Abstract
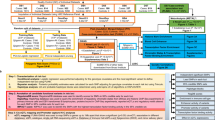
We have genotyped 14,436 nonsynonymous SNPs (nsSNPs) and 897 major histocompatibility complex (MHC) tag SNPs from 1,000 independent cases of ankylosing spondylitis (AS), autoimmune thyroid disease (AITD), multiple sclerosis (MS) and breast cancer (BC). Comparing these data against a common control dataset derived from 1,500 randomly selected healthy British individuals, we report initial association and independent replication in a North American sample of two new loci related to ankylosing spondylitis, ARTS1 and IL23R, and confirmation of the previously reported association of AITD with TSHR and FCRL3. These findings, enabled in part by increased statistical power resulting from the expansion of the control reference group to include individuals from the other disease groups, highlight notable new possibilities for autoimmune regulation and suggest that IL23R may be a common susceptibility factor for the major 'seronegative' diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Notes
Ian N Bruce and Michael R Stratton: *See Supplementary Note for details.
References
Rioux, J.D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Sladek, R. et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445, 881–885 (2007).
Easton, D.F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Libioulle, C. et al. Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS Genet. 3, e58 (2007).
Zanke, B.W. et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat. Genet. 39, 989–994 (2007).
Haiman, C.A. et al. Multiple regions within 8q24 independently affect risk for prostate cancer. Nat. Genet. 39, 638–644 (2007).
Gudmundsson, J. et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat. Genet. 39, 977–983 (2007).
Moffatt, M.F. et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448, 470–473 (2007).
Zeggini, E. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316, 1336–1341 (2007).
Scott, L.J. et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316, 1341–1345 (2007).
Saxena, R. et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316, 1331–1336 (2007).
WTCCC. Genome-wide association studies of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–683 (2007).
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Nat. Genet. 39, 207–211 (2007).
Jorgenson, E. & Witte, J.S. Coverage and power in genomewide association studies. Am. J. Hum. Genet. 78, 884–888 (2006).
Smyth, D.J. et al. A genome-wide association study of nonsynonymous SNPs identifies a type 1 diabetes locus in the interferon-induced helicase (IFIH1) region. Nat. Genet. 38, 617–619 (2006).
Miretti, M.M. et al. A high-resolution linkage-disequilibrium map of the human major histocompatibility complex and first generation of tag single-nucleotide polymorphisms. Am. J. Hum. Genet. 76, 634–646 (2005).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Sims, A.M. et al. Non-B27 MHC associations of ankylosing spondylitis. Genes Immun. 8, 115–123 (2007).
McGinnis, R., Shifman, S. & Darvasi, A. Power and efficiency of the TDT and case-control design for association scans. Behav. Genet. 32, 135–144 (2002).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Thomas, D.C. & Clayton, D.G. Betting odds and genetic associations. J. Natl. Cancer Inst. 96, 421–423 (2004).
Jiang, H.R. et al. Sialoadhesin promotes the inflammatory response in experimental autoimmune uveoretinitis. J. Immunol. 177, 2258–2264 (2006).
Dechairo, B.M. et al. Association of the TSHR gene with Graves' disease: the first disease specific locus. Eur. J. Hum. Genet. 13, 1223–1230 (2005).
Hiratani, H. et al. Multiple SNPs in intron 7 of thyrotropin receptor are associated with Graves' disease. J. Clin. Endocrinol. Metab. 90, 2898–2903 (2005).
Miettinen, O.S. Proportion of disease caused or prevented by a given exposure, trait or intervention. Am. J. Epidemiol. 99, 325–332 (1974).
Duerr, R.H. et al. A Genome-Wide Association Study Identifies IL23R as an Inflammatory Bowel Disease Gene. Science (2006).
Tremelling, M. et al. IL23R variation determines susceptibility but not disease phenotype in inflammatory bowel disease. Gastroenterology 132, 1657–1664 (2007).
Cargill, M. et al. A large-scale genetic association study confirms IL12B and leads to the identification of IL23R as psoriasis-risk genes. Am. J. Hum. Genet. 80, 273–290 (2007).
