Abstract
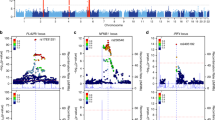
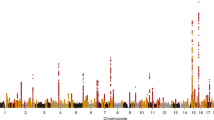
The increased burden of chronic kidney and end-stage kidney diseases (ESKD) in populations of African ancestry has been largely unexplained. To identify genetic variants predisposing to idiopathic and HIV-1–associated focal segmental glomerulosclerosis (FSGS), we carried out an admixture-mapping linkage-disequilibrium genome scan on 190 African American individuals with FSGS and 222 controls. We identified a chromosome 22 region with a genome-wide logarithm of the odds (lod) score of 9.2 and a peak lod of 12.4 centered on MYH9, a functional candidate gene expressed in kidney podocytes. Multiple MYH9 SNPs and haplotypes were recessively associated with FSGS, most strongly a haplotype spanning exons 14 through 23 (OR = 5.0, 95% CI = 3.5–7.1; P = 4 × 10−23, n = 852). This association extended to hypertensive ESKD (OR = 2.2, 95% CI = 1.5–3.4; n = 433), but not type 2 diabetic ESKD (n = 476). Genetic variation at the MYH9 locus substantially explains the increased burden of FSGS and hypertensive ESKD among African Americans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Coresh, J. et al. Prevalence of chronic kidney disease in the United States. J. Am. Med. Assoc. 298, 2038–2047 (2007).
Kiberd, B.A. & Clase, C.M. Cumulative risk for developing end-stage renal disease in the US population. J. Am. Soc. Nephrol. 13, 1635–1644 (2002).
Cowie, C.C. et al. Disparities in incidence of diabetic end-stage renal disease according to race and type of diabetes. N. Engl. J. Med. 321, 1074–1079 (1989).
Toto, R.D. Proteinuria and hypertensive nephrosclerosis in African Americans. Kidney Int. Suppl. 66, S102–S104 (2004).
Fernandez, M. et al. A multiethnic, multicenter cohort of patients with systemic lupus erythematosus (SLE) as a model for the study of ethnic disparities in SLE. Arthritis Rheum. 57, 576–584 (2007).
Kitiyakara, C., Eggers, P. & Kopp, J.B. Twenty-one-year trend in ESRD due to focal segmental glomerulosclerosis in the United States. Am. J. Kidney Dis. 44, 815–825 (2004).
Kopp, J.B. & Winkler, C. HIV-associated nephropathy in African Americans. Kidney Int. Suppl. 63, S43–S49 (2003).
Schwartz, E.J. & Klotman, P.E. Pathogenesis of human immunodeficiency virus (HIV)-associated nephropathy. Semin. Nephrol. 18, 436–445 (1998).
Powe, N.R. To have and have not: health and health care disparities in chronic kidney disease. Kidney Int. 64, 763–772 (2003).
Barisoni, L., Schnaper, H.W. & Kopp, J.B. A proposed taxonomy for the podocytopathies: a reassessment of the primary nephrotic diseases. Clin. J. Am. Soc. Nephrol. 2, 529–542 (2007).
Kitiyakara, C., Kopp, J.B. & Eggers, P. Trends in the epidemiology of focal segmental glomerulosclerosis. Semin. Nephrol. 23, 172–182 (2003).
Eggers, P.W. & Kimmel, P.L. Is there an epidemic of HIV infection in the US ESRD program? J. Am. Soc. Nephrol. 15, 2477–2485 (2004).
Freedman, M.L. et al. Admixture mapping identifies 8q24 as a prostate cancer risk locus in African-American men. Proc. Natl. Acad. Sci. USA 103, 14068–14073 (2006).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Zhu, X. et al. Admixture mapping for hypertension loci with genome-scan markers. Nat. Genet. 37, 177–181 (2005).
Reich, D. et al. A whole-genome admixture scan finds a candidate locus for multiple sclerosis susceptibility. Nat. Genet. 37, 1113–1118 (2005).
