, and 10,000 concatenated alignments constructed from 150 randomly chosen genes were used to estimate lineage-specific mean values. The human GO annotation download from Ensembl was used to assign GO categories to 8,923 orthologs. The binomial test9 was used to identify GO categories with more than 20 orthologs that had an excess of nonsynonymous changes in either yak or cattle lineages. To detect genes evolving under positive selection in either yak or cattle (Supplementary Table 17), we used the optimized branch-site model68 in which likelihood ratio test (LRT) P values were computed assuming that the null distribution was a 50:50 mixture of a chi-squared distribution with 1 degree of freedom and a point mass at zero. Fisher's exact tests were used to test for over-represented functional categories among positively selected genes8. For each category C and set of PSGs S, a 2 × 2 contingency table was constructed for the numbers of genes assigned or not assigned to C and within or outside S. Then, (one-sided) P values for the independence of rows and columns were computed by Fisher's exact test. In addition, the distributions of LRT P values among the genes assigned and not assigned to C were compared by a (one-sided) Mann-Whitney U test. To test whether the unique mutations in yak, which resulted in the detected signal of positive evolution in the yak lineage, were specific to yak, we amplified and sequenced the DNA fragments encompassing the candidate yak-specific mutations in 15 genes (including the 8 genes shown in Fig. 3b,c) in 5 randomly selected yaks and 5 cattle. All mutations were confirmed to be specific to yak.
Accession codes.
The yak whole-genome shotgun project has been deposited at the DNA Data Bank of Japan (DDBJ), the European Molecular Biology Laboratory (EMBL) nucleotide sequencing database and GenBank under the same accession, AGSK00000000. The version of the genome described in this paper is the first version, AGSK01000000 (available at DDBJ, EMBL and GenBank). The mitochondrial sequence has been deposited at GenBank under accession JQ692071. All short-read data have been deposited at the Short Read Archive (SRA) under accession SRA047288. Raw sequencing data for the transcriptome have been deposited in the Gene Expression Omnibus (GEO) under accession GSE33300.
Accession codes
Primary accessions
DDBJ/GenBank/EMBL
EMBL/GenBank/DDBJ
Gene Expression Omnibus
NCBI Reference Sequence
Sequence Read Archive
References
Wiener, G., Han, J.L. & Long, R.J. The Yak 2nd edn. (Regional Office for Asia and the Pacific Food and Agriculture Organization of the United Nations, Bangkok, 2003).
Will, D.H., Hicks, J.L., Card, C.S. & Alexander, A.F. Inherited susceptibility of cattle to high-altitude pulmonary hypertension. J. Appl. Physiol. 38, 491–494 (1975).
Weir, E.K., Tucker, A., Reeves, J.T., Will, D.H. & Grover, R.F. The genetic factor influencing pulmonary hypertension in cattle at high altitude. Cardiovasc. Res. 8, 745–749 (1974).
Hecht, H.H., Kuida, H., Lange, R.L., Horne, J.L. & Brown, A.M. Brisket disease. III. Clinical features and hemodynamic observations in altitude-dependent right heart failure of cattle. Am. J. Med. 32, 171–183 (1962).
Dolt, K.S. et al. cDNA cloning, gene organization and variant specific expression of HIF-1α in high altitude yak (Bos grunniens). Gene 386, 73–80 (2007).
Shao, B. et al. Morphological adaptations of yak (Bos grunniens) tongue to the foraging environment of the Qinghai-Tibetan Plateau. J. Anim. Sci. 88, 2594–2603 (2010).
Wang, H. et al. Comparison of nitrogen metabolism in Yak (Bos grunniens) and indigenous cattle (Bos taurus) on the Qinghai-Tibetan Plateau. Asian-Aust. J. Anim. Sci. 24, 766–773 (2011).
Kosiol, C. et al. Patterns of positive selection in six mammalian genomes. PLoS Genet. 4, e1000144 (2008).
Chimpanzee Sequencing and Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87 (2005).
Elsik, C.G. et al. The genome sequence of taurine cattle: a window to ruminant biology and evolution. Science 324, 522–528 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Wade, C.M. et al. Genome sequence, comparative analysis, and population genetics of the domestic horse. Science 326, 865–867 (2009).
Wang, Z. et al. Phylogeographical analyses of domestic and wild yaks based on mitochondrial DNA: new data and reappraisal. J. Biogeogr. 37, 2332–2344 (2010).
Guo, S. et al. Origin of mitochondrial DNA diversity of domestic yaks. BMC Evol. Biol. 6, 73 (2006).
Larkin, D.M. et al. Breakpoint regions and homologous synteny blocks in chromosomes have different evolutionary histories. Genome Res. 19, 770–777 (2009).
Sudmant, P.H. et al. Diversity of human copy number variation and multicopy genes. Science 330, 641–646 (2010).