Simmonds, M.J. et al. Contribution of single nucleotide polymorphisms within FCRL3 and MAP3K7IP2 to the pathogenesis of Graves' disease. J. Clin. Endocrinol. Metab. 91, 1056–1061 (2006).
Kochi, Y. et al. A functional variant in FCRL3, encoding Fc receptor-like 3, is associated with rheumatoid arthritis and several autoimmunities. Nat. Genet. 37, 478–485 (2005).
Capon, F. et al. Fine mapping of the PSORS4 psoriasis susceptibility region on chromosome 1q21. J. Invest. Dermatol. 116, 728–730 (2001).
Dai, K.Z. et al. The T cell regulator gene SH2D2A contributes to the genetic susceptibility of multiple sclerosis. Genes Immun. 2, 263–268 (2001).
Chang, S.C., Momburg, F., Bhutani, N. & Goldberg, A.L. The ER aminopeptidase, ERAP1, trims precursors to lengths of MHC class I peptides by a “molecular ruler” mechanism. Proc. Natl. Acad. Sci. USA 102, 17107–17112 (2005).
Saveanu, L. et al. Concerted peptide trimming by human ERAP1 and ERAP2 aminopeptidase complexes in the endoplasmic reticulum. Nat. Immunol. 6, 689–697 (2005).
Brown, M.A. et al. HLA class I associations of ankylosing spondylitis in the white population in the United Kingdom. Ann. Rheum. Dis. 55, 268–270 (1996).
Cui, X., Rouhani, F.N., Hawari, F. & Levine, S.J. Shedding of the type II IL-1 decoy receptor requires a multifunctional aminopeptidase, aminopeptidase regulator of TNF receptor type 1 shedding. J. Immunol. 171, 6814–6819 (2003).
Cui, X., Rouhani, F.N., Hawari, F. & Levine, S.J. An aminopeptidase, ARTS-1, is required for interleukin-6 receptor shedding. J. Biol. Chem. 278, 28677–28685 (2003).
Cui, X. et al. Identification of ARTS-1 as a novel TNFR1-binding protein that promotes TNFR1 ectodomain shedding. J. Clin. Invest. 110, 515–526 (2002).
Cua, D.J. et al. Interleukin-23 rather than interleukin-12 is the critical cytokine for autoimmune inflammation of the brain. Nature 421, 744–748 (2003).
Murphy, C.A. et al. Divergent pro- and antiinflammatory roles for IL-23 and IL-12 in joint autoimmune inflammation. J. Exp. Med. 198, 1951–1957 (2003).
Hue, S. et al. Interleukin-23 drives innate and T cell-mediated intestinal inflammation. J. Exp. Med. 203, 2473–2483 (2006).
Mannon, P.J. et al. Anti-interleukin-12 antibody for active Crohn's disease. N. Engl. J. Med. 351, 2069–2079 (2004).
Cohen, J.C. et al. Multiple rare alleles contribute to low plasma levels of HDL cholesterol. Science 305, 869–872 (2004).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Wigginton, J.E., Cutler, D.J. & Abecasis, G.R. A note on exact tests of Hardy-Weinberg equilibrium. Am. J. Hum. Genet. 76, 887–893 (2005).
Armitage, P. Test for linear trend in proportions and frequencies. Biometrics 11, 375–386 (1955).
Purcell, S. et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Purcell, S., Daly, M.J. & Sham, P.C. WHAP: haplotype-based association analysis. Bioinformatics 23, 255–256 (2007).
Marchini, J., Cardon, L.R., Phillips, M.S. & Donnelly, P. The effects of human population structure on large genetic association studies. Nat. Genet. 36, 512–517 (2004).
Acknowledgements
We would like to thank the Wellcome Trust for supporting this study, and all the individuals and controls who participated in this study.
AITD: We thank the collection coordinators, J. Carr-Smith and all contributors to the AITD national DNA collection of index cases and family members from centres including Birmingham, Bournemouth, Cambridge, Cardiff, Exeter, Leeds, Newcastle and Sheffield. Principal leads for the AITD UK national collection are S.C. Gough (Birmingham), S.H.S. Pearce (Newcastle), B. Vaidya (Exeter), J.H. Lazarus (Cardiff), A. Allahabadia (Sheffield), M. Armitage (Bournemouth), P.J. Grant (Leeds) and V.K. Chatterjee (Cambridge).
