Abstract
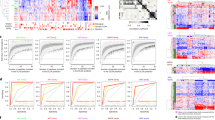
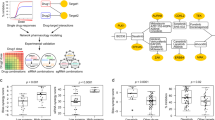
Targeted drugs are effective when they directly inhibit strong disease drivers, but only a small fraction of diseases feature defined actionable drivers. Alternatively, network-based approaches can uncover new therapeutic opportunities. Applying an integrated phenotypic screening, chemical and phosphoproteomics strategy, here we describe the anaplastic lymphoma kinase (ALK) inhibitor ceritinib as having activity across several ALK-negative lung cancer cell lines and identify new targets and network-wide signaling effects. Combining pharmacological inhibitors and RNA interference revealed a polypharmacology mechanism involving the noncanonical targets IGF1R, FAK1, RSK1 and RSK2. Mutating the downstream signaling hub YB1 protected cells from ceritinib. Consistent with YB1 signaling being known to cause taxol resistance, combination of ceritinib with paclitaxel displayed strong synergy, particularly in cells expressing high FAK autophosphorylation, which we show to be prevalent in lung cancer. Together, we present a systems chemical biology platform for elucidating multikinase inhibitor polypharmacology mechanisms, subsequent design of synergistic drug combinations, and identification of mechanistic biomarker candidates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sawyers, C.L. et al. Imatinib induces hematologic and cytogenetic responses in patients with chronic myelogenous leukemia in myeloid blast crisis: results of a phase II study. Blood 99, 3530–3539 (2002).
Kwak, E.L. et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med. 363, 1693–1703 (2010).
Flanagan, M.E. et al. Discovery of CP-690,550: a potent and selective Janus kinase (JAK) inhibitor for the treatment of autoimmune diseases and organ transplant rejection. J. Med. Chem. 53, 8468–8484 (2010).
Palla, G., Derényi, I., Farkas, I. & Vicsek, T. Uncovering the overlapping community structure of complex networks in nature and society. Nature 435, 814–818 (2005).
Farkas, I.J. et al. Network-based tools for the identification of novel drug targets. Sci. Signal. 4, pt3 (2011).
Hopkins, A.L. Network pharmacology: the next paradigm in drug discovery. Nat. Chem. Biol. 4, 682–690 (2008).
Barabási, A.-L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Paraiso, K.H.T. et al. Recovery of phospho-ERK activity allows melanoma cells to escape from BRAF inhibitor therapy. Br. J. Cancer 102, 1724–1730 (2010).
Knight, Z.A., Lin, H. & Shokat, K.M. Targeting the cancer kinome through polypharmacology. Nat. Rev. Cancer 10, 130–137 (2010).
Lombardo, L.J. et al. Discovery of N-(2-chloro-6-methyl- phenyl)-2-(6-(4-(2-hydroxyethyl)- piperazin-1-yl)-2-methylpyrimidin-4- ylamino)thiazole-5-carboxamide (BMS-354825), a dual Src/Abl kinase inhibitor with potent antitumor activity in preclinical assays. J. Med. Chem. 47, 6658–6661 (2004).
Rubbi, L. et al. Global phosphoproteomics reveals crosstalk between Bcr-Abl and negative feedback mechanisms controlling Src signaling. Sci. Signal. 4, ra18 (2011).
Frett, B. et al. Fragment-based discovery of a dual pan-RET/VEGFR2 kinase inhibitor optimized for single-agent polypharmacology. Angew. Chem. Int. Edn Engl. 54, 8717–8721 (2015).
Davis, M.I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046–1051 (2011).
Godl, K. et al. An efficient proteomics method to identify the cellular targets of protein kinase inhibitors. Proc. Natl. Acad. Sci. USA 100, 15434–15439 (2003).
Bantscheff, M. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat. Biotechnol. 25, 1035–1044 (2007).
Ong, S.-E. et al. Identifying the proteins to which small-molecule probes and drugs bind in cells. Proc. Natl. Acad. Sci. USA 106, 4617–4622 (2009).
Remsing Rix, L.L. et al. GSK3 alpha and beta are new functionally relevant targets of tivantinib in lung cancer cells. ACS Chem. Biol. 9, 353–358 (2014).
Marsilje, T.H. et al. Synthesis, structure-activity relationships, and in vivo efficacy of the novel potent and selective anaplastic lymphoma kinase (ALK) inhibitor 5-chloro-N2-(2-isopropoxy-5-methyl-4-(piperidin-4-yl)phenyl)-N4-(2-(isopropylsulfonyl)phenyl)pyrimidine-2,4-diamine (LDK378) currently in phase 1 and phase 2 clinical trials. J. Med. Chem. 56, 5675–5690 (2013).
Sabbatini, P. et al. GSK1838705A inhibits the insulin-like growth factor-1 receptor and anaplastic lymphoma kinase and shows antitumor activity in experimental models of human cancers. Mol. Cancer Ther. 8, 2811–2820 (2009).
