Abstract
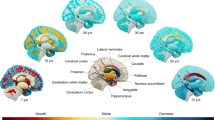
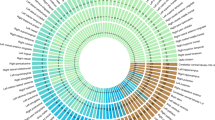
During aging, intracranial volume remains unchanged and represents maximally attained brain size, while various interacting biological phenomena lead to brain volume loss. Consequently, intracranial volume and brain volume in late life reflect different genetic influences. Our genome-wide association study (GWAS) in 8,175 community-dwelling elderly persons did not reveal any associations at genome-wide significance (P < 5 × 10−8) for brain volume. In contrast, intracranial volume was significantly associated with two loci: rs4273712 (P = 3.4 × 10−11), a known height-associated locus on chromosome 6q22, and rs9915547 (P = 1.5 × 10−12), localized to the inversion on chromosome 17q21. We replicated the associations of these loci with intracranial volume in a separate sample of 1,752 elderly persons (P = 1.1 × 10−3 for 6q22 and 1.2 × 10−3 for 17q21). Furthermore, we also found suggestive associations of the 17q21 locus with head circumference in 10,768 children (mean age of 14.5 months). Our data identify two loci associated with head size, with the inversion at 17q21 also likely to be involved in attaining maximal brain size.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
27 April 2012
In the version of this article initially published online, the percentage of females in the AGES study in Table 1 was left blank. This number should be 59%. The error has been corrected for the PDF and HTML versions of this article.
08 May 2013
In the version of this article initially published, Thorkild I.A. Sørensen was listed incorrectly as a contributing member of the EGG Consortium. The error has been corrected for the HTML and PDF versions of this article.
References
Gale, C.R., O'Callaghan, F.J., Bredow, M. & Martyn, C.N. The influence of head growth in fetal life, infancy, and childhood on intelligence at the ages of 4 and 8 years. Pediatrics 118, 1486–1492 (2006).
Gale, C.R., O'Callaghan, F.J., Godfrey, K.M., Law, C.M. & Martyn, C.N. Critical periods of brain growth and cognitive function in children. Brain 127, 321–329 (2004).
DeCarli, C. et al. Measures of brain morphology and infarction in the framingham heart study: establishing what is normal. Neurobiol. Aging 26, 491–510 (2005).
Ikram, M.A. et al. Brain tissue volumes in the general elderly population. The Rotterdam Scan Study. Neurobiol. Aging 29, 882–890 (2008).
Carmelli, D. et al. Evidence for genetic variance in white matter hyperintensity volume in normal elderly male twins. Stroke 29, 1177–1181 (1998).
Atwood, L.D. et al. Genetic variation in white matter hyperintensity volume in the Framingham Study. Stroke 35, 1609–1613 (2004).
Psaty, B.M. et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium. Design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ. Cardiovasc. Genet. 2, 73–80 (2009).
Seshadri, S. et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. J. Am. Med. Assoc. 303, 1832–1840 (2010).
Ikram, M.A. et al. Genomewide association studies of stroke. N. Engl. J. Med. 360, 1718–1728 (2009).
Taal, H.R. et al. Common variants at 12q15 and 12q24 are associated with infant head circumference. Nat. Genet. published online (15 April 2012); doi:10.1038/ng.2238.
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Di Maria, E. et al. The H1 haplotype of the tau gene (MAPT) is associated with mild cognitive impairment. J. Alzheimers Dis. 19, 909–914 (2010).
Zody, M.C. et al. Evolutionary toggling of the MAPT 17q21.31 inversion region. Nat. Genet. 40, 1076–1083 (2008).
Conrad, C. et al. Molecular evolution and genetics of the Saitohin gene and tau haplotype in Alzheimer's disease and argyrophilic grain disease. J. Neurochem. 89, 179–188 (2004).
Levecque, C. et al. Association of polymorphisms in the Tau and Saitohin genes with Parkinson's disease. J. Neurol. Neurosurg. Psychiatry 75, 478–480 (2004).
Gijselinck, I., Van Broeckhoven, C. & Cruts, M. Granulin mutations associated with frontotemporal lobar degeneration and related disorders: an update. Hum. Mutat. 29, 1373–1386 (2008).