Dassanayake, M. et al. Transcription strength and halophytic lifestyle. Trends Plant Sci. 16, 1–3 (2011).
Dassanayake, M. et al. The genome of the extremophile crucifer Thellungiella parvula. Nat. Genet. 43, 913–918 (2011).
Li, H. et al. TreeFam: a curated database of phylogenetic trees of animal gene families. Nucleic Acids Res. 34, D572–D580 (2006).
Vassilatis, D.K. et al. The G protein–coupled receptor repertoires of human and mouse. Proc. Natl. Acad. Sci. USA 100, 4903–4908 (2003).
Gracey, A.Y., Troll, J.V. & Somero, G.N. Hypoxia-induced gene expression profiling in the euryoxic fish Gillichthys mirabilis. Proc. Natl. Acad. Sci. USA 98, 1993–1998 (2001).
Salnikow, K. et al. The involvement of hypoxia-inducible transcription factor-1–dependent pathway in nickel carcinogenesis. Cancer Res. 63, 3524–3530 (2003).
Bakewell, M.A., Shi, P. & Zhang, J. More genes underwent positive selection in chimpanzee evolution than in human evolution. Proc. Natl. Acad. Sci. USA 104, 7489–7494 (2007).
Beall, C.M. Two routes to functional adaptation: Tibetan and Andean high-altitude natives. Proc. Natl. Acad. Sci. USA 104 (suppl. 1), 8655–8660 (2007).
Pugh, C.W. & Ratcliffe, P.J. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat. Med. 9, 677–684 (2003).
Storz, J.F. & Moriyama, H. Mechanisms of hemoglobin adaptation to high altitude hypoxia. High Alt. Med. Biol. 9, 148–157 (2008).
Prentki, M. & Madiraju, S.R. Glycerolipid metabolism and signaling in health and disease. Endocr. Rev. 29, 647–676 (2008).
Das, J. The role of mitochondrial respiration in physiological and evolutionary adaptation. Bioessays 28, 890–901 (2006).
Yee Koh, M., Spivak-Kroizman, T.R. & Powis, G. HIF-1 regulation: not so easy come, easy go. Trends Biochem. Sci. 33, 526–534 (2008).
Srour, N. et al. TACE/ADAM-17 maturation and activation of sheddase activity require proprotein convertase activity. FEBS Lett. 554, 275–283 (2003).
Zhou, J., Schmid, T. & Brune, B. Tumor necrosis factor–α causes accumulation of a ubiquitinated form of hypoxia inducible factor-1α through a nuclear factor-κB–dependent pathway. Mol. Biol. Cell 14, 2216–2225 (2003).
Sogawa, K. et al. Inhibition of hypoxia-inducible factor 1 activity by nitric oxide donors in hypoxia. Proc. Natl. Acad. Sci. USA 95, 7368–7373 (1998).
Sousa, M.S., Latini, F.R., Monteiro, H.P. & Cerutti, J.M. Arginase 2 and nitric oxide synthase: pathways associated with the pathogenesis of thyroid tumors. Free Radic. Biol. Med. 49, 997–1007 (2010).
Rosenberg, G.A. Matrix metalloproteinases and their multiple roles in neurodegenerative diseases. Lancet Neurol. 8, 205–216 (2009).
Simonson, T.S. et al. Genetic evidence for high-altitude adaptation in Tibet. Science 329, 72–75 (2010).
Han, X.T. et al. Effects of high altitude and season on fasting heat production in the yak Bos grunniens or Poephagus grunniens. Br. J. Nutr. 88, 189–197 (2002).
Armentano, L.E. Ruminant hepatic metabolism of volatile fatty acids, lactate and pyruvate. J. Nutr. 122, 838–842 (1992).
Russell, J.B. & Rychlik, J.L. Factors that alter rumen microbial ecology. Science 292, 1119–1122 (2001).
Allen, M.S., Bradford, B.J. & Oba, M. Board invited review: the hepatic oxidation theory of the control of feed intake and its application to ruminants. J. Anim. Sci. 87, 3317–3334 (2009).
Weimer, P.J., Russell, J.B. & Muck, R.E. Lessons from the cow: what the ruminant animal can teach us about consolidated bioprocessing of cellulosic biomass. Bioresour. Technol. 100, 5323–5331 (2009).
Hersey, S.J. & Sachs, G. Gastric acid secretion. Physiol. Rev. 75, 155–189 (1995).
Yeh, J.C., Ong, E. & Fukuda, M. Molecular cloning and expression of a novel β-1, 6-N-acetylglucosaminyltransferase that forms core 2, core 4, and I branches. J. Biol. Chem. 274, 3215–3221 (1999).
Moon, Y.A. & Horton, J.D. Identification of two mammalian reductases involved in the two-carbon fatty acyl elongation cascade. J. Biol. Chem. 278, 7335–7343 (2003).
