Abstract
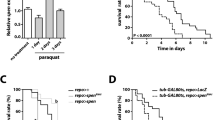
Paraquat, a herbicide linked to Parkinson's disease, generates reactive oxygen species (ROS), which causes cell death. Because the source of paraquat-induced ROS production remains unknown, we conducted a CRISPR-based positive-selection screen to identify metabolic genes essential for paraquat-induced cell death. Our screen uncovered three genes, POR (cytochrome P450 oxidoreductase), ATP7A (copper transporter), and SLC45A4 (sucrose transporter), required for paraquat–induced cell death. Furthermore, our results revealed POR as the source of paraquat-induced ROS production. Thus, our study highlights the use of functional genomic screens for uncovering redox biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dinis-Oliveira, R.J. et al. Paraquat poisonings: mechanisms of lung toxicity, clinical features, and treatment. Crit. Rev. Toxicol. 38, 13–71 (2008).
Suntres, Z.E. Role of antioxidants in paraquat toxicity. Toxicology 180, 65–77 (2002).
Blanco-Ayala, T., Andérica-Romero, A.C. & Pedraza-Chaverri, J. New insights into antioxidant strategies against paraquat toxicity. Free Radic. Res. 48, 623–640 (2014).
Kamel, F. Epidemiology. Paths from pesticides to Parkinson's. Science 341, 722–723 (2013).
Franco, R., Li, S., Rodriguez-Rocha, H., Burns, M. & Panayiotidis, M.I. Molecular mechanisms of pesticide-induced neurotoxicity: Relevance to Parkinson's disease. Chem. Biol. Interact. 188, 289–300 (2010).
Pezzoli, G. & Cereda, E. Exposure to pesticides or solvents and risk of Parkinson disease. Neurology 80, 2035–2041 (2013).
Tanner, C.M. et al. Rotenone, paraquat, and Parkinson's disease. Environ. Health Perspect. 119, 866–872 (2011).
Wang, A. et al. Parkinson's disease risk from ambient exposure to pesticides. Eur. J. Epidemiol. 26, 547–555 (2011).
Manning-Bog, A.B. et al. The herbicide paraquat causes up-regulation and aggregation of alpha-synuclein in mice: paraquat and alpha-synuclein. J. Biol. Chem. 277, 1641–1644 (2002).
McCormack, A.L. et al. Environmental risk factors and Parkinson's disease: selective degeneration of nigral dopaminergic neurons caused by the herbicide paraquat. Neurobiol. Dis. 10, 119–127 (2002).
Hong, S.Y., Lee, J.S., Sun, I.O., Lee, K.Y. & Gil, H.W. Prediction of patient survival in cases of acute paraquat poisoning. PLoS One 9, e111674 (2014).
Cochemé, H.M. & Murphy, M.P. Chapter 22 the uptake and interactions of the redox cycler paraquat with mitochondria. Methods Enzymol. 456, 395–417 (2009).
Cochemé, H.M. & Murphy, M.P. Complex I is the major site of mitochondrial superoxide production by paraquat. J. Biol. Chem. 283, 1786–1798 (2008).
Gray, J.P. et al. Paraquat increases cyanide-insensitive respiration in murine lung epithelial cells by activating an NAD(P)H:paraquat oxidoreductase: identification of the enzyme as thioredoxin reductase. J. Biol. Chem. 282, 7939–7949 (2007).
Shimada, H., Hirai, K., Simamura, E. & Pan, J. Mitochondrial NADH-quinone oxidoreductase of the outer membrane is responsible for paraquat cytotoxicity in rat livers. Arch. Biochem. Biophys. 351, 75–81 (1998).
Yee, C., Yang, W. & Hekimi, S. The intrinsic apoptosis pathway mediates the pro-longevity response to mitochondrial ROS in C. elegans. Cell 157, 897–909 (2014).
Shukla, A.K. et al. Heat shock protein-70 (Hsp-70) suppresses paraquat-induced neurodegeneration by inhibiting JNK and caspase-3 activation in Drosophila model of Parkinson's disease. PLoS One 9, e98886 (2014).
Rao, S.S. et al. Suberoylanilide hydroxamic acid attenuates paraquat-induced pulmonary fibrosis by preventing Smad7 from deacetylation in rats. J. Thorac. Dis. 8, 2485–2494 (2016).
Sun, I.O., Shin, S.H., Yoon, H.J. & Lee, K.Y. Predicting the probability of survival in acute paraquat poisoning. Kidney Res. Clin. Pract. 35, 102–106 (2016).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Wang, M. et al. Three-dimensional structure of NADPH-cytochrome P450 reductase: prototype for FMN- and FAD-containing enzymes. Proc. Natl. Acad. Sci. USA 94, 8411–8416 (1997).
Dierick, H.A., Adam, A.N., Escara-Wilke, J.F. & Glover, T.W. Immunocytochemical localization of the Menkes copper transport protein (ATP7A) to the trans-Golgi network. Hum. Mol. Genet. 6, 409–416 (1997).
Petris, M.J. & Mercer, J.F. The Menkes protein (ATP7A; MNK) cycles via the plasma membrane both in basal and elevated extracellular copper using a C-terminal di-leucine endocytic signal. Hum. Mol. Genet. 8, 2107–2115 (1999).
He, L., Vasiliou, K. & Nebert, D.W. Analysis and update of the human solute carrier (SLC) gene superfamily. Hum. Genomics 3, 195–206 (2009).
Rhee, S.G., Chang, T.S., Jeong, W. & Kang, D. Methods for detection and measurement of hydrogen peroxide inside and outside of cells. Mol. Cells 29, 539–549 (2010).
