Abstract
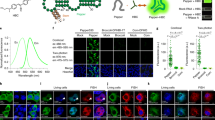
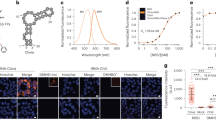
The discovery of functional RNAs that are critical for normal and disease physiology continues to expand at a breakneck pace. Many RNA functions are controlled by the formation of specific structures, and an understanding of each structural component is necessary to elucidate its function. Measuring solvent accessibility intracellularly with experimental ease is an unmet need in the field. Here, we present a novel method for probing nucleobase solvent accessibility, Light Activated Structural Examination of RNA (LASER). LASER depends on light activation of a small molecule, nicotinoyl azide (NAz), to measure solvent accessibility of purine nucleobases. In vitro, this technique accurately monitors solvent accessibility and identifies rapid structural changes resulting from ligand binding in a metabolite-responsive RNA. LASER probing can further identify cellular RNA–protein interactions and unique intracellular RNA structures. Our photoactivation technique provides an adaptable framework to structurally characterize solvent accessibility of RNA in many environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
22 January 2018
In the version of this article initially published online, the submission date was incorrectly stated as 21 June 2016. The correct date is 15 June 2017. The error has been corrected in the PDF and HTML versions of this article.
References
Mattick, J.S. The functional genomics of noncoding RNA. Science 309, 1527–1528 (2005).
Chappell, J. et al. The centrality of RNA for engineering gene expression. Biotechnol. J. 8, 1379–1395 (2013).
Sharp, P.A. The centrality of RNA. Cell 136, 577–580 (2009).
Tijerina, P., Mohr, S. & Russell, R. DMS footprinting of structured RNAs and RNA-protein complexes. Nat. Protoc. 2, 2608–2623 (2007).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
McGinnis, J.L., Duncan, C.D. & Weeks, K.M. High-throughput SHAPE and hydroxyl radical analysis of RNA structure and ribonucleoprotein assembly. Methods Enzymol. 468, 67–89 (2009).
Tullius, T.D. & Greenbaum, J.A. Mapping nucleic acid structure by hydroxyl radical cleavage. Curr. Opin. Chem. Biol. 9, 127–134 (2005).
Adilakshmi, T., Lease, R.A. & Woodson, S.A. Hydroxyl radical footprinting in vivo: mapping macromolecular structures with synchrotron radiation. Nucleic Acids Res. 34, e64 (2006).
Ramaswamy, P. & Woodson, S.A. S16 throws a conformational switch during assembly of 30S 5′ domain. Nat. Struct. Mol. Biol. 16, 438–445 (2009).
Batey, R.T., Rambo, R.P. & Doudna, J.A. Tertiary motifs in RNA structure and folding. Angew. Chem. Int. Ed. Engl. 38, 2326–2343 (1999).
Moras, D. & Poterszman, A. Getting into the major groove. Protein-RNA interactions. Curr. Biol. 6, 530–532 (1996).
Varani, L., Spillantini, M.G., Goedert, M. & Varani, G. Structural basis for recognition of the RNA major groove in the tau exon 10 splicing regulatory element by aminoglycoside antibiotics. Nucleic Acids Res. 28, 710–719 (2000).
Chen, L. & Frankel, A.D. A peptide interaction in the major groove of RNA resembles protein interactions in the minor groove of DNA. Proc. Natl. Acad. Sci. USA 92, 5077–5081 (1995).
Lawley, P.D. & Brookes, P. Further studies on the alkylation of nucleic acids and their constituent nucleotides. Biochem. J. 89, 127–138 (1963).
Lee, B. et al. Comparison of SHAPE reagents for mapping RNA structures inside living cells. RNA 23, 169–174 (2017).
Klán, P. et al. Photoremovable protecting groups in chemistry and biology: reaction mechanisms and efficacy. Chem. Rev. 113, 119–191 (2013).
Baker, A.S. & Deiters, A. Optical control of protein function through unnatural amino acid mutagenesis and other optogenetic approaches. ACS Chem. Biol. 9, 1398–1407 (2014).
Song, C.-X. & He, C. Bioorthogonal labeling of 5-hydroxymethylcytosine in genomic DNA and diazirine-based DNA photo-cross-linking probes. Acc. Chem. Res. 44, 709–717 (2011).
Wild, D. A novel pathway to the ultimate mutagens of aromatic amino and nitro compounds. Environ. Health Perspect. 88, 27–31 (1990).
Xue, J., Du, L., Zhu, R., Huang, J. & Phillips, D.L. Direct time-resolved spectroscopic observation of arylnitrenium ion reactions with guanine-containing DNA oligomers. J. Org. Chem. 79, 3610–3614 (2014).
Kuska, M.S. et al. Structural influence of C8-phenoxy-guanine in the NarI recognition DNA sequence. Chem. Res. Toxicol. 26, 1397–1408 (2013).
