Abstract
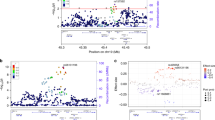
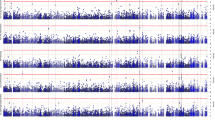
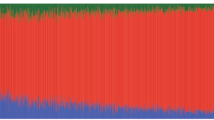
Aging is associated with reductions in hippocampal volume that are accelerated by Alzheimer's disease and vascular risk factors. Our genome-wide association study (GWAS) of dementia-free persons (n = 9,232) identified 46 SNPs at four loci with P values of <4.0 × 10−7. In two additional samples (n = 2,318), associations were replicated at 12q14 within MSRB3-WIF1 (discovery and replication; rs17178006; P = 5.3 × 10−11) and at 12q24 near HRK-FBXW8 (rs7294919; P = 2.9 × 10−11). Remaining associations included one SNP at 2q24 within DPP4 (rs6741949; P = 2.9 × 10−7) and nine SNPs at 9p33 within ASTN2 (rs7852872; P = 1.0 × 10−7); along with the chromosome 12 associations, these loci were also associated with hippocampal volume (P < 0.05) in a third younger, more heterogeneous sample (n = 7,794). The SNP in ASTN2 also showed suggestive association with decline in cognition in a largely independent sample (n = 1,563). These associations implicate genes related to apoptosis (HRK), development (WIF1), oxidative stress (MSR3B), ubiquitination (FBXW8) and neuronal migration (ASTN2), as well as enzymes targeted by new diabetes medications (DPP4), indicating new genetic influences on hippocampal size and possibly the risk of cognitive decline and dementia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ridha, B.H. et al. Tracking atrophy progression in familial Alzheimer's disease: a serial MRI study. Lancet Neurol. 5, 828–834 (2006).
Small, G.W. Use of neuroimaging to detect early brain changes in people at genetic risk for Alzheimer's disease. Adv. Drug Deliv. Rev. 54, 1561–1566 (2002).
den Heijer, T. et al. Type 2 diabetes and atrophy of medial temporal lobe structures on brain MRI. Diabetologia 46, 1604–1610 (2003).
Seshadri, S. et al. Association of plasma total homocysteine levels with subclinical brain injury: cerebral volumes, white matter hyperintensity, and silent brain infarcts at volumetric magnetic resonance imaging in the Framingham Offspring Study. Arch. Neurol. 65, 642–649 (2008).
Sullivan, E.V., Pfefferbaum, A., Swan, G.E. & Carmelli, D. Heritability of hippocampal size in elderly twin men: equivalent influence from genes and environment. Hippocampus 11, 754–762 (2001).
Psaty, B.M. et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ. Cardiovasc. Genet. 2, 73–80 (2009).
Zapala, M.A. et al. Adult mouse brain gene expression patterns bear an embryologic imprint. Proc. Natl. Acad. Sci. USA 102, 10357–10362 (2005).
Sborgi, L., Barrera-Vilarmau, S., Obregon, P. & de Alba, E. Characterization of a novel interaction between Bcl-2 members Diva and Harakiri. PLoS ONE 5, e15575 (2010).
Lukiw, W.J. & Bazan, N.G. Inflammatory, apoptotic, and survival gene signaling in Alzheimer's disease. A review on the bioactivity of neuroprotectin D1 and apoptosis. Mol. Neurobiol. 42, 10–16 (2010).
Inohara, N., Ding, L., Chen, S. & Nunez, G. harakiri, a novel regulator of cell death, encodes a protein that activates apoptosis and interacts selectively with survival-promoting proteins Bcl-2 and Bcl-XL . EMBO J. 16, 1686–1694 (1997).
Imaizumi, K. et al. The cell death–promoting gene DP5, which interacts with the BCL2 family, is induced during neuronal apoptosis following exposure to amyloid β protein. J. Biol. Chem. 274, 7975–7981 (1999).
