Abstract
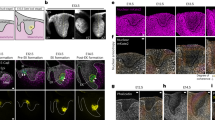
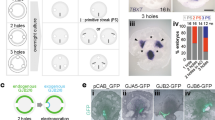
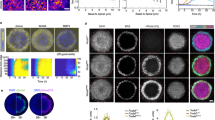
We present direct evidence of an activator-inhibitor system in the generation of the regularly spaced transverse ridges of the palate. We show that new ridges, called rugae, that are marked by stripes of expression of Shh (encoding Sonic hedgehog), appear at two growth zones where the space between previously laid rugae increases. However, inter-rugal growth is not absolutely required: new stripes of Shh expression still appeared when growth was inhibited. Furthermore, when a ruga was excised, new Shh expression appeared not at the cut edge but as bifurcating stripes branching from the neighboring stripe of Shh expression, diagnostic of a Turing-type reaction-diffusion mechanism. Genetic and inhibitor experiments identified fibroblast growth factor (FGF) and Shh as components of an activator-inhibitor pair in this system. These findings demonstrate a reaction-diffusion mechanism that is likely to be widely relevant in vertebrate development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Turing, A.M. The chemical basis of morphogenesis: a reaction-diffusion model for development. Phil. Trans. R. Soc. Lond. 237, 37–72 (1952).
Kondo, S. & Miura, T. Reaction-diffusion model as a framework for understanding biological pattern formation. Science 329, 1616–1620 (2010).
Asai, R., Taguchi, E., Kume, Y., Saito, M. & Kondo, S. Zebrafish Leopard gene as a component of the putative reaction-diffusion system. Mech. Dev. 89, 87–92 (1999).
Meinhardt, H. The Algorithmic Beauty of Seashells 4th edn. (Springer-Verlag, 2009).
Kulesa, P.M. et al. On a model mechanism for the spatial patterning of teeth primordia in the alligator. J. Theor. Biol. 180, 287–297 (1996).
Miura, T., Shiota, K., Morriss-Kay, G. & Maini, P.K. Mixed-mode pattern in Doublefoot mutant mouse limb—Turing reaction-diffusion model on a growing domain during limb development. J. Theor. Biol. 240, 562–573 (2006).
Jiang, T.X., Jung, H.S., Widelitz, R.B. & Chuong, C.M. Self-organization of periodic patterns by dissociated feather mesenchymal cells and the regulation of size, number and spacing of primordia. Development 126, 4997–5009 (1999).
Sick, S., Reinker, S., Timmer, J. & Schlake, T. WNT and DKK determine hair follicle spacing through a reaction-diffusion mechanism. Science 314, 1447–1450 (2006).
Baker, R.E., Schnell, S. & Maini, P.K. Waves and patterning in developmental biology: vertebrate segmentation and feather bud formation as case studies. Int. J. Dev. Biol. 53, 783–794 (2009).
Newman, S.A. & Bhat, R. Dynamical patterning modules: a “pattern language” for development and evolution of multicellular form. Int. J. Dev. Biol. 53, 693–705 (2009).
Goldbeter, A. & Pourquie, O. Modeling the segmentation clock as a network of coupled oscillations in the Notch, Wnt and FGF signaling pathways. J. Theor. Biol. 252, 574–585 (2008).
Axelrod, J.D. Delivering the lateral inhibition punchline: it's all about the timing. Sci. Signal. 3, pe38 (2010).
Cohen, M., Georgiou, M., Stevenson, N.L., Miodownik, M. & Baum, B. Dynamic filopodia transmit intermittent Delta-Notch signaling to drive pattern refinement during lateral inhibition. Dev. Cell 19, 78–89 (2010).
Pantalacci, S., Semon, M., Martin, A., Chevret, P. & Laudet, V. Heterochronic shifts explain variations in a sequentially developing repeated pattern: palatal ridges of muroid rodents. Evol. Dev. 11, 422–433 (2009).
Ikemi, N., Kawata, M. & Yasuda, M. All-trans-retinoic acid-induced variant patterns of palatal rugae in Crj:SD rat fetuses and their potential as indicators for teratogenicity. Reprod. Toxicol. 9, 369–377 (1995).
Pospieszny, N., Janeczek, M. & Klećkowska, J. Morphology of the incisive papilla (Papilla incisiva) of pigs during different stages of their prenatal period. EJPAU 6(1), Veterinary#04 (2003).
Pantalacci, S. et al. Patterning of palatal rugae through sequential addition reveals an anterior/posterior boundary in palatal development. BMC Dev. Biol. 8, 116 (2008).
Welsh, I.C. & O'Brien, T.P. Signaling integration in the rugae growth zone directs sequential SHH signaling center formation during the rostral outgrowth of the palate. Dev. Biol. 336, 53–67 (2009).