Ankylosing spondylitis: We thank the Arthritis Research Campaign (UK). MAB is funded by the National Health and Medical Research Council (Australia). TASC is funded by the National Institute of Arthritis and Musculoskeletal and Skin Diseases grants 1PO1-052915-01,RO1 AR046208 and RO1-AR048465, as well as by University of Texas at Houston CTSA grant UL1RR024148, Cedars-Sinai GCRC grant MO1-RR00425, The Rosalind Russell Center for Arthritis Research at The University of California San Francisco, and the Intramural Research Program, National Institute of Arthritis and Musculoskeletal and Skin Diseases, US National Institutes of Health. We thank R. Jin for technical assistance and L. Diekman, L. Guthrie, F. Lin and S. Morgan for their study coordination.
Breast cancer: The breast cancer samples were clinically and molecularly curated with the assistance of A. Renwick, A. Hall, A. Elliot, H. Jayatilake, T. Chagtai, R. Barfoot, P. Kelly and K. Spanova. Our research is supported by United States Army Medical Research and Material Command grant no. W81XWH-05-1-0204, The Institute of Cancer Research and Cancer Research UK.
MS: Our work has been supported by the Wellcome Trust (grant ref. 057097), the Medical Research Council (UK) (grant ref. G0000648), the Multiple Sclerosis Society of Great Britain and Northern Ireland (grant ref. 730/02) and the National Institutes of Health (USA) (grant ref. 049477). A.G. is a postdoctoral fellow of the Research Foundation–Flanders (FWO–Vlaanderen).
L. Galver and P. Ng at Illumina and J. Morrison at the Sanger Institute contributed in the design of the nsSNP array. We thank the DNA team of the JDRF/WT DIL and T. Dibling, C. Hind and D. Simpkin at the Sanger Institute for carrying out the genotyping. We also thank S. Bingham and the WTCCC Inflammatory Bowel Disease group for genotyping the ARTS1 markers in their replication samples.
The complete list of authors is as follows: Wellcome Trust Case Control Consortium Management Committee: Paul R Burton1, David G Clayton2, Lon R Cardon3,55, Nick Craddock4, Panos Deloukas5, Audrey Duncanson6, Dominic P Kwiatkowski3,5, Mark I McCarthy3,7, Willem H Ouwehand8,9, Nilesh J Samani10, John A Todd2, Peter Donnelly (Chair)11; Analysis Committee: Jeffrey C Barrett3, Paul R Burton1, Dan Davison11, Peter Donnelly11, Doug Easton12, David M Evans3, Hin-Tak Leung2, Jonathan L Marchini11, Andrew P Morris3, Chris CA Spencer11, Martin D Tobin1, Lon R Cardon3,55, David G Clayton2; UK Blood Services & University of Cambridge Controls: Antony P Attwood5,8, James P Boorman8,9, Barbara Cant8, Ursula Everson13, Judith M Hussey14, Jennifer D Jolley8, Alexandra S Knight8, Kerstin Koch8, Elizabeth Meech15, Sarah Nutland2, Christopher V Prowse16, Helen E Stevens2, Niall C Taylor8, Graham R Walters17, Neil M Walker2, Nicholas A Watkins8,9, Thilo Winzer8, John A Todd2, Willem H Ouwehand8,9; 1958 Birth Cohort Controls: Richard W Jones18, Wendy L McArdle18, Susan M Ring18, David P Strachan19, Marcus Pembrey18,20. Bipolar Disorder (Aberdeen): Gerome Breen21, David St Clair21; (Birmingham): Sian Caesar22, Katharine Gordon-Smith22,23, Lisa Jones22; (Cardiff): Christine Fraser23, Elaine K Green23, Detelina