Abstract
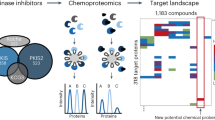
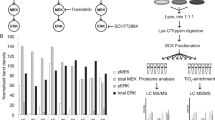
We describe a strategy for comprehending signaling pathways that are active in lung cancer cells and that are targeted by dasatinib using chemical proteomics to identify direct interacting proteins combined with immunoaffinity purification of tyrosine-phosphorylated peptides corresponding to activated tyrosine kinases. We identified nearly 40 different kinase targets of dasatinib. These include SRC-family kinase (SFK) members (LYN, SRC, FYN, LCK and YES), nonreceptor tyrosine kinases (FRK, BRK and ACK) and receptor tyrosine kinases (Ephrin receptors, DDR1 and EGFR). Using quantitative phosphoproteomics, we identified peptides corresponding to autophosphorylation sites of these tyrosine kinases that are inhibited in a concentration-dependent manner by dasatinib. Using drug-resistant gatekeeper mutants, we show that SFKs (particularly SRC and FYN), as well as EGFR, are relevant targets for dasatinib action. The combined mass spectrometry–based approach described here provides a system-level view of dasatinib action in cancer cells and suggests both functional targets and a rationale for combinatorial therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Blume-Jensen, P. & Hunter, T. Oncogenic kinase signalling. Nature 411, 355–365 (2001).
Paez, J.G. et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304, 1497–1500 (2004).
Lynch, T.J. et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 350, 2129–2139 (2004).
Puri, N. et al. A selective small molecule inhibitor of c-Met, PHA665752, inhibits tumorigenicity and angiogenesis in mouse lung cancer xenografts. Cancer Res. 67, 3529–3534 (2007).
Song, L. et al. Dasatinib (BMS-354825) selectively induces apoptosis in lung cancer cells dependent on epidermal growth factor receptor signaling for survival. Cancer Res. 66, 5542–5548 (2006).
Pao, W. et al. EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc. Natl. Acad. Sci. USA 101, 13306–13311 (2004).
Kim, L.C., Song, L. & Haura, E.B. Src kinases as therapeutic targets for cancer. Nat. Rev. Clin. Oncol. 6, 587–595 (2009).
Yeatman, T.J. A renaissance for SRC. Nat. Rev. Cancer 4, 470–480 (2004).
Ishizawar, R. & Parsons, S.J. c-Src and cooperating partners in human cancer. Cancer Cell 6, 209–214 (2004).
Tice, D.A., Biscardi, J.S., Nickles, A.L. & Parsons, S.J. Mechanism of biological synergy between cellular Src and epidermal growth factor receptor. Proc. Natl. Acad. Sci. USA 96, 1415–1420 (1999).
Maa, M.C., Leu, T.H., McCarley, D.J., Schatzman, R.C. & Parsons, S.J. Potentiation of epidermal growth factor receptor-mediated oncogenesis by c-Src: implications for the etiology of multiple human cancers. Proc. Natl. Acad. Sci. USA 92, 6981–6985 (1995).
Fu, Y.N. et al. EGFR mutants found in non-small cell lung cancer show different levels of sensitivity to suppression of Src: implications in targeting therapy. Oncogene 27, 957–965 (2008).
McDermott, U. et al. Identification of genotype-correlated sensitivity to selective kinase inhibitors by using high-throughput tumor cell line profiling. Proc. Natl. Acad. Sci. USA 104, 19936–19941 (2007).
Zhang, Q. et al. SRC family kinases mediate epidermal growth factor receptor ligand cleavage, proliferation, and invasion of head and neck cancer cells. Cancer Res. 64, 6166–6173 (2004).
Zhang, J. et al. SRC-family kinases are activated in non-small cell lung cancer and promote the survival of epidermal growth factor receptor-dependent cell lines. Am. J. Pathol. 170, 366–376 (2007).
Margolis, B. et al. High-efficiency expression/cloning of epidermal growth factor-receptor-binding proteins with Src homology 2 domains. Proc. Natl. Acad. Sci. USA 89, 8894–8898 (1992).
Osherov, N. & Levitzki, A. Epidermal-growth-factor-dependent activation of the src-family kinases. Eur. J. Biochem. 225, 1047–1053 (1994).
Martin, G.S. The hunting of the Src. Nat. Rev. Mol. Cell Biol. 2, 467–475 (2001).
Fabian, M.A. et al. A small molecule-kinase interaction map for clinical kinase inhibitors. Nat. Biotechnol. 23, 329–336 (2005).
Karaman, M.W. et al. A quantitative analysis of kinase inhibitor selectivity. Nat. Biotechnol. 26, 127–132 (2008).
Bantscheff, M. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat. Biotechnol. 25, 1035–1044 (2007).