Robb, E.L. et al. Selective superoxide generation within mitochondria by the targeted redox cycler MitoParaquat. Free Radic. Biol. Med. 89, 883–894 (2015).
Huang, C.P., Fofana, M., Chan, J., Chang, C.J. & Howell, S.B. Copper transporter 2 regulates intracellular copper and sensitivity to cisplatin. Metallomics 6, 654–661 (2014).
Krall, J., Bagley, A.C., Mullenbach, G.T., Hallewell, R.A. & Lynch, R.E. Superoxide mediates the toxicity of paraquat for cultured mammalian cells. J. Biol. Chem. 263, 1910–1914 (1988).
Kuo, Y.M., Zhou, B., Cosco, D. & Gitschier, J. The copper transporter CTR1 provides an essential function in mammalian embryonic development. Proc. Natl. Acad. Sci. USA 98, 6836–6841 (2001).
Zelko, I.N., Mariani, T.J. & Folz, R.J. Superoxide dismutase multigene family: a comparison of the CuZn-SOD (SOD1), Mn-SOD (SOD2), and EC-SOD (SOD3) gene structures, evolution, and expression. Free Radic. Biol. Med. 33, 337–349 (2002).
Van Remmen, H. et al. Multiple deficiencies in antioxidant enzymes in mice result in a compound increase in sensitivity to oxidative stress. Free Radic. Biol. Med. 36, 1625–1634 (2004).
Saito, M., Thomas, C.E. & Aust, S.D. Paraquat and ferritin-dependent lipid peroxidation. J. Free Radic. Biol. Med. 1, 179–185 (1985).
Han, J.F., Wang, S.L., He, X.Y., Liu, C.Y. & Hong, J.Y. Effect of genetic variation on human cytochrome p450 reductase-mediated paraquat cytotoxicity. Toxicol. Sci. 91, 42–48 (2006).
Kelner, M.J. & Bagnell, R. Paraquat resistance associated with reduced NADPH reductase in an energy-dependent paraquat-accumulating cell line. Arch. Biochem. Biophys. 274, 366–374 (1989).
Clejan, L. & Cederbaum, A.I. Synergistic interactions between NADPH-cytochrome P-450 reductase, paraquat, and iron in the generation of active oxygen radicals. Biochem. Pharmacol. 38, 1779–1786 (1989).
Dinis-Oliveira, R.J. et al. Paraquat exposure as an etiological factor of Parkinson's disease. Neurotoxicology 27, 1110–1122 (2006).
Ishihara, Y., Shiba, D. & Shimamoto, N. Enhancement of DMNQ-induced hepatocyte toxicity by cytochrome P450 inhibition. Toxicol. Appl. Pharmacol. 214, 109–117 (2006).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Acknowledgements
We thank the Genome Technology Core of the Whitehead Institute for deep sequencing the genomic DNA samples. We thank the Flow Cytometry Core of Northwestern University for providing the BD LSRFortessa cell analyzer to assess cell viability. We are grateful to M.P. Murphy (University of Cambridge) for providing us MitoParaquat. This work was supported by a National Institute of Aging grant (5P01AG049665) to N.S.C. C.R.R. was supported by a National Institutes of Health postdoctoral training grant (T32 HL076139-11). H.K. was supported by a National Institutes of Health pre-doctoral training grant (T32 CA9560-30). K.B. was supported by grants from the National Institutes of Health (K22 CA1936600), a Searle Scholar Award, an Irma T. Hirschl/Monique Weill–Caulier Trust Award, and the Sidney Kimmel Foundation. D.M.S. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.R.R., K.B., D.M.S., and N.S.C. initiated the project and designed the research plan. C.R.R. and K.B. conducted the PQ positive- and negative-selection CRISPR-based screens. T.W. performed the bioinformatics analysis of the deep sequencing data. C.R.R. and H.K. generated clonal knockout and cDNA overexpression Jurkat and A549 cell lines. C.R.R. and H.K. prepared genomic DNA for sequencing the knockout clones. H.K. performed all western blot analyses and the A549 viability assays. C.R.R. assessed Jurkat cell viability. I.M.-R. performed the ROS measurements using CM-H2DCFDA and Amplex red and assessed SOD1 activity. P.G. performed the mass spectrometry experiments to assess PQ uptake. C.R.R. and N.S.C. wrote the manuscript, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–2, Supplementary Figures 1–10 (PDF 22728 kb)
Rights and permissions
About this article
Cite this article
Reczek, C., Birsoy, K., Kong, H. et al. A CRISPR screen identifies a pathway required for paraquat-induced cell death. Nat Chem Biol 13, 1274–1279 (2017). https://doi.org/10.1038/nchembio.2499
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2499
This article is cited by
-
α-lipoic acid modulates prostate cancer cell growth and bone cell differentiation
Scientific Reports (2024)
-
Photosynthesis, Salicylic Acid Content and Enzyme Activity of Triticum aestivum L. Influenced by the Use of a Seaweed Biostimulant Based on Solieria chordalis
Journal of Plant Growth Regulation (2024)
-
Mitochondrial electron transport chain is necessary for NLRP3 inflammasome activation
Nature Immunology (2022)
-
Cannabinoid Analogue WIN 55212–2 Protects Paraquat-Induced Lung Injury and Enhances Macrophage M2 Polarization
Inflammation (2022)
-
Photocatalytic degradation of paraquat dichloride in the presence of ZnO.WO3 composite
International Journal of Environmental Science and Technology (2022)