Grozeva23, Marian L Hamshere23, Peter A Holmans23, Ian R Jones23, George Kirov23, Valentina Moskivina23, Ivan Nikolov23, Michael C O’Donovan23, Michael J Owen23, Nick Craddock23; (London): David A Collier24, Amanda Elkin24, Anne Farmer24, Richard Williamson24, Peter McGuffin24; (Newcastle): Allan H Young25, I Nicol Ferrier25; Coronary Artery Disease (Leeds): Stephen G Ball26, Anthony J Balmforth26, Jennifer H Barrett26, Timothy D Bishop26, Mark M Iles26, Azhar Maqbool26, Nadira Yuldasheva26, Alistair S Hall26; (Leicester): Peter S Braund10, Paul R Burton1, Richard J Dixon10, Massimo Mangino10, Suzanne Stevens10, Martin D Tobin1, John R Thompson1, Nilesh J Samani10; Crohn’s Disease (Cambridge): Francesca Bredin27, Mark Tremelling27, Miles Parkes27; (Edinburgh): Hazel Drummond28, Charles W Lees28, Elaine R Nimmo28, Jack Satsangi28; (London): Sheila A Fisher29, Alastair Forbes30, Cathryn M Lewis29, Clive M Onnie29, Natalie J Prescott29, Jeremy Sanderson31, Christopher G Matthew29; (Newcastle): Jamie Barbour32, M Khalid Mohiuddin32, Catherine E Todhunter32, John C Mansfield32; (Oxford): Tariq Ahmad33, Fraser R Cummings33, Derek P Jewell33; Hypertension (Aberdeen): John Webster34; (Cambridge): Morris J Brown35, David G Clayton2; (Evry, France): Mark G Lathrop36; (Glasgow): John Connell37, Anna Dominiczak37; (Leicester): Nilesh J Samani10; (London): Carolina A Braga Marcano38, Beverley Burke38, Richard Dobson38, Johannie Gungadoo38, Kate L Lee38, Patricia B Munroe38, Stephen J Newhouse38, Abiodun Onipinla38, Chris Wallace38, Mingzhan Xue38, Mark Caulfield38; (Oxford): Martin Farrall39; Rheumatoid Arthritis: Anne Barton40, The Biologics in RA Genetics and Genomics Study Syndicate (BRAGGS) Steering Committee*, Ian N Bruce40, Hannah Donovan40, Steve Eyre40, Paul D Gilbert40, Samantha L Hilder40, Anne M Hinks40, Sally L John40, Catherine Potter40, Alan J Silman40, Deborah PM Symmons40, Wendy Thomson40, Jane Worthington40; Type 1 Diabetes: David G Clayton2, David B Dunger2,41, Sarah Nutland2, Helen E Stevens2, Neil M Walker2, Barry Widmer2,41, John A Todd2; Type 2 Diabetes (Exeter): Timothy M Frayling42,43, Rachel M Freathy42,43, Hana Lango42,43, John R B Perry42,43, Beverley M Shields43, Michael N Weedon42,43, Andrew T Hattersley42,43; (London): Graham A Hitman44; (Newcastle): Mark Walker45; (Oxford): Kate S Elliott3,7, Christopher J Groves7, Cecilia M Lindgren3,7, Nigel W Rayner3,7, Nicolas J Timpson3,46, Eleftheria Zeggini3,7, Mark I McCarthy3,7, Tuberculosis (Gambia): Melanie Newport47, Giorgio Sirugo47, (Oxford): Emily Lyons3, Fredrik Vannberg3, Adrian V S Hill3; Ankylosing Spondylitis: Linda A Bradbury48, Claire Farrar49, Jennifer J Pointon49, Paul Wordsworth49, Matthew A Brown48,49; Autoimmune Thyroid Disease: Jayne A Franklyn50, Joanne M Heward50, Matthew J Simmonds50, Stephen CL Gough50; Breast Cancer: Sheila Seal51, Breast Cancer Susceptibility Collaboration (UK)*, Michael R Stratton51,52, Nazneen Rahman51; Multiple Sclerosis: Maria Ban53, An Goris53, Stephen J Sawcer53, Alastair Compston53; Gambian Controls (Gambia): David Conway47, Muminatou Jallow47, Melanie Newport47, Giorgio Sirugo47; (Oxford): Kirk A Rockett3, Dominic P