Rix, U. et al. Chemical proteomic profiles of the BCR-ABL inhibitors imatinib, nilotinib, and dasatinib reveal novel kinase and nonkinase targets. Blood 110, 4055–4063 (2007).
Peri, S. et al. Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res. 13, 2363–2371 (2003).
Rix, U. & Superti-Furga, G. Target profiling of small molecules by chemical proteomics. Nat. Chem. Biol. 5, 616–624 (2009).
Hantschel, O. et al. The Btk tyrosine kinase is a major target of the Bcr-Abl inhibitor dasatinib. Proc. Natl. Acad. Sci. USA 104, 13283–13288 (2007).
Legate, K.R., Montanez, E., Kudlacek, O. & Fassler, R. ILK, PINCH and parvin: the tIPP of integrin signalling. Nat. Rev. Mol. Cell Biol. 7, 20–31 (2006).
Heroult, M., Schaffner, F. & Augustin, H.G. Eph receptor and ephrin ligand-mediated interactions during angiogenesis and tumor progression. Exp. Cell Res. 312, 642–650 (2006).
Ford, C.E. et al. Expression and mutation analysis of the discoidin domain receptors 1 and 2 in non-small cell lung carcinoma. Br. J. Cancer 96, 808–814 (2007).
Rikova, K. et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131, 1190–1203 (2007).
Stover, D.R., Becker, M., Liebetanz, J. & Lydon, N.B. Src phosphorylation of the epidermal growth factor receptor at novel sites mediates receptor interaction with Src and P85 alpha. J. Biol. Chem. 270, 15591–15597 (1995).
Mueller, K.L., Hunter, L.A., Ethier, S.P. & Boerner, J.L. Met and c-Src cooperate to compensate for loss of epidermal growth factor receptor kinase activity in breast cancer cells. Cancer Res. 68, 3314–3322 (2008).
Kamalati, T., Jolin, H.E., Fry, M.J. & Crompton, M.R. Expression of the BRK tyrosine kinase in mammary epithelial cells enhances the coupling of EGF signalling to PI 3-kinase and Akt, via erbB3 phosphorylation. Oncogene 19, 5471–5476 (2000).
Ostrander, J.H., Daniel, A.R., Lofgren, K., Kleer, C.G. & Lange, C.A. Breast tumor kinase (protein tyrosine kinase 6) regulates heregulin-induced activation of ERK5 and p38 MAP kinases in breast cancer cells. Cancer Res. 67, 4199–4209 (2007).
Shen, F., Lin, Q., Gu, Y., Childress, C. & Yang, W. Activated Cdc42-associated kinase 1 is a component of EGF receptor signaling complex and regulates EGF receptor degradation. Mol. Biol. Cell 18, 732–742 (2007).
Nur-E-Kamal, A. et al. Requirement of activated Cdc42-associated kinase for survival of v-Ras-transformed mammalian cells. Mol. Cancer Res. 3, 297–305 (2005).
Park, S.I. et al. Targeting SRC family kinases inhibits growth and lymph node metastases of prostate cancer in an orthotopic nude mouse model. Cancer Res. 68, 3323–3333 (2008).
Du, J. et al. Bead-based profiling of tyrosine kinase phosphorylation identifies SRC as a potential target for glioblastoma therapy. Nat. Biotechnol. 27, 77–83 (2009).
Remsing Rix, L.L. et al. Global target profile of the kinase inhibitor bosutinib in primary chronic myeloid leukemia cells. Leukemia 23, 477–485 (2009).
Gandhi, J. et al. Alterations in genes of the EGFR signaling pathway and their relationship to EGFR tyrosine kinase inhibitor sensitivity in lung cancer cell lines. PLoS One 4, e4576 (2009).
Pao, W. et al. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med. 2, e73 (2005).
Kobayashi, S. et al. Transcriptional profiling identifies cyclin D1 as a critical downstream effector of mutant epidermal growth factor receptor signaling. Cancer Res. 66, 11389–11398 (2006).
Yang, L. et al. Akt/protein kinase B signaling inhibitor-2, a selective small molecule inhibitor of Akt signaling with antitumor activity in cancer cells overexpressing Akt. Cancer Res. 64, 4394–4399 (2004).
van der Horst, E.H. et al. Metastatic properties and genomic amplification of the tyrosine kinase gene ACK1. Proc. Natl. Acad. Sci. USA 102, 15901–15906 (2005).
Yokoyama, N. & Miller, W.T. Biochemical properties of the Cdc42-associated tyrosine kinase ACK1. Substrate specificity, authphosphorylation, and interaction with Hck. J. Biol. Chem. 278, 47713–47723 (2003).