Nalls, M.A. et al. Admixture mapping of white cell count: genetic locus responsible for lower white blood cell count in the Health ABC and Jackson Heart studies. Am. J. Hum. Genet. 82, 81–87 (2008).
Reich, D. et al. Admixture mapping of an allele affecting interleukin 6 soluble receptor and interleukin 6 levels. Am. J. Hum. Genet. 80, 716–726 (2007).
Smith, M.W. et al. A high density admixture map for disease gene discovery in African Americans. Am. J. Hum. Genet. 74, 1001–1013 (2004).
Patterson, N. et al. Methods for high-density admixture mapping of disease genes. Am. J. Hum. Genet. 74, 979–1000 (2004).
Zhu, X., Tang, H. & Risch, N. Admixture mapping and the role of population structure for localizing disease genes. Adv. Genet. 60, 547–569 (2008).
Arrondel, C. et al. Expression of the nonmuscle myosin heavy chain IIA in the human kidney and screening for MYH9 mutations in Epstein and Fechtner syndromes. J. Am. Soc. Nephrol. 13, 65–74 (2002).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Price, A.L. et al. Discerning the ancestry of European Americans in genetic association studies. PLoS Genet. 4, e236 (2008).
Wiggins, R.C. The spectrum of podocytopathies: a unifying view of glomerular diseases. Kidney Int. 71, 1205–1214 (2007).
Winn, M.P. Approach to the evaluation of heritable diseases and update on familial focal segmental glomerulosclerosis. Nephrol. Dial. Transplant. 18 (Suppl. 6), vi14–vi20 (2003).
McKenzie, L.M. et al. NPHS2 variation in sporadic focal segmental glomerulosclerosis. J. Am. Soc. Nephrol. 18, 2987–2995 (2007).
Orloff, M.S. et al. Variants in the Wilms' tumor gene are associated with focal segmental glomerulosclerosis in the African American population. Physiol. Genomics 21, 212–221 (2005).
Kao, W.H.L. et al. MYH9 is associated with nondiabetic end-stage renal disease in African Americans. Nat. Genet. advance online publication, doi:10.1038/ng.232 (14 September 2008).
Sellers, J.R. Myosins: a diverse superfamily. Biochim. Biophys. Acta 1496, 3–22 (2000).
Dong, F. et al. Genotype-phenotype correlation in MYH9-related thrombocytopenia. Br. J. Haematol. 130, 620–627 (2005).
Ghiggeri, G.M. et al. Genetics, clinical and pathological features of glomerulonephritis associated with mutations of nonmuscle myosin IIA (Fechtner syndrome). Am. J. Kidney Dis. 41, 95–104 (2003).
Matsushita, T. et al. Targeted disruption of mouse ortholog of the human MYH9 responsible for macrothrombocytopenia with different organ involvement: hematological, nephrological, and otological studies of heterozygous KO mice. Biochem. Biophys. Res. Commun. 325, 1163–1171 (2004).
Kaplan, J.M. et al. Mutations in ACTN4, encoding alpha-actinin-4, cause familial focal segmental glomerulosclerosis. Nat. Genet. 24, 251–256 (2000).
Kim, J.M. et al. CD2-associated protein haploinsufficiency is linked to glomerular disease susceptibility. Science 300, 1298–1300 (2003).
Asanuma, K. et al. Synaptopodin orchestrates actin organization and cell motility via regulation of RhoA signalling. Nat. Cell Biol. 8, 485–491 (2006).
Freedman, B.I., Iskandar, S.S. & Appel, R.G. The link between hypertension and nephrosclerosis. Am. J. Kidney Dis. 25, 207–221 (1995).
Zarif, L. et al. Inaccuracy of clinical phenotyping parameters for hypertensive nephrosclerosis. Nephrol. Dial. Transplant. 15, 1801–1807 (2000).