Guan, Q.H., Pei, D.S., Xu, T.L. & Zhang, G.Y. Brain ischemia/reperfusion-induced expression of DP5 and its interaction with Bcl-2, thus freeing Bax from Bcl-2/Bax dimmers are mediated by c-Jun N-terminal kinase (JNK) pathway. Neurosci. Lett. 393, 226–230 (2006).
Rohn, T.T. The role of caspases in Alzheimer's disease; potential novel therapeutic opportunities. Apoptosis 15, 1403–1409 (2010).
Segref, A. & Hoppe, T. Think locally: control of ubiquitin-dependent protein degradation in neurons. EMBO Rep. 10, 44–50 (2009).
Riederer, B.M., Leuba, G., Vernay, A. & Riederer, I.M. The role of the ubiquitin proteasome system in Alzheimer's disease. Exp. Biol. Med. (Maywood) 236, 268–276 (2011).
Liao, E.H., Hung, W., Abrams, B. & Zhen, M. An SCF-like ubiquitin ligase complex that controls presynaptic differentiation. Nature 430, 345–350 (2004).
Litterman, N. et al. An OBSL1-Cul7Fbxw8 ubiquitin ligase signaling mechanism regulates Golgi morphology and dendrite patterning. PLoS Biol. 9, e1001060 (2011).
Veyrieras, J.B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Myers, A.J. et al. A survey of genetic human cortical gene expression. Nat. Genet. 39, 1494–1499 (2007).
Stranger, B.E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005).
Kalmijn, S. et al. Total homocysteine and cognitive decline in a community-based sample of elderly subjects: the Rotterdam Study. Am. J. Epidemiol. 150, 283–289 (1999).
Prins, N.D. et al. Homocysteine and cognitive function in the elderly: the Rotterdam Scan Study. Neurology 59, 1375–1380 (2002).
den Heijer, T. et al. Homocysteine and brain atrophy on MRI of non-demented elderly. Brain 126, 170–175 (2003).
González, S. et al. Serum selenium is associated with plasma homocysteine concentrations in elderly humans. J. Nutr. 134, 1736–1740 (2004).
Kim, H. et al. Downregulation of Wnt/β-catenin signaling causes degeneration of hippocampal neurons in vivo. Neurobiol. Aging 32, 2316.e1–2316.e15 (2011).
Gogolla, N., Galimberti, I., Deguchi, Y. & Caroni, P. Wnt signaling mediates experience-related regulation of synapse numbers and mossy fiber connectivities in the adult hippocampus. Neuron 62, 510–525 (2009).
Chibnik, L.B. et al. CR1 is associated with amyloid plaque burden and age-related cognitive decline. Ann. Neurol. 69, 560–569 (2011).
Okamoto, O.K. et al. Whole transcriptome analysis of the hippocampus: toward a molecular portrait of epileptogenesis. BMC Genomics 11, 230 (2010).
Henrich-Noack, P., Prehn, J.H. & Krieglstein, J. TGF-β1 protects hippocampal neurons against degeneration caused by transient global ischemia. Dose-response relationship and potential neuroprotective mechanisms. Stroke 27, 1609–1614, discussion 1615 (1996).
Kubben, N. et al. Identification of differential protein interactors of lamin A and progerin. Nucleus 1, 513–525 (2010).
Mentlein, R. Dipeptidyl-peptidase IV (CD26)—role in the inactivation of regulatory peptides. Regul. Pept. 85, 9–24 (1999).
Gault, V.A. & Holscher, C. GLP-1 agonists facilitate hippocampal LTP and reverse the impairment of LTP induced by β-amyloid. Eur. J. Pharmacol. 587, 112–117 (2008).
Bernstein, H.G., Schon, E., Ansorge, S., Rose, I. & Dorn, A. Immunolocalization of dipeptidyl aminopeptidase (DAP IV) in the developing human brain. Int. J. Dev. Neurosci. 5, 237–242 (1987).
Lamers, D. et al. Dipeptidyl peptidase 4 is a novel adipokine potentially linking obesity to the metabolic syndrome. Diabetes 60, 1917–1925 (2011).
Jagust, W., Harvey, D., Mungas, D. & Haan, M. Central obesity and the aging brain. Arch. Neurol. 62, 1545–1548 (2005).