Gijselinck, I. et al. Progranulin locus deletion in frontotemporal dementia. Hum. Mutat. 29, 53–58 (2008).
Hsuchou, H., Kastin, A.J., Wu, X., Tu, H. & Pan, W. Corticotropin-releasing hormone receptor–1 in cerebral microvessels changes during development and influences urocortin transport across the blood-brain barrier. Endocrinology 151, 1221–1227 (2010).
Mortimer, J.A., Snowdon, D.A. & Markesbery, W.R. Head circumference, education and risk of dementia: findings from the Nun Study. J. Clin. Exp. Neuropsychol. 25, 671–679 (2003).
Koolen, D.A. et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat. Genet. 38, 999–1001 (2006).
Koolen, D.A. et al. Clinical and molecular delineation of the 17q21.31 microdeletion syndrome. J. Med. Genet. 45, 710–720 (2008).
Stefansson, H. et al. A common inversion under selection in Europeans. Nat. Genet. 37, 129–137 (2005).
Donnelly, M.P. et al. The distribution and most recent common ancestor of the 17q21 inversion in humans. Am. J. Hum. Genet. 86, 161–171 (2010).
Roth, G. & Dicke, U. Evolution of the brain and intelligence. Trends Cogn. Sci. 9, 250–257 (2005).
Adams, J.C. & Tucker, R.P. The thrombospondin type 1 repeat (TSR) superfamily: diverse proteins with related roles in neuronal development. Dev. Dyn. 218, 280–299 (2000).
von Rotz, R.C., Kins, S., Hipfel, R., von der Kammer, H. & Nitsch, R.M. The novel cytosolic RING finger protein dactylidin is up-regulated in brains of patients with Alzheimer's disease. Eur. J. Neurosci. 21, 1289–1298 (2005).
Price, A.L. et al. Nat. Genet. 38, 904–909 (2006).
Acknowledgements
Aging Gene-Environment Susceptibility–Reykjavik Study (AGES-RS): Research was funded by the US National Institute on Aging (NIA) (N01-AG-12100), with contributions from the US National Eye Institute (NEI), National Institute on Deafness and Other Communication Disorders (NIDCD) and the NHLBI, the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association), and the Althingi (the Icelandic Parliament).
Atherosclerosis Risk in Communities Study (ARIC): The authors thank the staff and participants of the ARIC study for their important contributions. Research is carried out as a collaborative study supported by the US NHLBI (N01-HC-55015, N01-HC-55016, N01-HC-55018, N01-HC-55019, N01-HC-55020, N01-HC-55021, N01-HC-55022, R01-HL087641 and R01-HL093029), the National Human Genome Research Institute (U01-HG004402) and the NIH (HHSN268200625226C). Infrastructure was partly supported by a component of the NIH and NIH Roadmap for Medical Research (UL1RR025005).
Austrian Stroke Prevention Study (ASPS): The authors thank the staff and participants of the ASPS for their valuable contributions. We thank B. Reinhart for her long-term administrative commitment and I.J. Semmler for technical assistance in creating the DNA bank. The research reported here was funded by the Austrian Science Fond (FWF) (P20545-P05 and P13180). The Medical University of Graz supports the databank of the ASPS.
Framingham Heart Study (FHS): This study is carried out by the US NHLBI of the NIH and Boston University School of Medicine. This work was supported by the Framingham Heart Study of the NHLBI (N01-HC-25195) and its contract with Affymetrix, Inc, for genotyping services (N02-HL-6-4278). A portion of this research used the Linux Cluster for Genetic Analysis (LinGA-II) funded by the Robert Dawson Evans Endowment of the Department of Medicine at Boston University School of Medicine and Boston Medical Center. This study was also supported by grants from the National Institute of Neurological Disorders and Stroke (NS17950), the NHLBI (HL093029) and the NIA (AG08122, AG16495, AG033193, AG033040, AG031287 and P30AG013846).