Shaw, A.T. et al. Ceritinib in ALK-rearranged non-small-cell lung cancer. N. Engl. J. Med. 370, 1189–1197 (2014).
Nishio, M. et al. Phase I study of ceritinib (LDK378) in Japanese patients with advanced, anaplastic lymphoma kinase-rearranged non-small-cell lung cancer or other tumors. J. Thorac. Oncol. 10, 1058–1066 (2015).
Friboulet, L. et al. The ALK inhibitor ceritinib overcomes crizotinib resistance in non-small cell lung cancer. Cancer Discov. 4, 662–673 (2014).
Fisher, T.L. & Blenis, J. Evidence for two catalytically active kinase domains in pp90rsk. Mol. Cell. Biol. 16, 1212–1219 (1996).
Bjørbaek, C., Zhao, Y. & Moller, D.E. Divergent functional roles for p90rsk kinase domains. J. Biol. Chem. 270, 18848–18852 (1995).
Vik, T.A. & Ryder, J.W. Identification of serine 380 as the major site of autophosphorylation of Xenopus pp90rsk. Biochem. Biophys. Res. Commun. 235, 398–402 (1997).
Romeo, Y., Zhang, X. & Roux, P.P. Regulation and function of the RSK family of protein kinases. Biochem. J. 441, 553–569 (2012).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Blondel, V.D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Creixell, P. et al. Kinome-wide decoding of network-attacking mutations rewiring cancer signaling. Cell 163, 202–217 (2015).
Wang, J., Chen, G., Li, M. & Pan, Y. Integration of breast cancer gene signatures based on graph centrality. BMC Syst. Biol. 5 (Suppl. 3), S10 (2011).
Linding, R. et al. NetworKIN: a resource for exploring cellular phosphorylation networks. Nucleic Acids Res. 36, D695–D699 (2008).
Clauset, A., Newman, M.E.J. & Moore, C. Finding community structure in very large networks. Phys. Rev. E 70, 066111 (2004).
Andersson, S., D'Arcy, P., Larsson, O. & Sehat, B. Focal adhesion kinase (FAK) activates and stabilizes IGF-1 receptor. Biochem. Biophys. Res. Commun. 387, 36–41 (2009).
Kang, Y. et al. Role of focal adhesion kinase in regulating YB-1-mediated paclitaxel resistance in ovarian cancer. J. Natl. Cancer Inst. 105, 1485–1495 (2013).
Shiota, M. et al. Targeting ribosomal S6 kinases/Y-box binding protein-1 signaling improves cellular sensitivity to taxane in prostate cancer. Prostate 74, 829–838 (2014).
Sumi, N.J., Kuenzi, B.M., Knezevic, C.E., Remsing Rix, L.L. & Rix, U. Chemoproteomics reveals novel protein and lipid kinase targets of clinical CDK4/6 inhibitors in lung cancer. ACS Chem. Biol. 10, 2680–2686 (2015).
Ott, G.R. et al. Discovery of clinical candidate CEP-37440, a selective inhibitor of focal adhesion kinase (FAK) and anaplastic lymphoma kinase (ALK). J. Med. Chem. 59, 7478–7496 (2016).
Konstantinidou, G. et al. RHOA-FAK is a required signaling axis for the maintenance of KRAS-driven lung adenocarcinomas. Cancer Discov. 3, 444–457 (2013).
de Hoon, M.J.L., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Ritz, C., Baty, F., Streibig, J.C. & Gerhard, D. Dose-response analysis using R. PLoS One 10, e0146021 (2015).
Ong, S.-E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Cox, J. et al. A practical guide to the MaxQuant computational platform for SILAC-based quantitative proteomics. Nat. Protoc. 4, 698–705 (2009).
Tyanova, S., Mann, M. & Cox, J. MaxQuant for in-depth analysis of large SILAC datasets. Methods Mol. Biol. 1188, 351–364 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Welsh, E.A., Eschrich, S.A., Berglund, A.E. & Fenstermacher, D.A. Iterative rank-order normalization of gene expression microarray data. BMC Bioinformatics 14, 153 (2013).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Zybailov, B. et al. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 5, 2339–2347 (2006).
Kuenzi, B.M. et al. APOSTL: an interactive galaxy pipeline for reproducible analysis of affinity proteomics data. J. Proteome Res. 15, 4747–4754 (2016).
Bastian, M., Heymann, S. & Jacomy Gephi: an open source software for exploring and manipulating networks. (Association for the Advancement of Artificial Intelligence, 2009).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal. Complex Syst. 2006, 1695 (2006).
Hanson, B. HiveR: 2D and 3D Hive Plots for R. R Package Version 0255 (2016).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag New York) 2009.
