Abstract
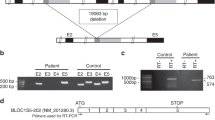
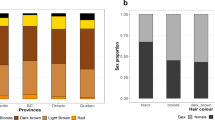
In horses, graying with age is an autosomal dominant trait associated with a high incidence of melanoma and vitiligo-like depigmentation. Here we show that the Gray phenotype is caused by a 4.6-kb duplication in intron 6 of STX17 (syntaxin-17) that constitutes a cis-acting regulatory mutation. Both STX17 and the neighboring NR4A3 gene are overexpressed in melanomas from Gray horses. Gray horses carrying a loss-of-function mutation in ASIP (agouti signaling protein) had a higher incidence of melanoma, implying that increased melanocortin-1 receptor signaling promotes melanoma development in Gray horses. The Gray horse provides a notable example of how humans have cherry-picked mutations with favorable phenotypic effects in domestic animals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sponenberg, D.P. Equine Coat Color Genetics 215 (Blackwell, Ames, Iowa, 2003).
Sutton, R.H. & Coleman, G.T. Melanoma and the Graying Horse (RIRDC Research Paper Series) 1–34 (Barton, Australia, 1997).
Fleury, C. et al. The study of cutaneous melanomas in Camargue-type gray-skinned horses (2): epidemiological survey. Pigment Cell Res. 13, 47–51 (2000).
Comfort, A. Coat-colour and longevity in thoroughbred mares. Nature 182, 1531–1532 (1958).
Seltenhammer, M.H. et al. Comparative histopathology of grey-horse-melanoma and human malignant melanoma. Pigment Cell Res. 17, 674–681 (2004).
Swinburne, J.E., Hopkins, A. & Binns, M.M. Assignment of the horse grey coat colour gene to ECA25 using whole genome scanning. Anim. Genet. 33, 338–342 (2002).
Henner, J. et al. Genetic mapping of the (G)-locus, responsible for the coat color phenotype “progressive greying with age” in horses (Equus caballus). Mamm. Genome 13, 535–537 (2002).
Locke, M.M., Penedo, M.C., Bricker, S.J., Millon, L.V. & Murray, J. Linkage of the grey coat colour locus to microsatellites on horse chromosome 25. Anim. Genet. 33, 329–337 (2002).
Pielberg, G., Mikko, S., Sandberg, K. & Andersson, L. Comparative linkage mapping of the grey coat colour gene in horses. Anim. Genet. 36, 390–395 (2005).
Rieder, S., Taourit, S., Mariat, D., Langlois, B. & Guerin, G. Mutations in the agouti (ASIP), the extension (MC1R), and the brown (TYRP1) loci and their association to coat color phenotypes in horses (Equus caballus). Mamm. Genome 12, 450–455 (2001).
Lu, D. et al. Agouti protein is an antagonist of the melanocyte-stimulating-hormone receptor. Nature 371, 799–802 (1994).
D'Orazio, J.A. et al. Topical drug rescue strategy and skin protection based on the role of Mc1r in UV-induced tanning. Nature 443, 340–344 (2006).
Marklund, L., Moller, M.J., Sandberg, K. & Andersson, L. A missense mutation in the gene for melanocyte-stimulating hormone receptor (MC1R) is associated with the chestnut coat color in horses. Mamm. Genome 7, 895–899 (1996).
Bonifacino, J.S. & Glick, B.S. The mechanisms of vesicle budding and fusion. Cell 116, 153–166 (2004).
Steegmaier, M. et al. Three novel proteins of the syntaxin/SNAP-25 family. J. Biol. Chem. 273, 34171–34179 (1998).
Zhang, Q., Li, J., Deavers, M., Abbruzzese, J.L. & Ho, L. The subcellular localization of syntaxin 17 varies among different cell types and is altered in some malignant cells. J. Histochem. Cytochem. 53, 1371–1382 (2005).
Maxwell, M.A. & Muscat, G.E. The NR4A subgroup: immediate early response genes with pleiotropic physiological roles. Nucl. Recept. Signal. 4, e002 (2006).
Nomiyama, T. et al. The NR4A orphan nuclear receptor NOR1 is induced by platelet-derived growth factor and mediates vascular smooth muscle cell proliferation. J. Biol. Chem. 281, 33467–33476 (2006).
Bailey, J.A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002).
Gray-Schopfer, V., Wellbrock, C. & Marais, R. Melanoma biology and new targeted therapy. Nature 445, 851–857 (2007).