Fogo, A. et al. Accuracy of the diagnosis of hypertensive nephrosclerosis in African Americans: a report from the African American Study of Kidney Disease (AASK) Trial. AASK Pilot Study Investigators. Kidney Int. 51, 244–252 (1997).
Marcantoni, C., Ma, L.J., Federspiel, C. & Fogo, A.B. Hypertensive nephrosclerosis in African Americans versus Caucasians. Kidney Int. 62, 172–180 (2002).
O'Brien, S.J. & Nelson, G.W. Human genes that limit AIDS. Nat. Genet. 36, 565–574 (2004).
Freedman, B.I. et al. A genome scan for ESRD in black families enriched for nondiabetic nephropathy. J. Am. Soc. Nephrol. 15, 2719–2727 (2004).
Bowden, D.W. et al. A genome scan for diabetic nephropathy in African Americans. Kidney Int. 66, 1517–1526 (2004).
Thorisson, G.A., Smith, A.V., Krishnan, L. & Stein, L.D. The International HapMap Project web site. Genome Res. 15, 1592–1593 (2005).
Reich, D. & Patterson, N. Will admixture mapping work to find disease genes? Phil. Trans. R. Soc. Lond. B 360, 1605–1607 (2005).
Tang, H., Peng, J., Wang, P. & Risch, N.J. Estimation of individual admixture: analytical and study design considerations. Genet. Epidemiol. 28, 289–301 (2005).
Gabriel, S.B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002).
Frazer, K.A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Stephens, M. & Scheet, P. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am. J. Hum. Genet. 76, 449–462 (2005).
Acknowledgements
We thank the subjects and controls who participated in the NIH Genetic Study and the Wake Forest ESRD Study, and the physicians who assisted by identifying cases for these studies. M. Branton, D. Smith, L. Bartlett and L. Howard provided clinical support. E. Binns, Y. Zhou and M. Jamba provided laboratory assistance. The following members of the NIH FSGS Genetics Study Group also contributed samples: F. Hutchinson (University of South Carolina); R. Mannon (Duke University); P. Nachman (University of North Carolina, Chapel Hill); T.K. Rao (State University of New York, Downstate) and M. Smith (Cincinnati Children's Hospital). Research blood donor samples were provided by S. Leitman (NIH). We thank S.J. O'Brien and B. Gold for discussions, N. Patterson for assistance with the ANCESTRYMAP program, and S. Leitman for assistance in identifying healthy volunteers.
The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the US government. The publisher or recipient acknowledges right of the US government to retain a nonexclusive, royalty-free license in and to any copyright covering the article.
This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. This project has been funded in whole or in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract N01-CO-12400, the Intramural Research Programs of the National Institute for Diabetes, Digestive, and Kidney Diseases (ZO-1 DK043308), and by a grant from the NIH (RO1 DK 070941 (B.I.F.)).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The following authors have filed a patent application for MYH9 risk variants and haplotypes: Randy Johnson, Cheryl Winkler, Michael Smith, Jeffrey Kopp and George Nelson.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1 and 2, Supplementary Tables 1–5 (PDF 753 kb)
Rights and permissions
About this article
Cite this article
Kopp, J., Smith, M., Nelson, G. et al. MYH9 is a major-effect risk gene for focal segmental glomerulosclerosis. Nat Genet 40, 1175–1184 (2008). https://doi.org/10.1038/ng.226
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.226
This article is cited by
-
Insights into human kidney function from the study of Drosophila
Pediatric Nephrology (2023)
-
The evolving story of apolipoprotein L1 nephropathy: the end of the beginning
Nature Reviews Nephrology (2022)
-
Apolipoprotein L1 gene variants and kidney disease in patients with HIV: a systematic review and meta-analysis
Journal of Nephrology (2022)
-
The glomerular filtration barrier: a structural target for novel kidney therapies
Nature Reviews Drug Discovery (2021)
-
A focus on the association of Apol1 with kidney disease in children
Pediatric Nephrology (2021)