Abstract
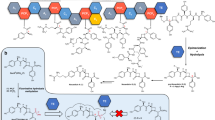
A previously determined crystal structure of the ternary complex of trehalose-6-phosphate synthase identified a putative transition state–like arrangement based on validoxylamine A 6′-O-phosphate and uridine diphosphate in the active site. Here linear free energy relationships confirm that these inhibitors are synergistic transition state mimics, supporting front-face nucleophilic attack involving hydrogen bonding between leaving group and nucleophile. Kinetic isotope effects indicate a highly dissociative oxocarbenium ion–like transition state. Leaving group 18O effects identified isotopically sensitive bond cleavages and support the existence of a hydrogen bond between the nucleophile and departing group. Brønsted analysis of nucleophiles and Taft analysis highlight participation of the nucleophile in the transition state, also consistent with a front-face mechanism. Together, these comprehensive, quantitative data substantiate this unusual enzymatic reaction mechanism. Its discovery should prompt useful reassessment of many biocatalysts and their substrates and inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lairson, L.L., Henrissat, B., Davies, G.J. & Withers, S.G. Glycosyltransferases: structures, functions and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008).
Kim, S.C., Singh, A.N. & Raushel, F.M. Analysis of the galactosyltransferase reaction by positional isotope exchange and secondary deuterium isotope effects. Arch. Biochem. Biophys. 267, 54–59 (1988).
Bruner, M. & Horenstein, B.A. Isotope trapping and kinetic isotope effect studies of rat liver α-(2–6)-sialyltransferase. Biochemistry 37, 289–297 (1998).
Vocadlo, D.J., Davies, G.J., Laine, R. & Withers, S.G. Catalysis by hen egg-white lysozyme proceeds via a covalent intermediate. Nature 412, 835–838 (2001).
Lairson, L.L. et al. Intermediate trapping on a mutant retaining alpha-galactosyltransferase identifies an unexpected aspartate residue. J. Biol. Chem. 279, 28339–28344 (2004).
Soya, N., Fang, Y., Palcic, M.M. & Klassen, J.S. Trapping and characterization of covalent intermediates of mutant retaining glycosyltransferases. Glycobiology 21, 547–552 (2011).
Monegal, A. & Planas, A. Chemical rescue of α3-galactosyltransferase. implications in the mechanism of retaining glycosyltransferase. J. Am. Chem. Soc. 128, 16030–16031 (2006).
Ly, H.D., Lougheed, B., Wakarchuk, W.W. & Withers, S.G. Mechanistic studies of a retaining α-galactosyltransferase from Neisseria meningitidis. Biochemistry 41, 5075–5085 (2002).
Gibson, R.P., Turkenburg, J.P., Charnock, S.J., Lloyd, R. & Davies, G.J. Insights into trehalose synthesis provided by the structure of the retaining glucosyltransferase OtsA. Chem. Biol. 9, 1337–1346 (2002).
Persson, K. et al. Crystal structure of the retaining galactosyltransferase LgtC from Neisseria meningitidis in complex with donor and acceptor analogs. Nat. Struct. Biol. 8, 166–175 (2001).
Gastinel, L.N. et al. Bovine α-1,3-galactosyltransferase catalytic domain structure and its relationship with ABO histo-blood group and glycosphingolipid glycosyltransferase. EMBO J. 20, 638–649 (2001).
Patenaude, S.I. et al. The structural basis for specificity in human ABO(H) blood group biosynthesis. Nat. Struct. Biol. 9, 685–690 (2002).
Martinez-Fleites, C. et al. Insights into the synthesis of lipopolysaccharide and antibiotics through the structures of two retaining glycosyltransferases from family GT4. Chem. Biol. 13, 1143–1152 (2006).
Jamaluddin, H., Tumbale, P., Withers, S.G., Acharya, K.R. & Brew, K. Conformational changes induced by binding UDP-2F-galactose to alpha-1,3-galactosyltransferase—implications for catalysis. J. Mol. Biol. 369, 1270–1281 (2007).
Vetting, M.W., Frantom, P.A. & Blanchard, J.S. Structural and enzymatic analysis of MshA from Corynebacterium glutamicum. J. Biol. Chem. 283, 15834–15844 (2008).
Cowdrey, W.A., Hughes, E.D., Ingold, C.K., Masterman, S. & Scott, A.D. 257. Reaction kinetics and the Walden inversion. Part VI. Relation of steric orientation to mechanism in substitutions involving halogen atoms and simple or substituted hydroxyl groups. J. Chem. Soc. 1252–1271 (1937).
Sinnott, M.L. & Jencks, W.P. Solvolysis of D-glucopyranosyl derivatives in mixtures of ethanol and 2,2,2-trifluoroethanol. J. Am. Chem. Soc. 102, 2026–2032 (1980).