Smith, A.G. et al. Melanocortin-1 receptor signaling markedly induces the expression of the NR4A nuclear receptor subgroup in melanocytic cells. J. Biol. Chem. 283, 12564–12570 (2008).
Dumaz, N. & Marais, R. Integrating signals between cAMP and the RAS/RAF/MEK/ERK signalling pathways. FEBS J. 272, 3491–3504 (2005).
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002).
van Dorssen, J. Über die genese der melanome in der haut bei Schimmelpferden. Inaugural dissertation, Univ. Amsterdam (1903).
Van Neste, D. & Tobin, D.J. Hair cycle and hair pigmentation: dynamic interactions and changes associated with aging. Micron 35, 193–200 (2004).
Steingrimsson, E., Copeland, N.G. & Jenkins, N.A. Melanocyte stem cell maintenance and hair graying. Cell 121, 9–12 (2005).
Nishimura, E.K., Granter, S.R. & Fisher, D.E. Mechanisms of hair graying: incomplete melanocyte stem cell maintenance in the niche. Science 307, 720–724 (2005).
Desser, H., Niebauer, G.W. & Gebhart, W. Polyamine and histamine contents in the blood of pigmented, depigmented and melanoma bearing Lipizzaner horses. Zentralbl. Veterinarmed. A 27, 45–53 (1980).
Acknowledgements
We thank H. Andersson, E.-M. Eriksson, S. Mikko and the directors of Piber, Lipica, Djakovo, Szilvasvarad and Topolcianky Lippizaner studs for valuable assistance with sample collections; T. Gunn for valuable discussions on agouti expression; U. Gustafson for expert technical assistance; D.F. Antczak for genomic DNA from Twilight; and J. Hansson (Radiumhemmet, Karolinska University Hospital) for the human melanoma cell lines. This work was supported by grants from the Swedish Cancer Society; the Olle Engkvist Foundation; the Swedish Foundation for Strategic Research; the Swedish Research Council for Environment, Agricultural Sciences and Spatial Planning; and the Uppsala Centre for Comparative Genomics.
Author information
Authors and Affiliations
Contributions
G.R.P. was responsible for marker development, positional cloning, characterization of the STX17 transcripts and real-time PCR analysis; A.G. was responsible for generation of antibodies to STX17, immunohistochemistry and northern blot analysis; E.S. was responsible for genotyping the Lipizzaner population material and analyzing allelic imbalance in melanoma tissue; I.C., M.H.S., T.D., R.B. and J.S. collected phenotypic data and blood samples from Lipizzaners; J.S. did the statistical analysis of genotype-phenotype relationships; J.L. and C.-H.H. took part in the functional characterization of STX17 and NR4A3; M.H.S. and M.V. established Gray melanoma cell lines and provided skin samples from Gray and non-Gray horses; M.B. provided samples from Gray tumors and helped isolate BAC clones; C.F. assisted with northern blot analysis; G.L. assisted with characterization of BAC clones; K.S. provided samples from Gray and non-Gray horses; S.S. and F.P. assisted with immunohistochemistry analysis; M.G., C.W. and K.L.-T. did the bioinformatics analysis of the horse genome assembly; L.A. planned the study and prepared the manuscript with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
A patent application has been filed on the basis of the data presented in this paper.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Note, Supplementary Tables 1–3, Supplementary Figures 1–3 (PDF 305 kb)
Rights and permissions
About this article
Cite this article
Rosengren Pielberg, G., Golovko, A., Sundström, E. et al. A cis-acting regulatory mutation causes premature hair graying and susceptibility to melanoma in the horse. Nat Genet 40, 1004–1009 (2008). https://doi.org/10.1038/ng.185
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.185
This article is cited by
-
Genome-wide detection of copy number variation in American mink using whole-genome sequencing
BMC Genomics (2022)
-
Genome-wide evaluation of copy gain and loss variations in three Afghan sheep breeds
Scientific Reports (2022)
-
Equine vitiligo-like depigmentation in grey horses is related to genes involved in immune response and tumor metastasis
BMC Veterinary Research (2021)
-
Beyond tradition and convention: benefits of non-traditional model organisms in cancer research
Cancer and Metastasis Reviews (2021)
-
Betulinic acid shows anticancer activity against equine melanoma cells and permeates isolated equine skin in vitro
BMC Veterinary Research (2020)