Landthaler, M. et al. Molecular characterization of human Argonaute-containing ribonucleoprotein complexes and their bound target mRNAs. RNA 14, 2580–2596 (2008).
Pratt, D. et al. NDEx, the network data exchange. Cell Syst. 1, 302–305 (2015).
Acknowledgements
This work was supported by the NIH/NCI R01 CA181746 (to U.R.), the NIH/NCI F99/K00 Predoctoral to Postdoctoral Transition Award F99 CA212456 (to B.M.K), the Moffitt NIH/NCI SPORE in Lung Cancer P50 CA119997 (to E.B.H.), Moffitt Pinellas Partners, and the H. Lee Moffitt Cancer Center and Research Institute. We wish to acknowledge the Moffitt Lung Cancer Center of Excellence and the Moffitt Chemical Biology (Chemistry Unit), Proteomics, Flow Cytometry, Molecular Genomics and Analytical Microscopy Core Facilities. Moffitt Core Facilities are supported by the National Cancer Institute (Award No. P30-CA076292) as a Cancer Center Support Grant. Proteomics is also supported by the Moffitt Foundation.
Author information
Authors and Affiliations
Contributions
B.M.K., L.L.R.R., J.M.K., E.B.H. and U.R. conceived and designed the project. L.L.R.R and F.K. conducted the drug screen, and B.M.K and U.R. analyzed the data. Chemistry was done by B.M.K. B.M.K. performed chemical proteomics experiments, and B.M.K and U.R. analyzed the data. Phosphoproteomics experiments were done by B.M.K. and B.F., and B.M.K., P.A.S., and U.R. analyzed the data. B.M.K., L.L.R.R. and A.T.B. performed the western blots. B.M.K. conducted bioinformatic and network analyses. SiRNA and rescue experiments were done by B.M.K. B.M.K. and L.L.R.R. performed synergy experiments. B.M.K. performed all IHC, T.A.B. scored the slides and B.M.K analyzed the data. B.M.K. and U.R. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–2 and Supplementary Figures 1–9. (PDF 4127 kb)
Supplementary Data Set 1
Chemical proteomics data set. Data was searched by Mascot and displayed are exclusive unique spectrum counts. A minimum of 2 exclusive unique spectra were required for protein ID. Data was filtered with 95% protein and peptide cutoffs. CT, competition. (XLSX 153 kb)
Supplementary Data Set 2
pY phosphoproteomics data set. Phosphotyrosine data was IRON normalized and filtered for PEP <= 0.1. Contaminants, reverse sequences and rows with all zero values were removed. (XLSX 116 kb)
Supplementary Data Set 3
pSTY phosphoproteomics data set. Global phosphoproteomics data was IRON normalized and filtered for PEP <= 0.1. Contaminants, reverse sequences and rows with all zero values were removed. (XLSX 737 kb)
Supplementary Data Set 4
KEGG pathways enriched in Modules 1–4. Pathway analysis was done using ClusterProfiler to search the KEGG database. (XLSX 13 kb)
Supplementary Data Set 5
ReKINect mutational profile analysis. ReKINect output was generated using H650 cell missense mutational data from the Cancer Cell Line Encyclopedia. (XLSX 15 kb)
Supplementary Data Set 6
KEGG pathways of mutated genes in Supplementary Data Set 5. Pathway analysis was done using ClusterProfiler to search the KEGG database. (XLSX 10 kb)
Supplementary Data Set 7
NetworKIN analysis for potential kinase–substrate interactions. Unfiltered NetworKIN output was generated from phosphosites present in ceritinib subnetwork in Figure 3b–d. (XLSX 278 kb)
Supplementary Data Set 8
pY peptide confirmation by extracted ion chromatogram (XIC). Phosphotyrosine peptide MS1 quantification was confirmed using skyline. (XLSX 14 kb)
Supplementary Data Set 9
pSTY peptide confirmation by XIC. Phosphopeptide MS1 quantification was confirmed using skyline. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Kuenzi, B., Remsing Rix, L., Stewart, P. et al. Polypharmacology-based ceritinib repurposing using integrated functional proteomics. Nat Chem Biol 13, 1222–1231 (2017). https://doi.org/10.1038/nchembio.2489
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2489
This article is cited by
-
PROTEOMAS: a workflow enabling harmonized proteomic meta-analysis and proteomic signature mapping
Journal of Cheminformatics (2023)
-
Ceritinib is a novel triple negative breast cancer therapeutic agent
Molecular Cancer (2022)
-
Progress in DNA Aptamers as Recognition Components for Protein Functional Regulation
Chemical Research in Chinese Universities (2022)
-
Mutations in ALK signaling pathways conferring resistance to ALK inhibitor treatment lead to collateral vulnerabilities in neuroblastoma cells
Molecular Cancer (2022)
-
Insulin-like growth factor receptor signaling in tumorigenesis and drug resistance: a challenge for cancer therapy
Journal of Hematology & Oncology (2020)