Abstract
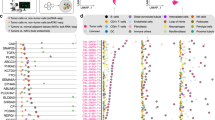
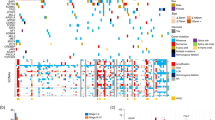
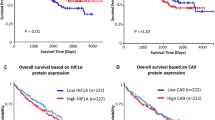
We sequenced whole exomes of ten clear cell renal cell carcinomas (ccRCCs) and performed a screen of ∼1,100 genes in 88 additional ccRCCs, from which we discovered 12 previously unidentified genes mutated at elevated frequencies in ccRCC. Notably, we detected frequent mutations in the ubiquitin-mediated proteolysis pathway (UMPP), and alterations in the UMPP were significantly associated with overexpression of HIF1α and HIF2α in the tumors (P = 0.01 and 0.04, respectively). Our findings highlight the potential contribution of UMPP to ccRCC tumorigenesis through the activation of the hypoxia regulatory network.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Rini, B.I., Campbell, S.C. & Escudier, B. Lancet 373, 1119–1132 (2009).
van Haaften, G. et al. Nat. Genet. 41, 521–523 (2009).
Dalgliesh, G.L. et al. Nature 463, 360–363 (2010).
Varela, I. et al. Nature 469, 539–542 (2011).
Gui, Y. et al. Nat. Genet. 43, 875–878 (2011).
Forbes, S.A. et al. Curr. Protoc. Hum. Genet. Chapter 10, Unit 10.11 (Wiley, 2008).
Getz, G. et al. Science 317, 1500 (2007).
Szymanska, K. et al. Cancer Lett. 293, 92–98 (2010).
Banks, R.E. et al. Cancer Res. 66, 2000–2011 (2006).
Harbour, J.W. et al. Science 330, 1410–1413 (2010).
Lamb, R.F. et al. Nat. Cell Biol. 2, 281–287 (2000).
Ding, L. et al. Nature 455, 1069–1075 (2008).
Sonoda, I. et al. Cancer Res. 64, 3741–3747 (2004).
Linehan, W.M. et al. Annu. Rev. Med. 61, 329–343 (2010).
Acknowledgements
This work was supported by the National Basic Research Program of China (973 program 2011CB809200), the National Natural Science Foundation of China (30725008, 30890032 and 30811130531), the Chinese 863 program (2006AA02A301, 2006AA02A302 and 2009AA022707), the Shenzhen Municipal Government of China (JC200903190767A, JC200903190772A, ZYC200903240076A, CXB200903110066A, ZYC200903240077A and ZYC200903240080A), the Promotion Program for Shenzhen Key Laboratory, Shenzhen, China (CXB200903090055A and CXB201005250016A) and the Biobank of Complex Diseases in Shenzhen (CXC201005260001A). This project was also funded by the Shenzhen municipal government and the local government of the Yantian District of Shenzhen.
Author information
Authors and Affiliations
Contributions
Jun Wang, Z.C., Jian Wang, H.Y., S.L. and Y. Li managed the project. Xiaokun Zhao, C. Liang, F.Z., Z.L., X. Li, L.Z., J.C., Y.W., Z.J., S. Wu, Z.Z., R. Yang, W.Y., Y. Liu, B.J., J.L. and Q.F. prepared the samples. X. Zhang, X.H., Xiao Liu, R.W., L.L., Xia Zhao, H.P. and K.K. performed the sequencing. G.G., Y.G., S.G., A.T., Y.H., W.J., M.H., S. Wan, C.C., M.J., T.J. and R. Ye performed the bioinformatic analysis. S.Y., P.S., P.H., J. Zou, F.F., Xingwang Liu and H.W. performed the validation of somatic mutations. L.S. and C. Li performed the immunohistochemistry analysis. Z.L., J.X. and J. Zhu performed the methylation analysis. G.G., Y.G., Y. Li, A.T. and Y.H. wrote the manuscript, and Y.H., Y.G., G.G., Y. Li and X.S. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–4 and Supplementary Tables 1, 2 and 6–9. (PDF 934 kb)
Supplementary Table 3
Supplementary Table 3 (XLSX 41 kb)
Supplementary Table 4
Supplementary Table 4 (XLSX 27 kb)
Supplementary Table 5
Supplementary Table 5 (XLSX 52 kb)
Rights and permissions
About this article
Cite this article
Guo, G., Gui, Y., Gao, S. et al. Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nat Genet 44, 17–19 (2012). https://doi.org/10.1038/ng.1014
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1014
This article is cited by
-
Targeting BAP1 with small compound inhibitor for colon cancer treatment
Scientific Reports (2023)
-
Hypoxia classifier for transcriptome datasets
BMC Bioinformatics (2022)
-
BAP1 promotes the repair of UV-induced DNA damage via PARP1-mediated recruitment to damage sites and control of activity and stability
Cell Death & Differentiation (2022)
-
Exploring synthetic lethal network for the precision treatment of clear cell renal cell carcinoma
Scientific Reports (2022)
-
Genomic profiling in renal cell carcinoma
Nature Reviews Nephrology (2020)