Abstract
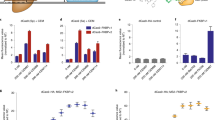
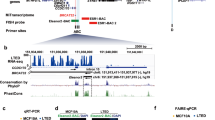
The ability to selectively activate or inhibit gene expression is fundamental to understanding complex cellular systems and developing therapeutics. Recent studies have demonstrated that duplex RNAs complementary to promoters within chromosomal DNA are potent gene silencing agents in mammalian cells. Here we report that chromosome-targeted RNAs also activate gene expression. We have identified multiple duplex RNAs complementary to the progesterone receptor (PR) promoter that increase expression of PR protein and RNA after transfection into cultured T47D or MCF7 human breast cancer cells. Upregulation of PR protein reduced expression of the downstream gene encoding cyclooygenase 2 but did not change concentrations of estrogen receptor, which demonstrates that activating RNAs can predictably manipulate physiologically relevant cellular pathways. Activation decreased over time and was sequence specific. Chromatin immunoprecipitation assays indicated that activation is accompanied by reduced acetylation at histones H3K9 and H3K14 and by increased di- and trimethylation at histone H3K4. These data show that, like proteins, hormones and small molecules, small duplex RNAs interact at promoters and can activate or repress gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Braasch, D.A. & Corey, D.R. Novel antisense strategies for controlling gene expression. Biochemistry 41, 4503–4510 (2002).
Eckstein, F. Small noncoding RNAs as magic bullets. Trends Biochem. Sci. 30, 445–452 (2005).
Dykxhoorn, D.M., Palliser, D. & Lieberman, J. The silent treatment: siRNAs as small molecule drugs. Gene Ther. 13, 541–552 (2006).
Arora, P.S., Ansari, A.Z., Best, T.P., Ptashne, M. & Dervan, P.B. Design of artificial transcriptional activators with rigid poly-L-proline linkers. J. Am. Chem. Soc. 124, 13067–13071 (2002).
Kwon, Y. et al. Small molecule transcription factor mimic. J. Am. Chem. Soc. 126, 15940–15941 (2004).
Liu, B., Han, Y., Ferdous, A., Corey, D.R. & Kodadek, T. Transcription activation by a PNA–peptide chimera in a mammalian cell extract. Chem. Biol. 10, 909–916 (2003).
Majmudar, C.Y. & Mapp, A.K. Chemical approaches to transcriptional regulation. Curr. Opin. Chem. Biol. 9, 467–474 (2005).
Janowski, B.A. et al. Inhibition of gene expression at transcription start sites using antigene RNAs (agRNAs). Nat. Chem. Biol. 1, 216–222 (2005).
Janowski, B.A. et al. Involvement of Ago1 and Ago2 in mammalian transcriptional silencing. Nat. Struct. Mol. Biol. 13, 787–792 (2006).
Janowski, B.A., Hu, J. & Corey, D.R. Antigene inhibition by peptide nucleic acids and duplex RNAs. Nat. Protoc. 1, 436–443 (2006).
Morris, K.V., Chan, S.W., Jacobsen, S.E. & Looney, D.J. Small interfering RNA–induced transcriptional silencing in human cells. Science 305, 1289–1292 (2004).
Ting, A.H., Schuebel, K.E., Herman, J.G. & Baylin, S.B. Short double-stranded RNA induces transcriptional gene silencing in human cancer cells in the absence of DNA methylation. Nat. Genet. 37, 906–910 (2005).
Suzuki, K. et al. Prolonged transcriptional silencing and CpG methylation induced by siRNAs targeted to the HIV-1 promoter region. J. RNAi Gene Silencing 1, 66–78 (2005).
Zhang, M.-X. et al. Regulation of endothelial nitric oxide synthase by small RNA. Proc. Natl. Acad. Sci. USA 102, 16967–16972 (2005).
Kim, D.H. et al. Argonaute-1 directs siRNA-mediated transcriptional gene silencing in human cells. Nat. Struct. Mol. Biol. 13, 793–797 (2006).
