Abstract
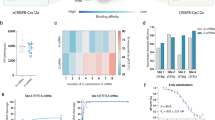
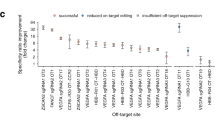
CRISPR–Cas9 is a versatile RNA-guided genome editing tool. Here we demonstrate that partial replacement of RNA nucleotides with DNA nucleotides in CRISPR RNA (crRNA) enables efficient gene editing in human cells. This strategy of partial DNA replacement retains on-target activity when used with both crRNA and sgRNA, as well as with multiple guide sequences. Partial DNA replacement also works for crRNA of Cpf1, another CRISPR system. We find that partial DNA replacement in the guide sequence significantly reduces off-target genome editing through focused analysis of off-target cleavage, measurement of mismatch tolerance and genome-wide profiling of off-target sites. Using the structure of the Cas9–sgRNA complex as a guide, the majority of the 3′ end of crRNA can be replaced with DNA nucleotide, and the 5 - and 3′-DNA-replaced crRNA enables efficient genome editing. Cas9 guided by a DNA–RNA chimera may provide a generalized strategy to reduce both the cost and the off-target genome editing in human cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
05 February 2018
In the HTML version of this article initially published online, the received date was incorrectly stated as 18 May 2017. The date should be 5 October 2017. This error has been corrected in the HTML version of the article.
References
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Doudna, J.A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cox, D.B., Platt, R.J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Swarts, D.C. et al. DNA-guided DNA interference by a prokaryotic Argonaute. Nature 507, 258–261 (2014).
Yuan, Y.R. et al. Crystal structure of A. aeolicus argonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol. Cell 19, 405–419 (2005).
Gabriel, R. et al. An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nat. Biotechnol. 29, 816–823 (2011).
Sander, J.D. et al. In silico abstraction of zinc finger nuclease cleavage profiles reveals an expanded landscape of off-target sites. Nucleic Acids Res. 41, e181 (2013).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Frock, R.L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nat. Biotechnol. 33, 179–186 (2015).
Kim, D. et al. Digenome-seq: genome-wide profiling of CRISPR–Cas9 off-target effects in human cells. Nat. Methods 12, 237–243 (2015).
Wang, X. et al. Unbiased detection of off-target cleavage by CRISPR-Cas9 and TALENs using integrase-defective lentiviral vectors. Nat. Biotechnol. 33, 175–178 (2015).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Tsai, S.Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Guilinger, J.P., Thompson, D.B. & Liu, D.R. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 32, 577–582 (2014).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Bolukbasi, M.F. et al. DNA-binding-domain fusions enhance the targeting range and precision of Cas9. Nat. Methods 12, 1150–1156 (2015).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Rahdar, M. et al. Synthetic CRISPR RNA-Cas9-guided genome editing in human cells. Proc. Natl. Acad. Sci. USA 112, E7110–E7117 (2015).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J.A. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Gilbert, L.A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Brinkman, E.K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Lesnik, E.A. & Freier, S.M. Relative thermodynamic stability of DNA, RNA, and DNA:RNA hybrid duplexes: relationship with base composition and structure. Biochemistry 34, 10807–10815 (1995).
Gyi, J.I., Lane, A.N., Conn, G.L. & Brown, T. The orientation and dynamics of the C2′-OH and hydration of RNA and DNA.RNA hybrids. Nucleic Acids Res. 26, 3104–3110 (1998).
Yin, H. et al. Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat. Biotechnol. 35, 1179–1187 (2017).
Yin, H. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat. Biotechnol. 34, 328–333 (2016).
Ran, F.A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Hou, Z. et al. Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitidis. Proc. Natl. Acad. Sci. USA 110, 15644–15649 (2013).
Lee, K. et al. Synthetically modified guide RNA and donor DNA are a versatile platform for CRISPR-Cas9 engineering. eLife 6, e25312 (2017).