Hamilton, A., Patterson, S., Porter, D., Gault, V.A. & Holscher, C. Novel GLP-1 mimetics developed to treat type 2 diabetes promote progenitor cell proliferation in the brain. J. Neurosci. Res. 89, 481–489 (2011).
Hamilton, A. & Holscher, C. Receptors for the incretin glucagon-like peptide–1 are expressed on neurons in the central nervous system. Neuroreport 20, 1161–1166 (2009).
During, M.J. et al. Glucagon-like peptide–1 receptor is involved in learning and neuroprotection. Nat. Med. 9, 1173–1179 (2003).
Wilson, P.M., Fryer, R.H., Fang, Y. & Hatten, M.E. Astn2, a novel member of the astrotactin gene family, regulates the trafficking of ASTN1 during glial-guided neuronal migration. J. Neurosci. 30, 8529–8540 (2010).
Gasser, U.E. & Hatten, M.E. Neuron-glia interactions of rat hippocampal cells in vitro: glial-guided neuronal migration and neuronal regulation of glial differentiation. J. Neurosci. 10, 1276–1285 (1990).
Lambert, J.C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat. Genet. 41, 1094–1099 (2009).
Harold, D. et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer's disease. Nat. Genet. 41, 1088–1093 (2009).
Hollingworth, P. et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nat. Genet. 43, 429–435 (2011).
Naj, A.C. et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer's disease. Nat. Genet. 43, 436–441 (2011).
Seshadri, S. et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. J. Am. Med. Assoc. 303, 1832–1840 (2010).
Corneveaux, J.J. et al. Association of CR1, CLU and PICALM with Alzheimer's disease in a cohort of clinically characterized and neuropathologically verified individuals. Hum. Mol. Genet. 19, 3295–3301 (2010).
van der Lijn, F., den Heijer, T., Breteler, M.M. & Niessen, W.J. Hippocampus segmentation in MR images using atlas registration, voxel classification, and graph cuts. Neuroimage 43, 708–720 (2008).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Johnson, A.D. & O'Donnell, C.J. An open access database of genome-wide association results. BMC Med. Genet. 10, 6 (2009).
Stein, J.L. et al. Identification of common variants associated with human hippocampal and intracranial volumes. Nat. Genet. published online (15 April 2012); doi:10.1038/ng.2250.
Acknowledgements
The authors thank the ENIGMA Consortium, which is represented by J.L.S., S.E.M., A.A.V., D.P.H., M.J.W., B.F., N.G.M. and P.M.T.
Aging Gene-Environment Susceptibility–Reykjavik Study (AGES): Research was funded by the US National Institute on Aging (NIA; N01-AG-12100), with contributions from the National Eye Institute (NEI), the National Institute on Deafness and Other Communication Disorders (NIDCD), the US National Heart, Lung, and Blood Institute (NHLBI), the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association) and the Althingi (the Icelandic Parliament).
The Atherosclerosis Risk in Communities Study (ARIC): The authors thank the staff and participants of the ARIC study for their important contributions. Research was carried out as a collaborative study supported by the US NHLBI (HHSN268201100005C, HHSN268201100006C, HHSN268201100007C, HHSN268201100008C, HHSN268201100009C, HHSN268201100010C, HHSN268201100011C, HHSN268201100012C, HL087641, HL59367, HL086694 and HL7825); the National Human Genome Research Institute (U01HG004402) and the NIH (HHSN268200625226C). Infrastructure was partly supported by a component of the NIH and the NIH Roadmap for Medical Research (UL1RR025005). This project was also supported by NHLBI grant HL093029.
The Cardiovascular Health Study (CHS): Coauthors were supported in part by US NHLBI grants (HL087652 and HL105756), as well as by NIA grants (AG20098 and AG05133).
The Austrian Stroke Prevention Study (ASPS): The authors thank the staff and the participants of ASPS for their valuable contributions. We thank B. Reinhart for her long-term administrative commitment and I.J. Semmler for technical assistance in creating the DNA bank. The research reported here was funded by the Austrian Science Fond (FWF; P20545-P05 and P13180). The Medical University of Graz supports the databank of ASPS.