Corey, D.R. Regulating mammalian transcription with RNA. Trends Biochem. Sci. 30, 655–658 (2005).
Morris, K.V. siRNA-mediated transcriptional gene silencing: the potential mechanism and a possible role in the histone code. Cell. Mol. Life Sci. 62, 3057–3066 (2005).
Kastner, P. et al. Two distinct estrogen-regulated promoters generate transcripts encoding the two functionally different human progesterone receptor isoforms A and B. EMBO J. 9, 1603–1614 (1990).
Misrahi, M. et al. Structure of the human progesterone receptor gene. Biochim. Biophys. Acta 1216, 289–292 (1993).
Jenster, G. et al. Steroid receptor induction of gene transcription: a two-step model. Proc. Natl. Acad. Sci. USA 94, 7879–7884 (1997).
Hurd, C. et al. Hormonal regulation of the p53 tumor suppressor protein in T47D human breast carcinoma cell line. J. Biol. Chem. 270, 28507–28510 (1995).
Janowski, B.A. et al. Inhibiting transcription of chromosomal DNA using antigene peptide nucleic acids. Nat. Chem. Biol. 1, 210–215 (2005).
Conneely, O.M., Jericevic, B.M. & Lydon, J.P. Progesterone receptors in mammary gland development and tumorigenesis. J. Mammary Gland Biol. Neoplasia 8, 205–214 (2003).
Reynolds, A. et al. Rational siRNA design for RNA interference. Nat. Biotechnol. 22, 326–330 (2004).
Huffman, K.E. & Corey, D.R. Inhibition of expression of major vault protein does not alter chemoresistance or drug localization in cancer cells. Biochemistry 44, 2253–2261 (2005).
Hardy, D.B., Janowski, B.A., Corey, D.R. & Mendelson, C.R. Progesterone receptor plays a major antiinflammatory role in human myometrial cells by antagonism of nuclear factor-κB activation of cyclooxygenase 2 expression. Mol. Endocrinol. 20, 2724–2733 (2006).
Read, L.D., Snider, C.E., Miller, J.S., Greene, G.L. & Katzenellenbogen, B.S. Ligand-modulated regulation of progesterone receptor messenger ribonucleic acid and protein in human breast cancer cell lines. Mol. Endocrinol. 2, 263–271 (1988).
Cho, H., Aronica, S.M. & Katzenellenbogen, B. Regulation of progesterone receptor gene expression in MCF-7 breast cancer cells: a comparison of the effects of cyclic adenosine 3′,5′-monophhosphate, estradiol, insulin-like growth factor-I and serum factors. Endocrinology 134, 658–664 (1994).
Alexander, I.E., Clarke, C.L., Shine, J. & Sutherland, R.L. Progestin inhibition of progesterone receptor gene expression in human breast cancer cells. Mol. Endocrinol. 3, 1377–1386 (1989).
Margueron, R., Trojer, P. & Reinberg, D. The key to development: interpreting the histone code. Curr. Opin. Genet. Dev. 15, 163–176 (2005).
Miao, F., Gonzalo, I.G., Lanting, L. & Natarajan, R. In vivo chromatin remodeling events leading to inflammatory gene transcription under diabetic conditions. J. Biol. Chem. 279, 18091–18097 (2004).
Ruh, M.F., Tian, S., Cox, L.K. & Ruh, T.S. The effect of histone acetylation on estrogen responsiveness in MCF-7 cells. Endocrine 11, 157–164 (1999).
Song, M.-R. & Ghosh, A. FGF2-induced chromatin remodelling regulates CNTF-mediated gene expression and astrocyte differentiation. Nat. Neurosci. 7, 229–235 (2004).
Williams-Ashman, H.G., Seidenfeld, J. & Galletti, P. Trends in the biochemical pharmacology of 5′-deoxy-5′-methylthioadenosine. Biochem. Pharmacol. 31, 277–288 (1982).