Voskresenska, V. et al. Photoaffinity labeling via nitrenium ion chemistry: protonation of the nitrene derived from 4-amino-3-nitrophenyl azide to afford reactive nitrenium ion pairs. J. Am. Chem. Soc. 131, 11535–11547 (2009).
Loeb, L.A. & Harris, C.C. Advances in chemical carcinogenesis: a historical review and prospective. Cancer Res. 68, 6863–6872 (2008).
Brachet, E., Ghosh, T., Ghosh, I. & Konig, B. Visible light C-H amidation of heteroarenes with benzoyl azides. Chem. Sci. 6, 987–992 (2015).
Kubicki, J. et al. Direct observation of acyl azide excited states and their decay processes by ultrafast time resolved infrared spectroscopy. J. Am. Chem. Soc. 131, 4212–4213 (2009).
Spitale, R.C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Desikan, V., Liu, Y., Toscano, J.P. & Jenks, W.S. Photochemistry of sulfilimine-based nitrene precursors: generation of both singlet and triplet benzoylnitrene. J. Org. Chem. 72, 6848–6859 (2007).
Humphreys, W.G., Kadlubar, F.F. & Guengerich, F.P. Mechanism of C8 alkylation of guanine residues by activated arylamines: evidence for initial adduct formation at the N7 position. Proc. Natl. Acad. Sci. USA 89, 8278–8282 (1992).
Reha, D. et al. Intercalators. 1. Nature of stacking interactions between intercalators (ethidium, daunomycin, ellipticine, and 4′,6-diaminide-2-phenylindole) and DNA base pairs. Ab initio quantum chemical, density functional theory, and empirical potential study. J. Am. Chem. Soc. 124, 3366–3376 (2002).
Wentrup, C., Reisinger, A. & Kvaskoff, D. 4-Pyridylnitrene and 2-pyrazinylcarbene. Beilstein J. Org. Chem. 9, 754–760 (2013).
Montange, R.K. & Batey, R.T. Structure of the S-adenosylmethionine riboswitch regulatory mRNA element. Nature 441, 1172–1175 (2006).
Winkler, W.C., Nahvi, A., Sudarsan, N., Barrick, J.E. & Breaker, R.R. An mRNA structure that controls gene expression by binding S-adenosylmethionine. Nat. Struct. Biol. 10, 701–707 (2003).
Hennelly, S.P. & Sanbonmatsu, K.Y. Tertiary contacts control switching of the SAM-I riboswitch. Nucleic Acids Res. 39, 2416–2431 (2011).
Heppell, B. et al. Molecular insights into the ligand-controlled organization of the SAM-I riboswitch. Nat. Chem. Biol. 7, 384–392 (2011).
Stoddard, C.D. et al. Free state conformational sampling of the SAM-I riboswitch aptamer domain. Structure 18, 787–797 (2010).
Mortimer, S.A., Johnson, J.S. & Weeks, K.M. Quantitative analysis of RNA solvent accessibility by N-silylation of guanosine. Biochemistry 48, 2109–2114 (2009).
Adams, P.L. et al. Crystal structure of a group I intron splicing intermediate. RNA 10, 1867–1887 (2004).
Kubota, M., Tran, C. & Spitale, R.C. Progress and challenges for chemical probing of RNA structure inside living cells. Nat. Chem. Biol. 11, 933–941 (2015).
Khatter, H., Myasnikov, A.G., Natchiar, S.K. & Klaholz, B.P. Structure of the human 80S ribosome. Nature 520, 640–645 (2015).
So, B.R. et al. A U1 snRNP-specific assembly pathway reveals the SMN complex as a versatile hub for RNP exchange. Nat. Struct. Mol. Biol. 23, 225–230 (2016).
Du, H. & Rosbash, M. The U1 snRNP protein U1C recognizes the 5′ splice site in the absence of base pairing. Nature 419, 86–90 (2002).
Kondo, Y., Oubridge, C., van Roon, A.M. & Nagai, K. Crystal structure of human U1 snRNP, a small nuclear ribonucleoprotein particle, reveals the mechanism of 5′ splice site recognition. eLife 4, e04986 (2015).
McConnell, T.S., Lokken, R.P. & Steitz, J.A. Assembly of the U1 snRNP involves interactions with the backbone of the terminal stem of U1 snRNA. RNA 9, 193–201 (2003).
Das, R., Laederach, A., Pearlman, S.M., Herschlag, D. & Altman, R.B. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005).
Tao, J., Perdew, J.P., Staroverov, V.N. & Scuseria, G.E. Climbing the density functional ladder: nonempirical meta-generalized gradient approximation designed for molecules and solids. Phys. Rev. Lett. 91, 146401 (2003).
Weigend, F. & Ahlrichs, R. Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy. Phys. Chem. Chem. Phys. 7, 3297–3305 (2005).