Rotterdam Study (RS): The authors are grateful to the study participants, the staff from the Rotterdam Study and the participating general practitioners and pharmacists. The authors thank P. Arp, M. Jhamai, M. Verkerk, L. Herrera and M. Peters for their help in creating the GWAS database and K. Estrada and M.V. Struchalin for their support in creation and analysis of imputed data. The generation and management of GWAS genotype data for the Rotterdam Study were supported by the Netherlands Organisation of Scientific Research (NWO) Investments (nr. 175.010.2005.011 and 911-03-012). This study is funded by the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), the Netherlands Genomics Initiative (NGI)- NWO project (nr. 050-060-810). The Rotterdam Study is funded by Erasmus Medical Center and Erasmus University Organization for Health Research and Development (ZonMw), the Research Institute for Diseases in the Elderly (RIDE), the Ministry of Education, Culture and Science, the Ministry for Health, Welfare and Sports, the European Commission (DG XII) and the Municipality of Rotterdam. The Rotterdam Scan Study is supported by the NWO (nrs. 918-46-615, 904-61-096, 904-61-133 and 948-00-010), the Nederlandse Hartstichting (2009B102) and the Internationaal Parkinson Fonds.
EGG Consortium: We gratefully acknowledge the contribution of general practitioners, hospitals, midwives and pharmacies in Rotterdam. Financial support was received from the Academy of Finland (104781, 120315, 1114194 and Center of Excellence in Complex Disease Genetics), University Hospital Oulu, Biocenter, University of Oulu, the US NHLBI (5R01HL087679-02, through the STAMPEED program 1RL1MH083268-01), the ENGAGE project and grant agreement HEALTH-F4-2007-201413, the UK MRC (G0500539 and PrevMetSyn/Salve/MRC) and the Wellcome Trust (GR069224). DNA extraction, sample quality control, biobank upkeep and aliquotting were performed at the National Public Health Institute, Biomedicum Helsinki (Helsinki, Finland) and supported financially by the Academy of Finland and Biocentrum Helsinki. The Generation R Study is conducted by the Erasmus Medical Center in close collaboration with the School of Law and Faculty of Social Sciences of the Erasmus University Rotterdam, the Municipal Health Service Rotterdam area, the Rotterdam Homecare Foundation and the Stichting Trombosedienst & Artsenlaboratorium Rijnmond (STAR). The Generation R Study is made possible by financial support from the Erasmus Medical Center, Rotterdam, the Erasmus University Rotterdam and the Netherlands Organization for Health Research and Development (ZonMw 21000074). V.J. received additional grants from the Netherlands Organization for Health Research and Development (ZonMw 90700303 and 916.10159). Additional support was provided by a grant from the Dutch Kidney Foundation (C08.2251).
The authors would like to thank all participating subjects and families from the Children's Hospital in Philadelphia. The research was financially supported by an Institute Development Award from the Children's Hospital of Philadelphia, a Research Development Award from the Cotswold Foundation and by the US NIH (1R01HD056465-01A1). The authors are grateful to the Raine Foundation, the RAINE Study families and the RAINE Study research staff. We gratefully acknowledge the assistance of the Western Australian Genetic Epidemiology Resource and the Western Australian DNA Bank (both National Health and Medical Research Council of Australia National Enabling Facilities). The authors also acknowledge the support of Healthway Western Australia, the National Health and Medical Research Council of Australia (572613) and the Canadian Institutes of Health Research (MOP 82893). We gratefully acknowledge the assistance of the Wind Over Water Foundation, the Telethon Institute for Child Health Research and the RAINE Medical Research Foundation of the University of Western Australia. The authors wish to acknowledge the following: Helmholtz Zentrum Muenchen–German Research Center for Environment and Health, Institute of Epidemiology; the Department of Pediatrics, University of Leipzig; Department of Pediatrics, Marien-Hospital; Bad Honnef; the Department of Human Exposure Research and Epidemiology, UFZ-Centre for Environmental Research Leipzig–Halle; the Department of Environmental Immunology, Zentrum für Umweltforschung (UFZ)-Centre for Environmental Research Leipzig–Halle; the Institut für Umweltmedizinische Forschung (IUF) and the Department of Pediatrics, Technical University. The UK MRC (74882), the Wellcome Trust (076467) and the University of Bristol provide core support for the Avon Longitudinal Study of Parents and Children (ALSPAC). We are extremely grateful to all the families who took part in the ALSPAC study, the midwives for their help in recruiting them and the whole ALSPAC team, which includes interviewers, computer and laboratory technicians, clerical workers, research scientists, volunteers, managers, receptionists and nurses.
