Abstract
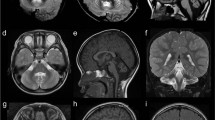
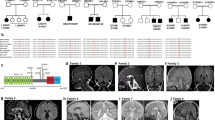
RNA exosomes are multi-subunit complexes conserved throughout evolution1 and are emerging as the major cellular machinery for processing, surveillance and turnover of a diverse spectrum of coding and noncoding RNA substrates essential for viability2. By exome sequencing, we discovered recessive mutations in EXOSC3 (encoding exosome component 3) in four siblings with infantile spinal motor neuron disease, cerebellar atrophy, progressive microcephaly and profound global developmental delay, consistent with pontocerebellar hypoplasia type 1 (PCH1; MIM 607596)3,4,5,6. We identified mutations in EXOSC3 in an additional 8 of 12 families with PCH1. Morpholino knockdown of exosc3 in zebrafish embryos caused embryonic maldevelopment, resulting in small brain size and poor motility, reminiscent of human clinical features, and these defects were largely rescued by co-injection with wild-type but not mutant exosc3 mRNA. These findings represent the first example of an RNA exosome core component gene that is responsible for a human disease and further implicate dysregulation of RNA processing in cerebellar and spinal motor neuron maldevelopment and degeneration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mitchell, P., Petfalski, E., Shevchenko, A., Mann, M. & Tollervey, D. The exosome: a conserved eukaryotic RNA processing complex containing multiple 3′→5′ exoribonucleases. Cell 91, 457–466 (1997).
Jensen, T.H. RNA exosome. Adv. Exp. Med. Biol. 702 (2010).
Norman, R.M. Cerebellar hypoplasia in Werdnig-Hoffmann disease. Arch. Dis. Child. 36, 96–101 (1961).
Goutières, F., Aicardi, J. & Farkas, E. Anterior horn cell disease associated with pontocerebellar hypoplasia in infants. J. Neurol. Neurosurg. Psychiatry 40, 370–378 (1977).
de Leén, G.A., Grover, W.D. & D'Cruz, C.A. Amyotrophic cerebellar hypoplasia: a specific form of infantile spinal atrophy. Acta Neuropathol. 63, 282–286 (1984).
Barth, P.G. Pontocerebellar hypoplasias. An overview of a group of inherited neurodegenerative disorders with fetal onset. Brain Dev. 15, 411–422 (1993).
Namavar, Y., Barth, P.G., Poll-The, B.T. & Baas, F. Classification, diagnosis and potential mechanisms in pontocerebellar hypoplasia. Orphanet J. Rare Dis. 6, 50 (2011).
Melki, J. et al. De novo and inherited deletions of the 5q13 region in spinal muscular atrophies. Science 264, 1474–1477 (1994).
Görgen-Pauly, U., Sperner, J., Reiss, I., Gehl, H.B. & Reusche, E. Familial pontocerebellar hypoplasia type I with anterior horn cell disease. Eur. J. Paediatr. Neurol. 3, 33–38 (1999).
Muntoni, F. et al. Clinical spectrum and diagnostic difficulties of infantile ponto-cerebellar hypoplasia type 1. Neuropediatrics 30, 243–248 (1999).
Ryan, M.M., Cooke-Yarborough, C.M., Procopis, P.G. & Ouvrier, R.A. Anterior horn cell disease and olivopontocerebellar hypoplasia. Pediatr. Neurol. 23, 180–184 (2000).
Rudnik-Schöneborn, S. et al. Extended phenotype of pontocerebellar hypoplasia with infantile spinal muscular atrophy. Am. J. Med. Genet. A 117A, 10–17 (2003).
Chou, S.M. et al. Infantile olivopontocerebellar atrophy with spinal muscular atrophy (infantile OPCA + SMA). Clin. Neuropathol. 9, 21–32 (1990).
Salman, M.S. et al. Pontocerebellar hypoplasia type 1: new leads for an earlier diagnosis. J. Child Neurol. 18, 220–225 (2003).
Gómez-Lado, C., Eiris-Punal, J., Vazquez-Lopez, M.E. & Castro-Gago, M. Pontocerebellar hypoplasia type I and mitochondrial pathology. Rev. Neurol. 45, 639–640 (2007).
Lev, D. et al. Infantile onset progressive cerebellar atrophy and anterior horn cell degeneration—a late onset variant of PCH-1? Eur. J. Paediatr. Neurol. 12, 97–101 (2008).
Szabó, N., Szabo, H., Hortobagyi, T., Turi, S. & Sztriha, L. Pontocerebellar hypoplasia type 1. Pediatr. Neurol. 39, 286–288 (2008).
Tsao, C.Y., Mendell, J., Sahenk, Z., Rusin, J. & Boue, D. Hypotonia, weakness, and pontocerebellar hypoplasia in siblings. Semin. Pediatr. Neurol. 15, 151–153 (2008).
Sanefuji, M. et al. Autopsy case of later-onset pontocerebellar hypoplasia type 1: pontine atrophy and pyramidal tract involvement. J. Child Neurol. 25, 1429–1434 (2010).
Renbaum, P. et al. Spinal muscular atrophy with pontocerebellar hypoplasia is caused by a mutation in the VRK1 gene. Am. J. Hum. Genet. 85, 281–289 (2009).
Namavar, Y. et al. Clinical, neuroradiological and genetic findings in pontocerebellar hypoplasia. Brain 134, 143–156 (2011).
Simonati, A., Cassandrini, D., Bazan, D. & Santorelli, F.M. TSEN54 mutation in a child with pontocerebellar hypoplasia type 1. Acta Neuropathol. 121, 671–673 (2011).
Budde, B.S. et al. tRNA splicing endonuclease mutations cause pontocerebellar hypoplasia. Nat. Genet. 40, 1113–1118 (2008).