Chau, C.M. & Lieberman, P.M. Dynamic chromatin boundaries delineate a latency control region of Epstein Barr virus. J. Virol. 78, 12308–12319 (2004).
Santos-Rosa, H. et al. Active genes are trimethylated at K4 of histone H3. Nature 419, 407–411 (2002).
Strahl, B.D., Ohba, R., Cook, R.G. & Allis, C.D. Methylation of histone H3 at lysine 4 is highly conserved and correlates with transcriptionally active nuclei in Tetrahymena. Proc. Natl. Acad. Sci. USA 96, 14967–14972 (1999).
Schneider, R. et al. Histone H3 lysine 4 methylation patterns in higher eukaryotic genes. Nat. Cell Biol. 6, 73–77 (2004).
Roh, T.Y., Cuddapah, S., Cui, K. & Zhao, K. The genomic landscape of histone modifications in human T cells. Proc. Natl. Acad. Sci. USA 103, 15782–15787 (2006).
Birmingham, A. et al. 3′ UTR see matches, but not overall identity, are associated with RNAi off targets. Nat. Methods 3, 199–204 (2006).
Wassenegger, M. et al. RNA-directed de novo methylation of genomic sequences in plants. Cell 76, 567–576 (1994).
Badia, E. et al. Long-term hydroxytamoxifen treatment of an MCF-7–derived breast cancer cell line irreversibly inhibits the expression of estrogenic genes through chromatin remodelling. Cancer Res. 60, 4130–4138 (2000).
Kuwabara, T. et al. A small modulatory dsRNA specifies the fate of adult neural stem cells. Cell 116, 779–793 (2004).
Li, L.C. et al. Small dsRNAs induce transcriptional activation in human cells. Proc. Natl. Acad. Sci. USA 103, 17337–17342 (2006).
Acknowledgements
We thank N.-B. Nguyen for skilled assistance. This work was supported by the US National Institutes of Health (NIGMS 60642 and 73042 to D.R.C., CA 10151 to K.E.H. and HD011149 for D.B.H.), the Susan G. Komen Breast Cancer Foundation (PDF0600877 to D.B.H.) and the Robert A. Welch Foundation (I–1244 to D.R.C.). We thank J. Schwartz, C. Mendelson and D. Shames for their helpful comments.
Author information
Authors and Affiliations
Contributions
B.A.J., S.T.Y., D.B.H., R.R. and K.E.H. designed and performed experiments. B.A.J. and D.R.C. supervised experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Increased expression of PR upon addition of selected RNAs in T47D breast cancer cells. (PDF 100 kb)
Supplementary Fig. 2
Reproducibility of probing the PR or MVP promoters with duplex RNAs. (PDF 162 kb)
Supplementary Fig. 3
Treatment of MCF7 cells with TSA yields increased COX-2 expression. (PDF 70 kb)
Supplementary Fig. 4
Contiguous and noncontiguous sequence similarity results (and common BLAST results) of PR11 and PR22. (PDF 125 kb)
Rights and permissions
About this article
Cite this article
Janowski, B., Younger, S., Hardy, D. et al. Activating gene expression in mammalian cells with promoter-targeted duplex RNAs. Nat Chem Biol 3, 166–173 (2007). https://doi.org/10.1038/nchembio860
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio860
This article is cited by
-
DHX9/DNA-tandem repeat-dependent downregulation of ciRNA-Fmn1 in the dorsal horn is required for neuropathic pain
Acta Pharmacologica Sinica (2023)
-
RNA activation in ticks
Scientific Reports (2023)
-
RNA-based translation activators for targeted gene upregulation
Nature Communications (2023)
-
Defining an optimal control for RNAi experiments with adult Schistosoma mansoni
Scientific Reports (2023)
-
Extracellular vesicle-derived miRNA as a novel regulatory system for bi-directional communication in gut-brain-microbiota axis
Journal of Translational Medicine (2021)