Erasmus Rucphen Family Study (ERF): We thank the participants from the Genetic Research in Isolated Populations in the Erasmus Rucphen Family Study who made this work possible. This study is financially supported by the Netherlands Organisation for Scientific Research (NWO), the Internationale Stichting Alzheimer Onderzoek (ISAO), the Hersenstichting Nederland (HSN) and the Centre for Medical Systems Biology (CMSB1 and CMSB2) in the framework of the Netherlands Genomics Initiative (NGI).
Framingham Heart Study (FHS): This work was supported by the National Heart, Lung and Blood Institute's Framingham Heart Study (contract N01-HC-25195) and its contract with Affymetrix, Inc, for genotyping services (contract N02-HL-6-4278). A portion of this research used the Linux Cluster for Genetic Analysis (LinGA-II) funded by the Robert Dawson Evans Endowment of the Department of Medicine at the Boston University School of Medicine and Boston Medical Center. Analyses reflect intellectual input and resource development from the Framingham Heart Study investigators participating in the SNP Health Association Resource (SHARe) project. This study was also supported by grants from the NINDS (NS17950) and the NIA (AG08122, AG16495, AG033193, AG013846 and AG031287).
The Religious Order Study and the Rush Memory and Aging Project: ROS and R-MAP data used in this study were obtained with support from the US NIA (grants P30AG10161, AG17917 and AG15819), the Illinois Department of Public Health and the Rush Clinical Translational Science Consortium and a gift from M. Dowd.
The Rotterdam Study (RS): The authors are grateful to the study participants, the staff from the Rotterdam Study and the participating general practitioners and pharmacists. The authors also thank P. Arp, M. Jhamai, M. Verkerk, L. Herrera and M. Peters for their help in creating the GWAS database and K. Estrada and M.V. Struchalin for their support in the creation and analysis of imputed data. The generation and management of GWAS genotype data for the Rotterdam Study are supported by NWO Investments (nr. 175.010.2005.011 and 911-03-012). This study is funded by the Research Institute for Diseases in the Elderly (RIDE2; 014-93-015) and the NGI-NWO project (nr. 050-060-810). The Rotterdam Study is funded by the Erasmus Medical Center and Erasmus University, Rotterdam, the Netherlands Organisation for Health Research and Development (ZonMw), RIDE2, the Dutch Ministry of Education, Culture and Science, the Dutch Ministry for Health, Welfare and Sports, the European Commission (DG XII) and the Municipality of Rotterdam. The Rotterdam Scan Study is supported by the NWO (project nrs. 918-46-615, 904-61-096, 904-61-133 and 948-00-010), Nederlandse Hartstichting (2009B102) and Internationaal Parkinson Fonds.
The Tasmanian Study of Gait and Cognition (TASCOG): This study is supported by project grants from the National Health and Medical Research Council of Australia (NHMRC; 403000, 491109 and 606543) and a grant from the Wicking Dementia Education and Research Centre, Hobart. V.S. is supported by an NHMRC–National Heart Foundation Career Development Fellowship (606544). M.A.B. is supported by an NHMRC Senior Principal Research Fellowship (APP1024879).
Three City Study (3C): We thank the staff and participants of the 3C Study for their important contributions. We also thank A. Boland for her technical help in preparing the DNA samples for analyses. The 3C Study is conducted under a partnership agreement between INSERM, Victor Segalen–Bordeaux II University and Sanofi-Aventis. The Fondation pour la Recherche Médicale funded the preparation and initiation of the study. The 3C Study is also supported by the Caisse Nationale Maladie des Travailleurs Salariés, Direction Générale de la Santé, Mutuelle Générale de l'Education Nationale (MGEN), Institut de la Longévité, Conseils Régionaux de Aquitaine et Bourgogne, Fondation de France and the French Ministry of Research–INSERM Programme Cohortes et Collections de Données Biologiques. This work was supported by the National Foundation for Alzheimer's Disease and Related Disorders, the Institut Pasteur de Lille and the Centre National de Génotypage.