Satoh, T., Kato, J., Nishida, K. & Kaziro, Y. Tyrosine phosphorylation of ACK in response to temperature shift-down, hyperosmotic shock, and epidermal growth factor stimulation. FEBS Lett. 386, 230–234 (1996).
Yang, W., Lin, Q., Zhao, J., Guan, J.L. & Cerione, R.A. The nonreceptor tyrosine kinase ACK2, a specific target for Cdc42 and a negative regulator of cell growth and focal adhesion complexes. J. Biol. Chem. 276, 43987–43993 (2001).
Coker, K.J., Staros, J.V. & Guyer, C.A. A kinase-negative epidermal growth factor receptor that retains the capacity to stimulate DNA synthesis. Proc. Natl. Acad. Sci. USA 91, 6967–6971 (1994).
Bandyopadhyay, A. et al. Inhibition of pulmonary and skeletal metastasis by a transforming growth factor-beta type I receptor kinase inhibitor. Cancer Res. 66, 6714–6721 (2006).
Takanami, I. Increased expression of integrin-linked kinase is associated with shorter survival in non-small cell lung cancer. BMC Cancer 5, 1 (2005).
Shah, N.P. et al. Transient potent BCR-ABL inhibition is sufficient to commit chronic myeloid leukemia cells irreversibly to apoptosis. Cancer Cell 14, 485–493 (2008).
Acknowledgements
We thank F. Lee (Bristol-Myers Squibb Oncology) for providing dasatinib, Genentech and OSI Pharmaceuticals for providing erlotinib, K. Rikova (Cell Signaling Technology) for providing NSCLC tumor spectral count data, Cell Signaling Technology for allowing reproduction of the kinome map, G. Durnberger for data uploading to PRIDE, J. Du (Broad Institute) and T. Golub (Broad Institute) for providing lentiviral constructs, T. Yoshida for helpful discussions and P. Johnston for administrative assistance. We thank W. Pao (Vanderbilt University) for the cDNA for L858R EGFR and L858R T790M EGFR and N. Mahajan (Moffitt Cancer Center) for the ACK. We thank J. Cheng (Moffitt Cancer Center) for the triciribine. The work was partially funded by grants from the US National Functional Genomics Center, the Moffitt Cancer Center SPORE in Lung Cancer (P50-CA119997), Joan's Legacy/LUNGevity Foundation (E.B.H.), the Austrian Federal Ministry of Science and Research within the GEN-AU program (GZ200.142/1-VI/I/2006 and GZ 200.145/1-VI/1/2006) and the Austrian Academy of Sciences. This work has been supported in part by the Proteomics Core, the Molecular Biology and Sequencing Core, and the Flow Cytometry Core at the H. Lee Moffitt Cancer Center & Research Institute.
Author information
Authors and Affiliations
Contributions
J.L. designed and performed the experiments, analyzed and interpreted the data, performed statistical analyses, made the figures and wrote the manuscript. U.R. performed all chemical proteomics experiments, analyzed and interpreted the data, performed statistical analyses, made the figures and wrote the manuscript. B.F. performed all extracted ion chromatogram experiments, analyzed and interpreted the data, performed statistical analyses, made the figures and wrote the manuscript. A.E. provided statistical and bioinformatics support for interpreting LC-MS/MS data. S.E. advised on the research, performed statistical and bioinformatics support for interpreting LC-MS/MS data and critically read the manuscript. J.G. helped perform pY peptide purification from lung cancer cell lines. Y.B. provided technical assistance with cell lines. L.S. provided technical help with mutagenesis. K.L.B. analyzed chemical proteomics experiments and operated mass spectrometers. J.C. analyzed chemical proteomics experimental data and performed bioinformatics analyses. J.K. advised on the research, supervised all US-based proteomics experiments and critically read the manuscript. G.S.-F. supervised the chemical proteomics experiments, advised on the research and critically read the manuscript. E.B.H. had overall responsibility for this research and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–11 and Supplementary Tables 1 and 2 (PDF 1706 kb)
Rights and permissions
About this article
Cite this article
Li, J., Rix, U., Fang, B. et al. A chemical and phosphoproteomic characterization of dasatinib action in lung cancer. Nat Chem Biol 6, 291–299 (2010). https://doi.org/10.1038/nchembio.332
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.332
This article is cited by
-
Preclinical pharmacokinetic and pharmacodynamic evaluation of dasatinib and ponatinib for the treatment of T-cell acute lymphoblastic leukemia
Leukemia (2023)
-
Biomimetic design of highly flexible metal oxide nanofibrous membranes with exceptional mechanical performance for superior phosphopeptide enrichment
Nano Research (2023)
-
KOPI: Kinase inhibitOr Proteome Impact analysis
Scientific Reports (2022)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)
-
Evasion of apoptosis by myofibroblasts: a hallmark of fibrotic diseases
Nature Reviews Rheumatology (2020)