Author information
Authors and Affiliations
Consortia
Contributions
Study concept and design was carried out by M.A.I., M.F., S. Seshadri, R.S., W.T.L., A.G.U., A.-L.H., E.A.P.S., J.H., L.J.P., S.F.A.G., N.J.T., G.D.S., A.H., A.v.d.L., M.A.v.B., V.W.V.J., V.G., P.A.W., C.M.v.D., T.H.M., H.S., L.J.L., M.M.B.B. and C.D. Acquisition of data was carried out by M.A.I., M.F., A.V.S., S.D., H.A.V., S. Sigurdsson, S.R., H.R.T., D.O.M.-K., L.H.C., W.J.N., A.B., A.P.Z., M.S., C.R.J., F.R., D.S.K., C.E.P., E.A.P.S., H.H., L.J.P., M.-R.J., S.F.A.G., B.S.P., M.A.N., R.A., H.G., T.B.H., W.M.M., M.W.V., M.A.v.B., D.C., B.G.W. and T.H.M. Statistical analysis and interpretation of the findings were performed by M.A.I., M.F., A.V.S., S. Seshadri, R.S., S.D., H.R.T., D.O.M.-K., A.L.D., F.R., E.T., S.F.A.G., M.A.N., R.A., V.W.V.J., C.M.v.D., H.S., L.J.L., M.M.B.B. and C.D. The manuscript was drafted by M.A.I., M.F., S. Seshadri, R.S., H.R.T., D.O.M.-K., L.J.L. and C.D. Critical revision of the manuscript was performed by A.V.S., S.D., H.A.V., S. Sigurdsson, S.R., L.H.C., W.T.L., W.J.N., A.L.D., A.B., A.P.Z., M.S., C.R.J., F.R., A.G.U., D.S.K., A.-L.H., C.E.P., E.T., E.A.P.S., H.H., J.H., L.J.P., M.-R.J., M.I.M., B.S.P., N.J.T., G.D.S., U.S., M.A.N., R.A., A.H., H.G., A.v.d.L., T.B.H., W.M.M., M.W.V., M.A.v.B., V.G., B.G.W., P.A.W., C.M.v.D., T.H.M., H.S. and M.M.B.B. A.V.S., S. Seshadri, R.S., W.T.L., W.J.N., A.G.U., M.-R.J., M.I.M., S.F.A.G., N.J.T., U.S., A.H., A.v.d.L., M.A.v.B., D.C., V.W.V.J., V.G., P.A.W., C.M.v.D., T.H.M., H.S., L.J.L., M.M.B.B. and C.D. obtained funding and supervised the studies.
Corresponding authors
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Tables 1–3 and Supplementary Note (PDF 548 kb)
Rights and permissions
About this article
Cite this article
the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium., Early Growth Genetics (EGG) Consortium. Common variants at 6q22 and 17q21 are associated with intracranial volume. Nat Genet 44, 539–544 (2012). https://doi.org/10.1038/ng.2245
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2245
This article is cited by
-
Identifying genes associated with brain volumetric differences through tissue specific transcriptomic inference from GWAS summary data
BMC Bioinformatics (2022)
-
Genetics of early-life head circumference and genetic correlations with neurological, psychiatric and cognitive outcomes
BMC Medical Genomics (2022)
-
Toward a global and reproducible science for brain imaging in neurotrauma: the ENIGMA adult moderate/severe traumatic brain injury working group
Brain Imaging and Behavior (2021)
-
The genetic regulation of size variation in the transcriptome of the cerebrum in the chicken and its role in domestication and brain size evolution
BMC Genomics (2020)
-
Translational research on reserve against neurodegenerative disease: consensus report of the International Conference on Cognitive Reserve in the Dementias and the Alzheimer’s Association Reserve, Resilience and Protective Factors Professional Interest Area working groups
BMC Medicine (2019)