Kwiatkowski3,5; DNA, Genotyping, Data QC and Informatics (Wellcome Trust Sanger Institute, Hinxton): Suzannah J Bumpstead5, Amy Chaney5, Kate Downes2,5, Mohammed JR Ghori5, Rhian Gwilliam5, Sarah E Hunt5, Michael Inouye5, Andrew Keniry5, Emma King5, Ralph McGinnis5, Simon Potter5, Rathi Ravindrarajah5, Pamela Whittaker5, Claire Widden5, David Withers5, Panos Deloukas5; (Cambridge): Hin-Tak Leung2, Sarah Nutland2, Helen E Stevens2, Neil M Walker2, John A Todd2; Statistics (Cambridge): Doug Easton12, David G Clayton2; (Leicester): Paul R Burton1, Martin D Tobin1; (Oxford): Jeffrey C Barrett3, David M Evans3, Andrew P Morris3, Lon R Cardon3,55; (Oxford): Niall J Cardin11, Dan Davison11, Teresa Ferreira11, Joanne Pereira-Gale11, Ingeleif B Hallgrimsdo' ttir11, Bryan N Howie11, Jonathan L Marchini11, Chris CA Spencer11, Zhan Su11, Yik Ying Teo3,11, Damjan Vukcevic11, Peter Donnelly11; Principal Investigators: David Bentley5,54, Matthew A Brown48,49, Lon R Cardon3,55, Mark Caulfield38, David G Clayton2, Alastair Compston53, Nick Craddock23, Panos Deloukas5, Peter Donnelly11, Martin Farrall39, Stephen CL Gough50, Alistair S Hall26, Andrew T Hattersley42,43, Adrian V S Hill3, Dominic P Kwiatkowski3,5, Christopher G Matthew29, Mark I McCarthy3,7, Willem H Ouwehand8,9, Miles Parkes27, Marcus Pembrey18,20, Nazneen Rahman51, Nilesh J Samani10, Michael R Stratton51,52, John A Todd2, Jane Worthington40; AITD Replication Group: Sarah L Mitchell50, Paul R Newby50, Oliver J Brand50, Jackie Carr-Smith50, Simon H S Pearce56, Stephen C L Gough50; IL23R replication: R McGinnis5, A Keniry5, P Deloukas5, TASC. The Australo-Anglo-American Spondylitis Consortium (TASC): John D Reveille57, Xiaodong Zhou57, Linda A Bradbury58, Anne-Marie Sims58, Alison Dowling58, Jacqueline Taylor58, Tracy Doan58, Lon R Cardon55,59, John C Davis60, Jennifer J Pointon61, Laurie Savage62, Michael MWard63, Thomas L Learch64, Michael H Weisman65, Paul Wordsworth61, Matthew A Brown58,61
Author information
Authors and Affiliations
Consortia
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 769 kb)
Rights and permissions
About this article
Cite this article
Wellcome Trust Case Control Consortium., The Australo-Anglo-American Spondylitis Consortium (TASC). Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat Genet 39, 1329–1337 (2007). https://doi.org/10.1038/ng.2007.17
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2007.17
This article is cited by
-
Sequence of Events in the Pathogenesis of Axial Spondyloarthritis: A Current Review—2023 SPARTAN Meeting Proceedings
Current Rheumatology Reports (2024)
-
Zur Rolle von HLA-B27 in der Pathogenese und Diagnostik der axialen Spondyloarthritis
Zeitschrift für Rheumatologie (2024)
-
MICA and NLRP3 gene polymorphisms interact synergistically affecting the risk of ankylosing spondylitis
Immunologic Research (2024)
-
RNA sequencing and bioinformatics analysis of differentially expressed genes in the peripheral serum of ankylosing spondylitis patients
Journal of Orthopaedic Surgery and Research (2023)
-
Quantitative proteomic screening uncovers candidate diagnostic and monitoring serum biomarkers of ankylosing spondylitis
Arthritis Research & Therapy (2023)