Rice, D.P., Rice, R. & Thesleff, I. Fgfr mRNA isoforms in craniofacial bone development. Bone 33, 14–27 (2003).
Hosokawa, R. et al. Epithelial-specific requirement of FGFR2 signaling during tooth and palate development. J. Exp. Zool. B. Mol. Dev. Evol. 312B, 343–350 (2009).
Simrick, S., Lickert, H. & Basson, M.A. Sprouty genes are essential for the normal development of epibranchial ganglia in the mouse embryo. Dev. Biol. 358, 147–155 (2011).
Porntaveetus, T., Oommen, S., Sharpe, P.T. & Ohazama, A. Expression of Fgf signalling pathway related genes during palatal rugae development in the mouse. Gene Expr. Patterns 10, 193–198 (2010).
Dassule, H.R., Lewis, P., Bei, M., Maas, R. & McMahon, A.P. Sonic hedgehog regulates growth and morphogenesis of the tooth. Development 127, 4775–4785 (2000).
Shoji, H., Iwasa, Y. & Kondo, S. Stripes, spots, or reversed spots in two-dimensional Turing systems. J. Theor. Biol. 224, 339–350 (2003).
Sinha, S. & Chen, J.K. Purmorphamine activates the Hedgehog pathway by targeting Smoothened. Nat. Chem. Biol. 2, 29–30 (2006).
Zhang, Z. et al. Rescue of cleft palate in Msx1-deficient mice by transgenic Bmp4 reveals a network of BMP and Shh signaling in the regulation of mammalian palatogenesis. Development 129, 4135–4146 (2002).
Lin, C. et al. The inductive role of Wnt–β-catenin signaling in the formation of oral apparatus. Dev. Biol. 356, 40–50 (2011).
Ishihara, S. & Kaneko, K. Turing pattern with proportion preservation. J. Theor. Biol. 238, 683–693 (2006).
Ahn, Y., Sanderson, B.W., Klein, O.D. & Krumlauf, R. Inhibition of Wnt signaling by Wise (Sostdc1) and negative feedback from Shh controls tooth number and patterning. Development 137, 3221–3231 (2010).
Sala, F.G. et al. FGF10 controls the patterning of the tracheal cartilage rings via Shh. Development 138, 273–282 (2011).
Martin, P. Tissue patterning in the developing mouse limb. Int. J. Dev. Biol. 34, 323–336 (1990).
Mootoosamy, R.C. & Dietrich, S. Distinct regulatory cascades for head and trunk myogenesis. Development 129, 573–583 (2002).
Basson, M.A. et al. Sprouty1 is a critical regulator of GDNF/RET-mediated kidney induction. Dev. Cell 8, 229–239 (2005).
Shim, K., Minowada, G., Coling, D.E. & Martin, G.R. Sprouty2, a mouse deafness gene, regulates cell fate decisions in the auditory sensory epithelium by antagonizing FGF signaling. Dev. Cell 8, 553–564 (2005).
Acknowledgements
We would like to thank A. Lander and M. Cohen for useful advice on models, G. Martin (University of California, San Francisco) for the Spry2 mutant mice, M. Kawasaki, Y. Otsuka-Tanaka and K. Kawasaki for assistance with in situ hybridization and M. Miodownik and M. Rubock for critical reading of the manuscript. This work was funded by a Medical Research Council (MRC; UK) grant (G0801154 to J.B.A.G. and M.T.C.).
Author information
Authors and Affiliations
Contributions
A.D.E. performed palate measurements and explant experiments. T.P., A.O., P.T.S., M.A.B., A.G.-L. and M.T.C. constructed and analyzed the mouse mutants. S.K. performed the modeling. A.D.E., M.T.C. and J.B.A.G. designed the explant experiments and wrote the manuscript, with contributions from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 2454 kb)
Rights and permissions
About this article
Cite this article
Economou, A., Ohazama, A., Porntaveetus, T. et al. Periodic stripe formation by a Turing mechanism operating at growth zones in the mammalian palate. Nat Genet 44, 348–351 (2012). https://doi.org/10.1038/ng.1090
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1090
This article is cited by
-
Novel Aspects in Pattern Formation Arise from Coupling Turing Reaction–Diffusion and Chemotaxis
Bulletin of Mathematical Biology (2024)
-
Turing structuring with multiple nanotwins to engineer efficient and stable catalysts for hydrogen evolution reaction
Nature Communications (2023)
-
Appearance and suppression of Turing patterns under a periodically forced feed
Communications Chemistry (2023)
-
Bat teeth illuminate the diversification of mammalian tooth classes
Nature Communications (2023)
-
“A Single and Indivisible Principle of Unity”: On Growth and Form in Context
Biological Theory (2023)