Li, Y. et al. The target of the NSD family of histone lysine methyltransferases depends on the nature of the substrate. J. Biol. Chem. 284, 34283–34295 (2009).
Conjard, A. et al. Inhibition of glutamine synthetase in the mouse kidney: a novel mechanism of adaptation to metabolic acidosis. J. Biol. Chem. 278, 38159–38166 (2003).
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature 463, 311–317 (2010).
Myers, E.W. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Salamov, A.A. & Solovyev, V.V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19 (suppl. 2), ii215–ii225 (2003).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Elsik, C.G. et al. Creating a honey bee consensus gene set. Genome Biol. 8, R13 (2007).
Mulder, N.J. et al. New developments in the InterPro database. Nucleic Acids Res. 35, D224–D228 (2007).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Posada, D. & Crandall, K.A. MODELTEST: testing the model of DNA substitution. Bioinformatics 14, 817–818 (1998).
Huelsenbeck, J.P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Yang, Z. PAML 4: Phylogenetic Analysis by Maximum Likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
De Bie, T., Cristianini, N., Demuth, J.P. & Hahn, M.W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271 (2006).
Rhesus Macaque Genome Sequencing and Analysis Consortium. Evolutionary and biomedical insights from the rhesus macaque genome. Science 316, 222–234 (2007).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Pruitt, K.D., Tatusova, T. & Maglott, D.R. NCBI Reference Sequence (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 33, D501–D504 (2005).
Hsu, F. et al. The UCSC Known Genes. Bioinformatics 22, 1036–1046 (2006).
Wilming, L.G. et al. The vertebrate genome annotation (Vega) database. Nucleic Acids Res. 36, D753–D760 (2008).
Löytynoja, A. & Goldman, N. An algorithm for progressive multiple alignment of sequences with insertions. Proc. Natl. Acad. Sci. USA 102, 10557–10562 (2005).
Zhang, J., Nielsen, R. & Yang, Z. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol. Biol. Evol. 22, 2472–2479 (2005).
Acknowledgements
The project was supported by the Natural Science Foundation of China (30725004 and 40972018), the Ministry of Science and Technology of China (2010DFB63500), the International Collaboration 111 Projects of China, the 985 and 211 Projects of Lanzhou University, the Shenzhen Municipal Government (ZYC200903240077A) and the Hundreds-Talent Program from the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
J.L. designed and managed the project. Jian Wang and Jun Wang led the genome sequencing. P.S. designed the related analyses of the gene families. Z.W. prepared the yak nucleic acid samples. G.Z., Y.L., W.Q., H,Y., Junyi Wang, X.Q., Y. Wang, X.Z. and Y.Y. performed the DNA sequencing. W.Q., Junyi Wang, S.L., S.P. and C.C. performed the genome assembly. W.Q., C.C., S.L., Yan Zhang, J.X., H.H. and G.Z. performed the genome annotation. Q.Q. and T.M. designed evolutionary analyses. T.M., Q.Q., Q.H., M.H., Yingmei Zhang, R.Z., X.L., L.W., H.M., K.W., D.Z., S.Z., R.L., T.Y., W.Q., C.C., G.Z. and Y. Wu performed evolutionary analyses. Z.Y. and P.S. independently verified all analyses. J.K., D.M.L., L.A., B.C., J.M., H.A.L. and W.L. performed the synteny analyses. Q.H., Q.Q., J.Z. and Z.H. carried out data submission and database construction. Q.Q. and J.L. wrote the paper. H.A.L., Jun Wang, P.S., D.N.C., G.Z., J.A.L., J.M. and Y. Zhong revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–16 and Supplementary Note (PDF 2056 kb)
Supplementary Table 17
Identification of lineage-specific accelerated GO categories (XLS 48 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-ShareAlike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Qiu, Q., Zhang, G., Ma, T. et al. The yak genome and adaptation to life at high altitude. Nat Genet 44, 946–949 (2012). https://doi.org/10.1038/ng.2343
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2343
This article is cited by
-
Whole-genome resequencing of major populations revealed domestication-related genes in yaks
BMC Genomics (2024)
-
Whole-genome resequencing of Chinese indigenous sheep provides insight into the genetic basis underlying climate adaptation
Genetics Selection Evolution (2024)
-
Chromosome-scale genome assembly of Lepus oiostolus (Lepus, Leporidae)
Scientific Data (2024)
-
The mitochondrial genome of the mountain wooly tapir, Tapirus pinchaque and a formal test of the effect of altitude on the adaptive evolution of mitochondrial protein coding genes in odd-toed ungulates
BMC Genomics (2023)
-
Genome and whole-genome resequencing of Cinnamomum camphora elucidate its dominance in subtropical urban landscapes
BMC Biology (2023)