Tvaroska, I. Molecular modeling of retaining glycosyltransferases in NMR Spectroscopy and Computer Modeling of Carbohydrates (eds. Vliegenthart, J.F.G. et al.) 285–301 (American Chemical Society, 2006).
Nidetzky, B. & Eis, C. α-Retaining glucosyl transfer catalysed by trehalose phosphorylase from Schizophyllum commune: mechanistic evidence obtained from steady-state kinetic studies with substrate analogues and inhibitors. Biochem. J. 360, 727–736 (2001).
Goedl, C., Griessler, R., Schwarz, A. & Nidetzky, B. Structure-function relationships for Schizophyllum commune trehalose phosphorylase and their implications for the catalytic mechanism of family GT-4 glycosyltransferase. Biochem. J. 397, 491–500 (2006).
Goedl, C. & Nidetzky, B. Sucrose phosphorylase harboring a redesigned, glycosyltransferase-like active site exhibits retaining glucosyl transfer in the absence of a covalent intermediate. ChemBioChem 10, 2333–2337 (2009).
Errey, J.C. et al. Mechanistic insight into enzymatic glycosyl transfer with retention of configuration through analysis of glycomimetic inhibitors. Angew. Chem. Int. Ed. Engl. 49, 1234–1237 (2010).
Coutinho, P.M., Deleury, E., Davies, G.J. & Henrissat, B. An evolving hierarchical family classification for glycosyltransferases. J. Mol. Biol. 328, 307–317 (2003).
Greig, I.R. The analysis of enzymic free energy relationships using kinetic and computational models. Chem. Soc. Rev. 39, 2272–2301 (2010).
Mader, M.M. & Bartlett, P.A. Binding energy and catalysis: the implications for transition-state analogs and catalytic antibodies. Chem. Rev. 97, 1281–1302 (1997).
Gosselin, S., Alhussaini, M., Streiff, M.B., Takabayashi, K. & Palcic, M.M. A continuous spectrophotometric assay for glycosyltransferases. Anal. Biochem. 220, 92–97 (1994).
Baxter, N.J. et al. A Trojan horse transition state analogue generated by MgF3− formation in an enzyme active site. Proc. Natl. Acad. Sci. USA 103, 14732–14737 (2006).
Berti, P.J. & McCann, J.A.B. Toward a detailed understanding of base excision repair enzymes: transition state and mechanistic analyses of N-glycoside hydrolysis and N-glycosyl transfer. Chem. Rev. 106, 506–555 (2006).
Northrop, D.B. Minimal kinetic mechanism and general equation for deuterium isotope effects on enzymic reactions: uncertaintly in detecting a rate-limiting step. Biochemistry 20, 4056–4061 (1981).
Werner, R.M. & Stivers, J.T. Kinetic isotope effects studies of the reaction catalyzed by uracil DNA glycosylase: evidence for an oxocarbenium ion-uracil anion intermediate. Biochemistry 39, 14054–14064 (2000).
Luo, M. & Schramm, V.L. Transition state structure of E. coli tRNA-specific adenosine deaminase. J. Am. Chem. Soc. 130, 2649–2655 (2008).
Cleland, W.W. The use of isotope effects to determine enzyme mechanisms. Arch. Biochem. Biophys. 433, 2–12 (2005).
Lee, J.K., Bain, D.A. & Berti, P.J. Probing the transition states of four glucoside hydrolyses with 13C kinetic isotope effects measured at natural abundance by NMR spectroscopy. J. Am. Chem. Soc. 126, 3769–3776 (2004).
Glad, S.S. & Jensen, F. Transition state looseness and α-secondary kinetic isotope effects. J. Am. Chem. Soc. 119, 227–232 (1997).
Sunko, D.E., Szele, I. & Hehre, W.J. Hyperconjugation and the angular dependence of β-deuterium isotope effects. J. Am. Chem. Soc. 99, 5000–5005 (1977).
Guthrie, R.D. & Jencks, W.P. IUPAC recommendations for the representation of reaction mechanisms. Acc. Chem. Res. 22, 343–349 (1989).
Huang, X., Tanaka, K.S.E. & Bennet, A.J. Glucosidase-catalyzed hydrolysis of α-D-glucopyranosyl pyridinium salts: kinetic evidence for nucleophilic involvement at the glucosidation transition state. J. Am. Chem. Soc. 119, 11147–11154 (1997).
Chan, J., Lewis, A.R., Gilbert, M., Karwaski, M.-F. & Bennet, A.J. A direct NMR method for the measurement of competitive kinetic isotope effects. Nat. Chem. Biol. 6, 405–407 (2010).