Schäfer, A., Horn, H. & Ahlrichs, R. Fully optimized contracted Gaussian basis sets for atoms Li to Kr. J. Chem. Phys. 97, 2571–2577 (1992).
Schäfer, A., Huber, C. & Ahlrichs, R. Fully optimized contracted Gaussian basis sets of triple zeta valence quality for atoms Li to Kr. J. Chem. Phys. 100, 5829–5835 (1994).
Eichkorn, K., Weigend, F., Treutler, O. & Ahlrichs, R. Auxiliary basis sets for main row atoms and transition metals and their use to approximate Coulomb potentials. Theor. Chem. Acc. 97, 119–124 (1997).
Weigend, F., Furche, F. & Ahlrichs, R. Gaussian basis sets of quadruple zeta valence quality for atoms H–Kr. J. Chem. Phys. 119, 12753–12762 (2003).
Grimme, S., Antony, J., Ehrlich, S. & Krieg, H. A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J. Chem. Phys. 132, 154104 (2010).
Schäfer, A., Klamt, A., Sattel, D., Lohrenz, J.C. & Eckert, F. COSMO Implementation in TURBOMOLE: Extension of an efficient quantum chemical code towards liquid systems. Phys. Chem. Chem. Phys. 2, 2187–2193 (2000).
Grimme, S. Supramolecular binding thermodynamics by dispersion-corrected density functional theory. Chem. Eur. J. 18, 9955–9964 (2012).
Furche, F. et al. Turbomole. Wiley Interdiscip. Rev. Comput. Mol. Sci. 4, 91–100 (2014).
Sierka, M., Hogekamp, A. & Ahlrichs, R. Fast evaluation of the Coulomb potential for electron densities using multipole accelerated resolution of identity approximation. J. Chem. Phys. 118, 9136–9148 (2003).
Weigend, F. Accurate Coulomb-fitting basis sets for H to Rn. Phys. Chem. Chem. Phys. 8, 1057–1065 (2006).
Gritsan, N.P. & Pritchina, E.A. Are aroylnitrenes species with a singlet ground state? Mendeleev. Commun. 11, 94–95 (2001).
Pritchina, E.A. et al. Matrix isolation, time-resolved IR, and computational study of the photochemistry of benzoyl azide. Phys. Chem. Chem. Phys. 5, 1010–1018 (2003).
Staroverov, V.N., Scuseria, G.E., Tao, J. & Perdew, J.P. Comparative assessment of a new nonempirical density functional: Molecules and hydrogen-bonded complexes. J. Chem. Phys. 119, 12129–12137 (2003).
Acknowledgements
We thank members of the Spitale lab for their careful reading and critique of the manuscript. The Spitale lab is supported by startup funds from the University of California, Irvine, and the NIH (1DP2GM119164 RCS) and 1RO1MH109588 (RCS). F.F. and the computational work are supported by the US Department of Energy under Award DE-SC0008694. C.M.H. acknowledges financial support from the National Science Foundation (DMR-1212842 and CHE-1609889), as well as generous allocations of computational resources at the Ohio Supercomputer Center. Femtosecond TRIR experiments were performed at The Ohio State University's Center for Chemical and Biophysical Dynamics.
Author information
Authors and Affiliations
Contributions
C.F. performed synthesis of the NAz probe and all positive controls, and did all analysis to determine reactivity. D.C. performed all RNA structure probing experiments. J.J. performed TRIR experiments with help from W.H.C. and C.M.H. M.M. performed density functional theory calculations and analyzed the results with C.F. and F.F. N.D. and I.R.C. assisted with HPLC and LC–MS experiments. J.J., W.H.C., and C.M.H. performed the CASSCF and B3LYP/6-31+G** using Gaussian09. C.F. and D.C. wrote the manuscript with help from R.C.S. All authors read and helped finalize the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–8, Supplementary Figures 1–16 (PDF 2533 kb)
Supplementary Data Set 1
Cartesian Coordinates of Structures presented in Supplementary Figures 1 and 2. (XLSX 69 kb)
Supplementary Data Set 2
Cartesian Coordinates of Structures presented in Supplementary Figures 3–6. (XLSX 47 kb)
Supplementary Data Set 3
LASER (+SAM) and corresponding solvent accessibility measurements at C8. Values are in A2. Solvent was contoured to 2 Å. (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Feng, C., Chan, D., Joseph, J. et al. Light-activated chemical probing of nucleobase solvent accessibility inside cells. Nat Chem Biol 14, 276–283 (2018). https://doi.org/10.1038/nchembio.2548
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2548
This article is cited by
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
Keth-seq for transcriptome-wide RNA structure mapping
Nature Chemical Biology (2020)
-
Epitranscriptomic technologies and analyses
Science China Life Sciences (2020)
-
A LASER-focused view into cells
Nature Chemical Biology (2018)
-
High-throughput determination of RNA structures
Nature Reviews Genetics (2018)