Author information
Authors and Affiliations
Consortia
Contributions
Study concept and design were performed by J.C.B., C. DeCarli, M.A.v.B., C. Dufouil, P.A., A.G.U., M.M.B.B., F.F., M.A.B., C.M.v.D., T.H.M., C.T., L.J.L., M.A.I. and S. Seshadri. Acquisition of data was carried out by F.v.d.L., F.C., H.S., V.S., M. Schuur, S. Sigurdsson, B.F.J.V., J.-C.L., C.R.J., J.S., D.F., T.d.H., B.M., L.H.C., C.E., P.D., K.A., M.A.v.B., A.B., C. Dufouil, J.H., W.J.N., G.K.G., P.A., K.B.F., T.G.P., B.A.O., A.G.U., R.A., A.E., R.J.B., J.C.v.S., M.M.B.B., M.W.V., P.A.W., A.v.d.L., V.G., M.A.B., D.A.B., C.M.v.D., T.H.M., R.S., C.T., L.J.L., M.A.I. and S. Seshadri. Statistical analysis and interpretation of the data were performed by J.C.B., C. DeCarli, A.V.S., F.v.d.L., M.F., S.D., J.M.S., H.S., V.S., M. Schuur, L.Y., S.-H.C., B.F.J.V., A.L.D., M. Struchalin, J.S., C.A.I.-V., N.A., R.F.A.G.d.B., M.C., R.T., L.B.C., G.K.G., P.A., A.E., R.J.B., P.L.D.J., M.A.B., D.A.B., R.S., L.J.L. and M.A.I. The manuscript was drafted by J.C.B., C. DeCarli, J.M.S., H.S., M. Schuur, B.M., P.L.D.J., L.J.L. and S. Seshadri, and critical revision of the manuscript was performed by J.C.B., C. DeCarli, F.v.d.L., M.F., S.D., J.M.S., H.S., V.S., S.-H.C., S. Sigurdsson, A.L.D., J.-C.L., J.S., C.A.I.-V., A.Z., T.d.H., L.H.C., C.E., P.D., C. Dufouil, M.C., R.T., W.J.N., L.B.C., A.H., A.P., K.B.F., T.G.P., J.L.S., S.E.M., A.A.V., D.P.H., M.J.W., B.F., N.G.M., P.M.T., M.A.N., A.G.U., A.E., R.J.B., O.L.L., T.B.H., V.C., M.M.B.B., J.T.B., M.W.V., D.K., F.F., P.A.W., A.v.d.L., V.G., W.T.L., M.A.B., C.M.v.D., T.H.M., R.S., C.T., L.J.L., M.A.I. and S. Seshadri. Funding was obtained by M.F., H.S., V.S., C. Dufouil, W.J.N., A.H., B.A.O., A.G.U., J.C.v.S., T.B.H., V.C., M.M.B.B., F.F., P.A.W., A.v.d.L., V.G., D.A.B., C.M.v.D., T.H.M., R.S., C.T., L.J.L. and S. Seshadri.
Corresponding author
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–7, Supplementary Figures 1–3 and Supplementary Note (PDF 1133 kb)
Rights and permissions
About this article
Cite this article
Enhancing Neuro Imaging Genetics through Meta-Analysis (ENIGMA) Consortium., the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium. Common variants at 12q14 and 12q24 are associated with hippocampal volume. Nat Genet 44, 545–551 (2012). https://doi.org/10.1038/ng.2237
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2237
This article is cited by
-
A fast non-parametric test of association for multiple traits
Genome Biology (2023)
-
Current Update on Categorization of Migraine Subtypes on the Basis of Genetic Variation: a Systematic Review
Molecular Neurobiology (2023)
-
Genetic and Environmental Contributions to Subcortical Gray Matter Microstructure and Volume in the Developing Brain
Behavior Genetics (2023)
-
Identifying genes associated with brain volumetric differences through tissue specific transcriptomic inference from GWAS summary data
BMC Bioinformatics (2022)
-
ENIGMA + COINSTAC: Improving Findability, Accessibility, Interoperability, and Re-usability
Neuroinformatics (2022)