Edvardson, S. et al. Deleterious mutation in the mitochondrial arginyl-transfer RNA synthetase gene is associated with pontocerebellar hypoplasia. Am. J. Hum. Genet. 81, 857–862 (2007).
Rankin, J. et al. Pontocerebellar hypoplasia type 6: a British case with PEHO-like features. Am. J. Med. Genet. A 152A, 2079–2084 (2010).
Brouwer, R. et al. Three novel components of the human exosome. J. Biol. Chem. 276, 6177–6184 (2001).
Liu, Q., Greimann, J.C. & Lima, C.D. Reconstitution, activities, and structure of the eukaryotic RNA exosome. Cell 127, 1223–1237 (2006).
Kani, S. et al. Proneural gene–linked neurogenesis in zebrafish cerebellum. Dev. Biol. 343, 1–17 (2010).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Wolfe, J.F., Adelstein, E. & Sharp, G.C. Antinuclear antibody with distinct specificity for polymyositis. J. Clin. Invest. 59, 176–178 (1977).
Allmang, C. et al. The yeast exosome and human PM-Scl are related complexes of 3′→5′ exonucleases. Genes Dev. 13, 2148–2158 (1999).
Yang, X.F. et al. CML28 is a broadly immunogenic antigen, which is overexpressed in tumor cells. Cancer Res. 62, 5517–5522 (2002).
Xie, L.H. et al. Activation of cytotoxic T lymphocytes against CML28-bearing tumors by dendritic cells transduced with a recombinant adeno-associated virus encoding the CML28 gene. Cancer Immunol. Immunother. 57, 1029–1038 (2008).
Kabashi, E. et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat. Genet. 40, 572–574 (2008).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Kwiatkowski, T.J. Jr. et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009).
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p–linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A.E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21–linked ALS-FTD. Neuron 72, 257–268 (2011).
Kobayashi, H. et al. Expansion of intronic GGCCTG hexanucleotide repeat in NOP56 causes SCA36, a type of spinocerebellar ataxia accompanied by motor neuron involvement. Am. J. Hum. Genet. 89, 121–130 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Biesecker, L.G. et al. The ClinSeq Project: piloting large-scale genome sequencing for research in genomic medicine. Genome Res. 19, 1665–1674 (2009).
Bhagwat, M. Searching NCBI's dbSNP database. Curr. Protoc. Bioinformatics Chapter 1, Unit 1.19 (2010).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science 322, 1851–1854 (2008).
Acknowledgements
The authors are grateful for the generosity of the families who participated in this study. The sequencing and analytical work were supported by the Bioinformatics and Genomics Core of the UCLA Muscular Dystrophy Core Center (US National Institutes of Health (NIH)/National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) P30AR057230) within the Center for Duchenne Muscular Dystrophy. We appreciate fibroblasts from three subjects from the Eunice Kennedy Shriver National Institute of Child Health & Human Development (NICHD) Brain and Tissue Bank for Developmental Disorders. We acknowledge the contribution of clinical data by P. Carpenter (University of California, Irvine). The work was supported by the NIH/National Eye Institute (NEI) (R01 EY015311 and NINDS R01 NS064183 to J.C.J.), Deutsche Forschungsgemeinschaft (Ru-746/1-1 to S.R.-S. and K.Z.), IZKF Aachen (N5-4 to S.R.-S. and J.S.), the Australian National Health and Medical Research Council (NHMRC) Centre for Research Excellence (M.M.R.), the Internal Grant Agency of the Ministry of Health of the Czech Republic (NS 10552-3 to P.S.) and Xunta de Galicia–Plan Galego de InvestigaCión, Desenvolvemento e Innovación Tecnolóxica (INCITE) (10PXIB9101280PR to M.-J.S.).
Author information
Authors and Affiliations
Contributions
S.F.N. and J.C.J. designed the study. M.Y., Z.C. and S.F.N. analyzed data from SNP genotyping and exome sequencing. S.R.-S., M.P.M., J.S., M.S.S., D.C., P.S., A.v.M., L.G.-N., A.J.K., M.C.-G., M.-J.S., M.S., P.B.S., N.S., R.C.K., H.V.V., K.Z. and M.M.R. provided and analyzed clinical material from subjects. H.M., J.E.H. and D.W.L. performed and analyzed data from Sanger sequencing of subjects and controls. H.M. and J.E.H. generated constructs and performed molecular genetics studies. J.W. performed and analyzed the functional studies in zebrafish. All authors contributed to the manuscript written by J.C.J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–5 and Supplementary Figures 1–4 (PDF 942 kb)
Rights and permissions
About this article
Cite this article
Wan, J., Yourshaw, M., Mamsa, H. et al. Mutations in the RNA exosome component gene EXOSC3 cause pontocerebellar hypoplasia and spinal motor neuron degeneration. Nat Genet 44, 704–708 (2012). https://doi.org/10.1038/ng.2254
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2254
This article is cited by
-
A missense variant in EXOSC8 causes exon skipping and expands the phenotypic spectrum of pontocerebellar hypoplasia type 1C
Journal of Human Genetics (2024)
-
Comprehensive whole-genome sequence analyses provide insights into the genomic architecture of cerebral palsy
Nature Genetics (2024)
-
A non-coding variant in the Kozak sequence of RARS2 strongly decreases protein levels and causes pontocerebellar hypoplasia
BMC Medical Genomics (2023)
-
Novel compound heterozygous variant of TOE1 results in a mild type of pontocerebellar hypoplasia type 7: an expansion of the clinical phenotype
neurogenetics (2022)
-
Diagnostic Approach to Cerebellar Hypoplasia
The Cerebellum (2021)