Deleavey, G.F. & Damha, M.J. Designing chemically modified oligonucleotides for targeted gene silencing. Chem. Biol. 19, 937–954 (2012).
Suter, S.R. et al. Controlling miRNA-like off-target effects of an siRNA with nucleobase modifications. Org. Biomol. Chem. 15, 10029–10036 (2017).
Jackson, A.L. et al. Position-specific chemical modification of siRNAs reduces “off-target” transcript silencing. RNA 12, 1197–1205 (2006).
Chen, J.S. et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature 550, 407–410 (2017).
Kiani, S. et al. Cas9 gRNA engineering for genome editing, activation and repression. Nat. Methods 12, 1051–1054 (2015).
Dahlman, J.E. et al. Orthogonal gene knockout and activation with a catalytically active Cas9 nuclease. Nat. Biotechnol. 33, 1159–1161 (2015).
He, K., Chou, E.T., Begay, S., Anderson, E.M. & van Brabant Smith, A. Conjugation and evaluation of triazole-linked single guide RNA for CRISPR-Cas9 gene editing. ChemBioChem 17, 1809–1812 (2016).
Zou, J. et al. Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell 5, 97–110 (2009).
Sanjana, N.E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Zhu, L.J. et al. GUIDEseq: a bioconductor package to analyze GUIDE-seq datasets for CRISPR-Cas nucleases. BMC Genomics 18, 379 (2017).
Acknowledgements
We thank T. Jacks, P. Sharp, Z. Weng, C. Mello, and E. Sontheimer for discussions, and Y. Li for technical assistance. We thank K. Joung (Massachusetts General Hospital and Harvard Medical School) for sharing U2OS-GFP-PEST cells and J. Smith and A. Sheel for proofreading. This work is supported by grants from the National Institutes of Health (NIH), 5R00CA169512, DP2HL137167 and P01HL131471 (to W.X.). H.Y. is supported by 5-U54-CA151884-04 (NIH) and Skoltech Center. W.X. was supported by the Lung Cancer Research Foundation, Hyundai Hope on Wheels, and ALS Association. This work is supported in part by Cancer Center Support (core) grant P30-CA14051 from the NIH. We thank the Swanson Biotechnology Center at MIT for technical support.
Author information
Authors and Affiliations
Contributions
H.Y. conceived of and designed the study and directed the project. H.Y., C.-Q.S, S.S., S.W., Q.W., J.D., S.-Y.K., L.J.Z., and S.A.W. performed experiments and analyzed data. H.Y. made the figures with C.-Q.S. V.K., W.X., and R.L.B., and R.L. provided conceptual advice. H.Y. wrote the manuscript with comments from all authors. W.X., D.G.A. and R.L. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
H.Y., C.-Q.S., W.X., D.G.A and R.L. have applied for patents related to this study. D.G.A. is a scientific co-founder of CRISPR Therapeutics.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–2 and Supplementary Figures 1–6 (PDF 1362 kb)
Supplementary Data Set 1
Details of GUIDE-seq analysis of native and 10 DNA crRNAs of three endogenous genes (XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Yin, H., Song, CQ., Suresh, S. et al. Partial DNA-guided Cas9 enables genome editing with reduced off-target activity. Nat Chem Biol 14, 311–316 (2018). https://doi.org/10.1038/nchembio.2559
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2559
This article is cited by
-
CRISPR/Cas9: a powerful tool in colorectal cancer research
Journal of Experimental & Clinical Cancer Research (2023)
-
An overview of genome engineering in plants, including its scope, technologies, progress and grand challenges
Functional & Integrative Genomics (2023)
-
Clonal tracking in cancer and metastasis
Cancer and Metastasis Reviews (2023)
-
Genome-wide detection of CRISPR editing in vivo using GUIDE-tag
Nature Communications (2022)
-
FrCas9 is a CRISPR/Cas9 system with high editing efficiency and fidelity
Nature Communications (2022)