Rosenberg, S. & Kirsch, J.F. Oxygen-18 leaving group kinetic isotope effects on the hydrolysis of nitrophenyl glycosides. 2. Lysozyme and beta-glucosidase: acid and alkaline hydrolysis. Biochemistry 20, 3196–3204 (1981).
Bennet, A.J. & Sinnott, M.L. Complete kinetic isotope effect description of transition states for acid-catalyzed hydrolyses of methyl alpha- and beta-glucopyranosides. J. Am. Chem. Soc. 108, 7287–7294 (1986).
Indurugalla, D. & Bennet, A.J. A kinetic isotope effect study on the hydrolysis reactions of methyl xylopyranosides and methyl 5-thioxylopyranosides: oxygen versus sulfur stabilization of carbenium ions. J. Am. Chem. Soc. 123, 10889–10898 (2001).
Du, X., Black, G.E., Lecchi, P., Abramson, F.P. & Sprang, S.R. Kinetic isotope effects in Ras-catalyzed GTP hydrolysis: evidence for a loose transition state. Proc. Natl. Acad. Sci. USA 101, 8858–8863 (2004).
Chenault, H.K., Mandes, R.F. & Hornberger, K.R. Synthetic utility of yeast hexokinase. Substrate specificity, cofactor regeneration, and product isolation. J. Org. Chem. 62, 331–336 (1997).
Hansch, C. & Leo, A. Substituent Constants for Correlation Analysis in Chemistry and Biology (John Wiley & Sons, New York, 1979).
Gibson, R.P., Tarling, C.A., Roberts, S., Withers, S.G. & Davies, G.J. The donor subsite of trehalose-6-phosphate synthase. J. Biol. Chem. 279, 1950–1955 (2004).
Gloster, T.M. & Davies, G.J. Glycosidase inhibition: assessing mimicry of the transition state. Org. Biomol. Chem. 8, 305–320 (2010).
Sinnott, M.L. Catalytic mechanism of enzymic glycosyl transfer. Chem. Rev. 90, 1171–1202 (1990).
Ye, J.-D., Li, N.-S., Dai, Q. & Piccirilli, J.A. The mechanism of RNA strand scission: an experiemtnal measure of the Bronsted coefficient, βnuc . Angew. Chem. Int. Ed. Engl. 46, 3714–3717 (2007).
Jones, C.S. & Kosman, D.J. Purification, properties, kinetics, and mechanism of β-N-acetylglucosaminidase from Aspergillus niger. J. Biol. Chem. 255, 11861–11869 (1980).
Macauley, M.S., Whitworth, G.E., Debowski, A.W., Chin, D. & Vocadlo, D.J. O-GlcNAcase uses substrate-assisted catalysis. J. Biol. Chem. 280, 25313–25322 (2005).
Acknowledgements
We thank the Bill and Melinda Gates Foundation, the Samsung Fellowship and the Biotechnology and Biological Sciences Research Council for financial support. R. Zhang and S.G. Withers (University of British Columbia) are acknowledged for providing 2-deoxy-2,2-difluoroglucose; R. Gibson (currently at the University of Liverpool) is thanked for initial construction of some OtsA mutants. B.G.D. and G.J.D. are both Royal Society Wolfson Research Merit Award recipients, and B.G.D. is supported by an Engineering and Physical Sciences Research Council Life Sciences Interface Platform grant.
Author information
Authors and Affiliations
Contributions
S.S.L. and B.G.D. designed the experiments. A.I. provided mutant otsA plasmids. S.S.L., S.Y.H. and J.C.E. expressed the mutants. S.S.L. and J.C.E. performed the kinetic measurements. S.S.L. performed all other experiments. S.S.L., B.G.D. and G.J.D. analyzed the experiments. B.G.D., S.S.L. and G.J.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–3, Tables 1 and 3, Supplementary Methods and Supplementary Results (PDF 2109 kb)
Rights and permissions
About this article
Cite this article
Lee, S., Hong, S., Errey, J. et al. Mechanistic evidence for a front-side, SNi-type reaction in a retaining glycosyltransferase. Nat Chem Biol 7, 631–638 (2011). https://doi.org/10.1038/nchembio.628
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.628
This article is cited by
-
The retaining β-Kdo glycosyltransferase WbbB uses a double-displacement mechanism with an intermediate adduct rearrangement step
Nature Communications (2022)
-
Inhibition of Clostridium difficile TcdA and TcdB toxins with transition state analogues
Nature Communications (2021)
-
Palladium-mediated enzyme activation suggests multiphase initiation of glycogenesis
Nature (2018)
-
A front-face 'SNi synthase' engineered from a retaining 'double-SN2' hydrolase
Nature Chemical Biology (2017)
-
Notch-modifying xylosyltransferase structures support an SNi-like retaining mechanism
Nature Chemical Biology (2015)